Chymotrypsin (EC 3.4.21.1, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides. Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid (tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme. Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, particularly those containing leucine and methionine at the P1 position.
Hydrolysis is a term used for both an electro-chemical process and a biological one. The hydrolysis of water is the separation of water molecules into hydrogen and oxygen atoms using electricity (electrolysis).

A peptide bond is an amide type of covalent chemical bond linking two consecutive alpha-amino acids from C1 of one alpha-amino acid and N2 of another along a peptide or protein chain.

Protein primary structure is the linear sequence of amino acids in a peptide or protein. By convention, the primary structure of a protein is reported starting from the amino-terminal (N) end to the carboxyl-terminal (C) end. Protein biosynthesis is most commonly performed by ribosomes in cells. Peptides can also be synthesized in the laboratory. Protein primary structures can be directly sequenced, or inferred from DNA sequences.

Trypsin is a serine protease from the PA clan superfamily, found in the digestive system of many vertebrates, where it hydrolyzes proteins. Trypsin is formed in the small intestine when its proenzyme form, the trypsinogen produced by the pancreas, is activated. Trypsin cleaves peptide chains mainly at the carboxyl side of the amino acids lysine or arginine, except when either is followed by proline. It is used for numerous biotechnological processes. The process is commonly referred to as trypsin proteolysis or trypsinisation, and proteins that have been digested/treated with trypsin are said to have been trypsinized. Trypsin was discovered in 1876 by Wilhelm Kühne.

A microarray is a multiplex lab-on-a-chip. It is a two-dimensional array on a solid substrate that assays (tests) large amounts of biological material using high-throughput screening miniaturized, multiplexed and parallel processing and detection methods. The concept and methodology of microarrays was first introduced and illustrated in antibody microarrays by Tse Wen Chang in 1983 in a scientific publication and a series of patents. The "gene chip" industry started to grow significantly after the 1995 Science Paper by the Ron Davis and Pat Brown labs at Stanford University. With the establishment of companies, such as Affymetrix, Agilent, Applied Microarrays, Arrayjet, Illumina, and others, the technology of DNA microarrays has become the most sophisticated and the most widely used, while the use of protein, peptide and carbohydrate microarrays is expanding.
Twaron is a para-aramid. It is a heat-resistant and strong synthetic fibre developed in the early 1970s by the Dutch company AKZO, division ENKA, later Akzo Industrial Fibers. The research name of the para-aramid fibre was originally Fiber X, but it was soon called Arenka. Although the Dutch para-aramid fiber was developed only a little later than DuPont's Kevlar, introduction of Twaron as a commercial product came much later than Kevlar due to financial problems at the AKZO company in the 1970s.

Bremelanotide is a peptide melanocortin receptor agonist which is under development by Palatin Technologies as a treatment for female sexual dysfunction.

In organic chemistry, peptide synthesis is the production of peptides, compounds where multiple amino acids are linked via amide bonds, also known as peptide bonds. Peptides are chemically synthesized by the condensation reaction of the carboxyl group of one amino acid to the amino group of another. Protecting group strategies are usually necessary to prevent undesirable side reactions with the various amino acid side chains. Chemical peptide synthesis most commonly starts at the carboxyl end of the peptide (C-terminus), and proceeds toward the amino-terminus (N-terminus). Protein biosynthesis in living organisms occurs in the opposite direction.

Melanotan II is a synthetic analogue of the peptide hormone α-melanocyte-stimulating hormone (α-MSH).

Deamidation is a chemical reaction in which an amide functional group in the side chain of the amino acids asparagine or glutamine is removed or converted to another functional group. Typically, asparagine is converted to aspartic acid or isoaspartic acid. Glutamine is converted to glutamic acid or pyroglutamic acid (5-oxoproline). In a protein or peptide, these reactions are important because they may alter its structure, stability or function and may lead to protein degradation. The resulting mass adduct is a +1 Da shift.

Bentiromide is a peptide used as a screening test for exocrine pancreatic insufficiency and to monitor the adequacy of supplemental pancreatic therapy. It is given by mouth as a noninvasive test. The amount of 4-aminobenzoic acid and its metabolites excreted in the urine is taken as a measure of the chymotrypsin-secreting activity of the pancreas. Headache and gastrointestinal disturbances have been reported in patients taking bentiromide. Bentiromide is not available in the United States or Canada

Davydov soliton is a quantum quasiparticle representing an excitation propagating along the protein α-helix self-trapped amide I. It is a solution of the Davydov Hamiltonian. It is named for the Soviet and Ukrainian physicist Alexander Davydov. The Davydov model describes the interaction of the amide I vibrations with the hydrogen bonds that stabilize the α-helix of proteins. The elementary excitations within the α-helix are given by the phonons which correspond to the deformational oscillations of the lattice, and the excitons which describe the internal amide I excitations of the peptide groups. Referring to the atomic structure of an α-helix region of protein the mechanism that creates the Davydov soliton can be described as follows: vibrational energy of the C=O stretching oscillators that is localized on the α-helix acts through a phonon coupling effect to distort the structure of the α-helix, while the helical distortion reacts again through phonon coupling to trap the amide I oscillation energy and prevent its dispersion. This effect is called self-localization or self-trapping. Solitons in which the energy is distributed in a fashion preserving the helical symmetry are dynamically unstable, and such symmetrical solitons once formed decay rapidly when they propagate. On the other hand, an asymmetric soliton which spontaneously breaks the local translational and helical symmetries possesses the lowest energy and is a robust localized entity.
The Bergmann degradation is a series of chemical reactions designed to remove a single amino acid from the carboxylic acid (C-terminal) end of a peptide. First demonstrated by Max Bergmann in 1934, it is a rarely used method for sequencing peptides. The later developed Edman degradation is an improvement upon the Bergmann degradation, instead cleaving the N-terminal amino acid of peptides to produce a hydantoin containing the desired amino acid. The Bergmann degradation combines the azide degradation of the Curtius rearrangement with Bergmann and Zervas' carbobenzoxy method, which they designed to occur under relatively mild conditions so as to allow for peptide sequencing. A single round of the Bergmann degradation yields an aldehyde containing the sought after amino acid residue and the remaining fragment of the original peptide in amide form.

Pyroglutamylated RFamide peptide receptor also known as orexigenic neuropeptide QRFP receptor or G-protein coupled receptor 103 (GPR103) is a protein that in humans is encoded by the QRFPR gene.

Methylisopropyllysergamide is an analogue of LSD that was originally discovered by Albert Hofmann at Sandoz during the original structure-activity research into LSD. It has subsequently been investigated in more detail by the team led by David E. Nichols at Purdue University. Methylisopropyllysergamide is a structural isomer of LSD, with the alkyl groups on the amide nitrogen having been subjected to a methylene shuffle. MIPLA and its ethylisopropyl homologue are the only simple N,N-dialkyl lysergamides that approach the potency of LSD itself, being around 1/3-1/2 the potency of LSD, while all other dialkyl analogues tested are only around 1/10 as potent as LSD, although some N-monoalkyl lysergamides such as the sec-butyl and t-butyl derivatives were also found to show an activity profile and potency comparable to LSD, and the mono-isopropyl derivative is only slightly weaker than MIPLA. Apart from its lower potency, the hallucinogenic effects of methylisopropyllysergamide are similar to those of LSD itself, and the main use for this drug has been in studies of the binding site at the 5-HT2A receptor through which LSD exerts most of its pharmacological effects.
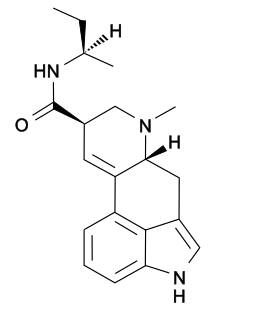
Lysergic acid 2-butyl amide (2-Butyllysergamide, LSB) is an analogue of LSD originally developed by Richard Pioch at Eli Lilly in the 1950s, but mostly publicised through research conducted by the team led by David E. Nichols at Purdue University. It is a structural isomer of LSD, with the two ethyl groups on the amide nitrogen having been replaced by a single sec-butyl group, joined at the 2-position. It is one of the few lysergamide derivatives to exceed the potency of LSD in animal drug discrimination assays, with the (R) isomer having an ED50 of 33nmol/kg for producing drug-appropriate responding, vs 48nmol/kg for LSD itself. The corresponding (R)-2-pentyl analogue has higher binding affinity for the 5-HT1A and 5-HT2A receptors, but is less potent in producing drug-appropriate responding, suggesting that the butyl compound has a higher efficacy at the receptor target. The drug discrimination assay for LSD in rats involves both 5-HT1A and 5-HT2A mediated components, and while lysergic acid 2-butyl amide is more potent than LSD as a 5-HT1A agonist, it is slightly less potent as a 5-HT2A agonist, and so would probably be slightly less potent than LSD as a hallucinogen in humans. The main use for this drug has been in studies of the binding site at the 5-HT2A receptor through which LSD exerts most of its pharmacological effects, with the stereoselective activity of these unsymmetric monoalkyl lysergamides foreshadowing the subsequent development of compounds such as lysergic acid 2,4-dimethylazetidide (LSZ).
Peptide T is an HIV entry inhibitor discovered in 1986 by researchers at the National Institutes of Health in the United States. Peptide T, and its modified analog Dala1-peptide T-amide (DAPTA), a drug in clinical trials, is a short peptide derived from the HIV envelope protein gp120 which blocks binding and infection of viral strains which use the CCR5 receptor to infect cells.

Dihexa, also known as N-hexanoic-Tyr-Ile-(6) aminohexanoic amide, is an oligopeptide drug derived from angiotensin IV that binds with high affinity to hepatocyte growth factor (HGF) and potentiates its activity at its receptor, c-Met. The compound has been found to potently improve cognitive function in animal models of Alzheimer's disease-like mental impairment. In an assay of neurotrophic activity, Dihexa was found to be seven orders of magnitude more potent than brain-derived neurotrophic factor.