Methylation, in the chemical sciences, is the addition of a methyl group on a substrate, or the substitution of an atom by a methyl group. Methylation is a form of alkylation, with a methyl group replacing a hydrogen atom. These terms are commonly used in chemistry, biochemistry, soil science, and biology.
Nitrogenases are enzymes (EC 1.18.6.1EC 1.19.6.1) that are produced by certain bacteria, such as cyanobacteria (blue-green bacteria) and rhizobacteria. These enzymes are responsible for the reduction of nitrogen (N2) to ammonia (NH3). Nitrogenases are the only family of enzymes known to catalyze this reaction, which is a step in the process of nitrogen fixation. Nitrogen fixation is required for all forms of life, with nitrogen being essential for the biosynthesis of molecules (nucleotides, amino acids) that create plants, animals and other organisms. They are encoded by the Nif genes or homologs. They are related to protochlorophyllide reductase.
Iron–sulfur proteins are proteins characterized by the presence of iron–sulfur clusters containing sulfide-linked di-, tri-, and tetrairon centers in variable oxidation states. Iron–sulfur clusters are found in a variety of metalloproteins, such as the ferredoxins, as well as NADH dehydrogenase, hydrogenases, coenzyme Q – cytochrome c reductase, succinate – coenzyme Q reductase and nitrogenase. Iron–sulfur clusters are best known for their role in the oxidation-reduction reactions of electron transport in mitochondria and chloroplasts. Both Complex I and Complex II of oxidative phosphorylation have multiple Fe–S clusters. They have many other functions including catalysis as illustrated by aconitase, generation of radicals as illustrated by SAM-dependent enzymes, and as sulfur donors in the biosynthesis of lipoic acid and biotin. Additionally, some Fe–S proteins regulate gene expression. Fe–S proteins are vulnerable to attack by biogenic nitric oxide, forming dinitrosyl iron complexes. In most Fe–S proteins, the terminal ligands on Fe are thiolate, but exceptions exist.

Aconitase is an enzyme that catalyses the stereo-specific isomerization of citrate to isocitrate via cis-aconitate in the tricarboxylic acid cycle, a non-redox-active process.

Methionine synthase (MS, MeSe, MTR) is primarily responsible for the regeneration of methionine from homocysteine in most individuals. In humans it is encoded by the MTR gene (5-methyltetrahydrofolate-homocysteine methyltransferase). Methionine synthase forms part of the S-adenosylmethionine (SAMe) biosynthesis and regeneration cycle, and is the enzyme responsible for linking the cycle to one-carbon metabolism via the folate cycle. There are two primary forms of this enzyme, the Vitamin B12 (cobalamin)-dependent (MetH) and independent (MetE) forms, although minimal core methionine synthases that do not fit cleanly into either category have also been described in some anaerobic bacteria. The two dominant forms of the enzymes appear to be evolutionary independent and rely on considerably different chemical mechanisms. Mammals and other higher eukaryotes express only the cobalamin-dependent form. In contrast, the distribution of the two forms in Archaeplastida (plants and algae) is more complex. Plants exclusively possess the cobalamin-independent form, while algae have either one of the two, depending on species. Many different microorganisms express both the cobalamin-dependent and cobalamin-independent forms.

Methyltransferases are a large group of enzymes that all methylate their substrates but can be split into several subclasses based on their structural features. The most common class of methyltransferases is class I, all of which contain a Rossmann fold for binding S-Adenosyl methionine (SAM). Class II methyltransferases contain a SET domain, which are exemplified by SET domain histone methyltransferases, and class III methyltransferases, which are membrane associated. Methyltransferases can also be grouped as different types utilizing different substrates in methyl transfer reactions. These types include protein methyltransferases, DNA/RNA methyltransferases, natural product methyltransferases, and non-SAM dependent methyltransferases. SAM is the classical methyl donor for methyltransferases, however, examples of other methyl donors are seen in nature. The general mechanism for methyl transfer is a SN2-like nucleophilic attack where the methionine sulfur serves as the leaving group and the methyl group attached to it acts as the electrophile that transfers the methyl group to the enzyme substrate. SAM is converted to S-Adenosyl homocysteine (SAH) during this process. The breaking of the SAM-methyl bond and the formation of the substrate-methyl bond happen nearly simultaneously. These enzymatic reactions are found in many pathways and are implicated in genetic diseases, cancer, and metabolic diseases. Another type of methyl transfer is the radical S-Adenosyl methionine (SAM) which is the methylation of unactivated carbon atoms in primary metabolites, proteins, lipids, and RNA.
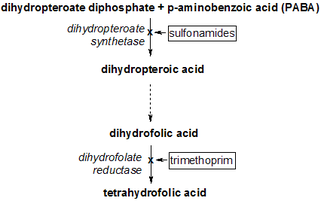
Dihydropteroate synthase (DHPS) is an enzyme classified under EC 2.5.1.15. It produces dihydropteroate in bacteria, but it is not expressed in most eukaryotes including humans. This makes it a useful target for sulfonamide antibiotics, which compete with the PABA precursor.
In enzymology, sarcosine dehydrogenase (EC 1.5.8.3) is a mitochondrial enzyme that catalyzes the chemical reaction N-demethylation of sarcosine to give glycine. This enzyme belongs to the family of oxidoreductases, specifically those acting on the CH-NH group of donor with other acceptors. The systematic name of this enzyme class is sarcosine:acceptor oxidoreductase (demethylating). Other names in common use include sarcosine N-demethylase, monomethylglycine dehydrogenase, and sarcosine:(acceptor) oxidoreductase (demethylating). Sarcosine dehydrogenase is closely related to dimethylglycine dehydrogenase, which catalyzes the demethylation reaction of dimethylglycine to sarcosine. Both sarcosine dehydrogenase and dimethylglycine dehydrogenase use FAD as a cofactor. Sarcosine dehydrogenase is linked by electron-transferring flavoprotein (ETF) to the respiratory redox chain. The general chemical reaction catalyzed by sarcosine dehydrogenase is:

Isopenicillin N synthase (IPNS) is a non-heme iron protein belonging to the 2-oxoglutarate (2OG)-dependent dioxygenases oxidoreductase family. This enzyme catalyzes the formation of isopenicillin N from δ-(L-α-aminoadipoyl)-L-cysteinyl-D-valine (LLD-ACV).
In enzymology, carbon monoxide dehydrogenase (CODH) (EC 1.2.7.4) is an enzyme that catalyzes the chemical reaction
Biotin synthase (BioB) is an enzyme that catalyzes the conversion of dethiobiotin (DTB) to biotin; this is the final step in the biotin biosynthetic pathway. Biotin, also known as vitamin B7, is a cofactor used in carboxylation, decarboxylation, and transcarboxylation reactions in many organisms including humans. Biotin synthase is an S-Adenosylmethionine (SAM) dependent enzyme that employs a radical mechanism to thiolate dethiobiotin, thus converting it to biotin.
5-methyltetrahydrosarcinapterin:corrinoid/iron-sulfur protein Co-methyltransferase is an enzyme with systematic name 5-methyltetrahydrosarcinapterin:corrinoid/iron-sulfur protein methyltransferase. This enzyme catalyses the following chemical reaction:
(Methyl-Co methylamine-specific corrinoid protein):coenzyme M methyltransferase is an enzyme with systematic name methylated monomethylamine-specific corrinoid protein:coenzyme M methyltransferase. This enzyme catalyses the following chemical reaction
Dimethylamine-corrinoid protein Co-methyltransferase is an enzyme with systematic name dimethylamine:5-hydroxybenzimidazolylcobamide Co-methyltransferase. This enzyme catalyses the following chemical reaction
Trimethylamine-corrinoid protein Co-methyltransferase is an enzyme with systematic name trimethylamine:5-hydroxybenzimidazolylcobamide Co-methyltransferase. This enzyme catalyses the following chemical reaction
Methylated-thiol-coenzyme M methyltransferase is an enzyme with systematic name methylated-thiol:coenzyme M methyltransferase. This enzyme catalyses the following chemical reaction:
(Methyl-Co tetramethylammonium-specific corrinoid protein):coenzyme M methyltransferase is an enzyme with systematic name methylated tetramethylammonium-specific corrinoid protein:coenzyme M methyltransferase. This enzyme catalyses the following chemical reaction

Acetyl-CoA synthase (ACS), not to be confused with acetyl-CoA synthetase or acetate-CoA ligase, is a nickel-containing enzyme involved in the metabolic processes of cells. Together with carbon monoxide dehydrogenase (CODH), it forms the bifunctional enzyme Acetyl-CoA Synthase/Carbon Monoxide Dehydrogenase (ACS/CODH) found in anaerobic microorganisms such as archaea and bacteria. The ACS/CODH enzyme works primarily through the Wood–Ljungdahl pathway which converts carbon dioxide to Acetyl-CoA. The recommended name for this enzyme is CO-methylating acetyl-CoA synthase.
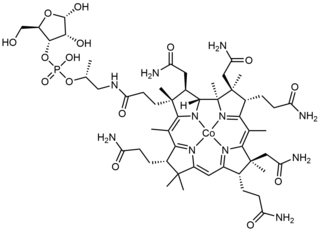
Cobamide is a naturally occurring chemical compound containing cobalt in the corrinoid family of macrocyclic complexes. Cobamide works as a coenzyme with some enzymes in bacteria. The cobalt atom may have a transferable methyl group attached. It is used for example in 5-methyltetrahydrosarcinapterin:corrinoid/iron-sulfur protein Co-methyltransferase.
Mercury methylation is the process of forming methylmercury (MeHg). The methylation of mercury can occur abiotically or biotically. Biotically, the primary methylators of mercury are sulfate-reducing and iron-reducing bacteria. Three mechanisms have been proposed for the biotic methylation of mercury by sulfate-reducing bacteria. Mercury methylation can be problematic as methylmercury is toxic and can be bio-magnified through the food web.









