
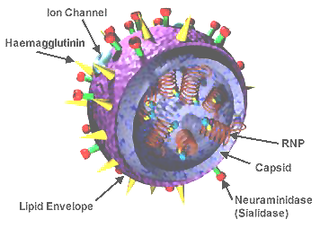
Influenza A virus (IAV) is a pathogen that causes the flu in birds and some mammals, including humans. It is an RNA virus whose subtypes have been isolated from wild birds. Occasionally, it is transmitted from wild to domestic birds, and this may cause severe disease, outbreaks, or human influenza pandemics.

Influenza hemagglutinin (HA) or haemagglutinin[p] is a homotrimeric glycoprotein found on the surface of influenza viruses and is integral to its infectivity.

Orthomyxoviridae is a family of negative-sense RNA viruses. It includes seven genera: Alphainfluenzavirus, Betainfluenzavirus, Gammainfluenzavirus, Deltainfluenzavirus, Isavirus, Thogotovirus, and Quaranjavirus. The first four genera contain viruses that cause influenza in birds and mammals, including humans. Isaviruses infect salmon; the thogotoviruses are arboviruses, infecting vertebrates and invertebrates. The Quaranjaviruses are also arboviruses, infecting vertebrates (birds) and invertebrates (arthropods).
Antigenic drift is a kind of genetic variation in viruses, arising from the accumulation of mutations in the virus genes that code for virus-surface proteins that host antibodies recognize. This results in a new strain of virus particles that is not effectively inhibited by the antibodies that prevented infection by previous strains. This makes it easier for the changed virus to spread throughout a partially immune population. Antigenic drift occurs in both influenza A and influenza B viruses.
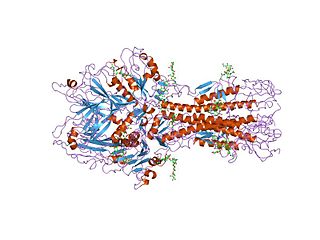
Hemagglutinin esterase (HEs) is a glycoprotein that certain enveloped viruses possess and use as an invading mechanism. HEs helps in the attachment and destruction of certain sialic acid receptors that are found on the host cell surface. Viruses that possess HEs include influenza C virus, toroviruses, and coronaviruses of the subgenus Embecovirus. HEs is a dimer transmembrane protein consisting of two monomers, each monomer is made of three domains. The three domains are: membrane fusion, esterase, and receptor binding domains.

Envelope glycoprotein GP120 is a glycoprotein exposed on the surface of the HIV envelope. It was discovered by Professors Tun-Hou Lee and Myron "Max" Essex of the Harvard School of Public Health in 1984. The 120 in its name comes from its molecular weight of 120 kDa. Gp120 is essential for virus entry into cells as it plays a vital role in attachment to specific cell surface receptors. These receptors are DC-SIGN, Heparan Sulfate Proteoglycan and a specific interaction with the CD4 receptor, particularly on helper T-cells. Binding to CD4 induces the start of a cascade of conformational changes in gp120 and gp41 that lead to the fusion of the viral membrane with the host cell membrane. Binding to CD4 is mainly electrostatic although there are van der Waals interactions and hydrogen bonds.

Thogotovirus is a genus of enveloped RNA viruses, one of seven genera in the virus family Orthomyxoviridae. Their single-stranded, negative-sense RNA genome has six or seven segments. Thogotoviruses are distinguished from most other orthomyxoviruses by being arboviruses – viruses that are transmitted by arthropods, in this case usually ticks. Thogotoviruses can replicate in both tick cells and vertebrate cells; one subtype has also been isolated from mosquitoes. A consequence of being transmitted by blood-sucking vectors is that the virus must spread systemically in the vertebrate host – unlike influenza viruses, which are transmitted by respiratory droplets and are usually confined to the respiratory system.

Influenza B virus is the only species in the genus Betainfluenzavirus in the virus family Orthomyxoviridae.

Influenza C virus is the only species in the genus Gammainfluenzavirus, in the virus family Orthomyxoviridae, which like other influenza viruses, causes influenza.

H5N1 genetic structure is the molecular structure of the H5N1 virus's RNA.

Murine coronavirus (M-CoV) is a virus in the genus Betacoronavirus that infects mice. Belonging to the subgenus Embecovirus, murine coronavirus strains are enterotropic or polytropic. Enterotropic strains include mouse hepatitis virus (MHV) strains D, Y, RI, and DVIM, whereas polytropic strains, such as JHM and A59, primarily cause hepatitis, enteritis, and encephalitis. Murine coronavirus is an important pathogen in the laboratory mouse and the laboratory rat. It is the most studied coronavirus in animals other than humans, and has been used as an animal disease model for many virological and clinical studies.
Antigenic variation or antigenic alteration refers to the mechanism by which an infectious agent such as a protozoan, bacterium or virus alters the proteins or carbohydrates on its surface and thus avoids a host immune response, making it one of the mechanisms of antigenic escape. It is related to phase variation. Antigenic variation not only enables the pathogen to avoid the immune response in its current host, but also allows re-infection of previously infected hosts. Immunity to re-infection is based on recognition of the antigens carried by the pathogen, which are "remembered" by the acquired immune response. If the pathogen's dominant antigen can be altered, the pathogen can then evade the host's acquired immune system. Antigenic variation can occur by altering a variety of surface molecules including proteins and carbohydrates. Antigenic variation can result from gene conversion, site-specific DNA inversions, hypermutation, or recombination of sequence cassettes. The result is that even a clonal population of pathogens expresses a heterogeneous phenotype. Many of the proteins known to show antigenic or phase variation are related to virulence.

Murine respirovirus, formerly Sendai virus (SeV) and previously also known as murine parainfluenza virus type 1 or hemagglutinating virus of Japan (HVJ), is an enveloped, 150-200 nm–diameter, negative sense, single-stranded RNA virus of the family Paramyxoviridae. It typically infects rodents and it is not pathogenic for humans or domestic animals

In virology, a spike protein or peplomer protein is a protein that forms a large structure known as a spike or peplomer projecting from the surface of an enveloped virus. The proteins are usually glycoproteins that form dimers or trimers.

Influenza, commonly known as "the flu", is an infectious disease caused by influenza viruses. Symptoms range from mild to severe and often include fever, runny nose, sore throat, muscle pain, headache, coughing, and fatigue. These symptoms begin from one to four days after exposure to the virus and last for about 2–8 days. Diarrhea and vomiting can occur, particularly in children. Influenza may progress to pneumonia, which can be caused by the virus or by a subsequent bacterial infection. Other complications of infection include acute respiratory distress syndrome, meningitis, encephalitis, and worsening of pre-existing health problems such as asthma and cardiovascular disease.
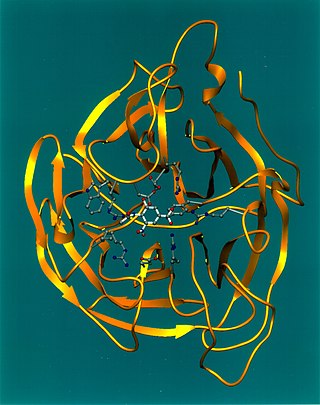
Viral neuraminidase is a type of neuraminidase found on the surface of influenza viruses that enables the virus to be released from the host cell. Neuraminidases are enzymes that cleave sialic acid groups from glycoproteins. Viral neuraminidase was discovered by Alfred Gottschalk at the Walter and Eliza Hall Institute in 1957. Neuraminidase inhibitors are antiviral agents that inhibit influenza viral neuraminidase activity and are of major importance in the control of influenza.

In molecular biology, hemagglutinins are receptor-binding membrane fusion glycoproteins produced by viruses in the Paramyxoviridae and Orthomyxoviridae families. Hemagglutinins are responsible for binding to receptors on red blood cells to initiate viral attachment and infection. The agglutination of red cells occurs when antibodies on one cell bind to those on others, causing amorphous aggregates of clumped cells.
A neutralizing antibody (NAb) is an antibody that defends a cell from a pathogen or infectious particle by neutralizing any effect it has biologically. Neutralization renders the particle no longer infectious or pathogenic. Neutralizing antibodies are part of the humoral response of the adaptive immune system against viruses, intracellular bacteria and microbial toxin. By binding specifically to surface structures (antigen) on an infectious particle, neutralizing antibodies prevent the particle from interacting with its host cells it might infect and destroy.
Virus quantification is counting or calculating the number of virus particles (virions) in a sample to determine the virus concentration. It is used in both research and development (R&D) in academic and commercial laboratories as well as in production situations where the quantity of virus at various steps is an important variable that must be monitored. For example, the production of virus-based vaccines, recombinant proteins using viral vectors, and viral antigens all require virus quantification to continually monitor and/or modify the process in order to optimize product quality and production yields and to respond to ever changing demands and applications. Other examples of specific instances where viruses need to be quantified include clone screening, multiplicity of infection (MOI) optimization, and adaptation of methods to cell culture.
A universal flu vaccine is a flu vaccine that is effective against all influenza strains regardless of the virus sub type, antigenic drift or antigenic shift. Hence it should not require modification from year to year. As of 2021 no universal flu vaccine had been approved for general use, several were in development, and one was in clinical trial.
















