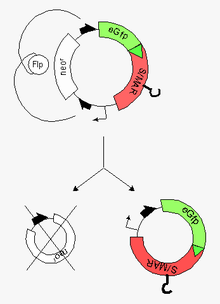
An episome is a special type of plasmid, which remains as a part of the eukaryotic genome without integration. Episomes manage this by replicating together with the rest of the genome and subsequently associating with metaphase chromosomes during mitosis. Episomes do not degrade, unlike standard plasmids, and can be designed so that they are not epigenetically silenced inside the eukaryotic cell nucleus. Episomes can be observed in nature in certain types of long-term infection by adeno-associated virus or Epstein-Barr virus. In 2004, it was proposed that non-viral episomes might be used in genetic therapy for long-term change in gene expression.
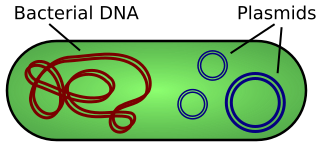
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; however, plasmids are sometimes present in archaea and eukaryotic organisms. In nature, plasmids often carry genes that benefit the survival of the organism and confer selective advantage such as antibiotic resistance. While chromosomes are large and contain all the essential genetic information for living under normal conditions, plasmids are usually very small and contain only additional genes that may be useful in certain situations or conditions. Artificial plasmids are widely used as vectors in molecular cloning, serving to drive the replication of recombinant DNA sequences within host organisms. In the laboratory, plasmids may be introduced into a cell via transformation. Synthetic plasmids are available for procurement over the internet.

Yeast artificial chromosomes (YACs) are genetically engineered chromosomes derived from the DNA of the yeast, Saccharomyces cerevisiae, which is then ligated into a bacterial plasmid. By inserting large fragments of DNA, from 100–1000 kb, the inserted sequences can be cloned and physically mapped using a process called chromosome walking. This is the process that was initially used for the Human Genome Project, however due to stability issues, YACs were abandoned for the use of bacterial artificial chromosome
A DNA construct is an artificially-designed segment of DNA borne on a vector that can be used to incorporate genetic material into a target tissue or cell. A DNA construct contains a DNA insert, called a transgene, delivered via a transformation vector which allows the insert sequence to be replicated and/or expressed in the target cell. This gene can be cloned from a naturally occurring gene, or synthetically constructed. The vector can be delivered using physical, chemical or viral methods. Typically, the vectors used in DNA constructs contain an origin of replication, a multiple cloning site, and a selectable marker. Certain vectors can carry additional regulatory elements based on the expression system involved.
Extrachromosomal DNA is any DNA that is found off the chromosomes, either inside or outside the nucleus of a cell. Most DNA in an individual genome is found in chromosomes contained in the nucleus. Multiple forms of extrachromosomal DNA exist, and, while some of these serve important biological functions, they can also play a role in diseases such as cancer.
Cre-Lox recombination is a site-specific recombinase technology, used to carry out deletions, insertions, translocations and inversions at specific sites in the DNA of cells. It allows the DNA modification to be targeted to a specific cell type or be triggered by a specific external stimulus. It is implemented both in eukaryotic and prokaryotic systems. The Cre-lox recombination system has been particularly useful to help neuroscientists to study the brain in which complex cell types and neural circuits come together to generate cognition and behaviors. NIH Blueprint for Neuroscience Research has created several hundreds of Cre driver mouse lines which are currently used by the worldwide neuroscience community.
Site-specific recombinase technologies are genome engineering tools that depend on recombinase enzymes to replace targeted sections of DNA.

In genetics, Flp-FRT recombination is a site-directed recombination technology, increasingly used to manipulate an organism's DNA under controlled conditions in vivo. It is analogous to Cre-lox recombination but involves the recombination of sequences between short flippase recognition target (FRT) sites by the recombinase flippase (Flp) derived from the 2 µ plasmid of baker's yeast Saccharomyces cerevisiae.

Gene delivery is the process of introducing foreign genetic material, such as DNA or RNA, into host cells. Gene delivery must reach the genome of the host cell to induce gene expression. Successful gene delivery requires the foreign gene delivery to remain stable within the host cell and can either integrate into the genome or replicate independently of it. This requires foreign DNA to be synthesized as part of a vector, which is designed to enter the desired host cell and deliver the transgene to that cell's genome. Vectors utilized as the method for gene delivery can be divided into two categories, recombinant viruses and synthetic vectors.
Fosmids are similar to cosmids but are based on the bacterial F-plasmid. The cloning vector is limited, as a host can only contain one fosmid molecule. Fosmids can hold DNA inserts of up to 40 kb in size; often the source of the insert is random genomic DNA. A fosmid library is prepared by extracting the genomic DNA from the target organism and cloning it into the fosmid vector. The ligation mix is then packaged into phage particles and the DNA is transfected into the bacterial host. Bacterial clones propagate the fosmid library. The low copy number offers higher stability than vectors with relatively higher copy numbers, including cosmids. Fosmids may be useful for constructing stable libraries from complex genomes. Fosmids have high structural stability and have been found to maintain human DNA effectively even after 100 generations of bacterial growth. Fosmid clones were used to help assess the accuracy of the Public Human Genome Sequence.
P1 is a temperate bacteriophage that infects Escherichia coli and some other bacteria. When undergoing a lysogenic cycle the phage genome exists as a plasmid in the bacterium unlike other phages that integrate into the host DNA. P1 has an icosahedral head containing the DNA attached to a contractile tail with six tail fibers. The P1 phage has gained research interest because it can be used to transfer DNA from one bacterial cell to another in a process known as transduction. As it replicates during its lytic cycle it captures fragments of the host chromosome. If the resulting viral particles are used to infect a different host the captured DNA fragments can be integrated into the new host's genome. This method of in vivo genetic engineering was widely used for many years and is still used today, though to a lesser extent. P1 can also be used to create the P1-derived artificial chromosome cloning vector which can carry relatively large fragments of DNA. P1 encodes a site-specific recombinase, Cre, that is widely used to carry out cell-specific or time-specific DNA recombination by flanking the target DNA with loxP sites.

A transplastomic plant is a genetically modified plant in which genes are inactivated, modified or new foreign genes are inserted into the DNA of plastids like the chloroplast instead of nuclear DNA.
A transfer DNA (T-DNA) binary system is a pair of plasmids consisting of a T-DNA binary vector and a virhelper plasmid. The two plasmids are used together to produce genetically modified plants. They are artificial vectors that have been derived from the naturally occurring Ti plasmid found in bacterial species of the genus Agrobacterium, such as A. tumefaciens. The binary vector is a shuttle vector, so-called because it is able to replicate in multiple hosts.
RMCE is a procedure in reverse genetics allowing the systematic, repeated modification of higher eukaryotic genomes by targeted integration, based on the features of site-specific recombination processes (SSRs). For RMCE, this is achieved by the clean exchange of a preexisting gene cassette for an analogous cassette carrying the "gene of interest" (GOI).
In molecular cloning, a vector is any particle used as a vehicle to artificially carry a foreign nucleic sequence – usually DNA – into another cell, where it can be replicated and/or expressed. A vector containing foreign DNA is termed recombinant DNA. The four major types of vectors are plasmids, viral vectors, cosmids, and artificial chromosomes. Of these, the most commonly used vectors are plasmids. Common to all engineered vectors are an origin of replication, a multicloning site, and a selectable marker.

Amphidinium is a genus of dinoflagellates. The type for the genus is Amphidinium operculatum Claparède & Lachmann. The genus includes the species Amphidinium carterae which is used as a model organism.
The Sleeping Beauty transposon system is a synthetic DNA transposon designed to introduce precisely defined DNA sequences into the chromosomes of vertebrate animals for the purposes of introducing new traits and to discover new genes and their functions. It is a Tc1/mariner-type system, with the transposase resurrected from multiple inactive fish sequences.

The term S/MAR, otherwise called SAR, or MAR, are sequences in the DNA of eukaryotic chromosomes where the nuclear matrix attaches. As architectural DNA components that organize the genome of eukaryotes into functional units within the cell nucleus, S/MARs mediate structural organization of the chromatin within the nucleus. These elements constitute anchor points of the DNA for the chromatin scaffold and serve to organize the chromatin into structural domains. Studies on individual genes led to the conclusion that the dynamic and complex organization of the chromatin mediated by S/MAR elements plays an important role in the regulation of gene expression.

Genetic engineering techniques allow the modification of animal and plant genomes. Techniques have been devised to insert, delete, and modify DNA at multiple levels, ranging from a specific base pair in a specific gene to entire genes. There are a number of steps that are followed before a genetically modified organism (GMO) is created. Genetic engineers must first choose what gene they wish to insert, modify, or delete. The gene must then be isolated and incorporated, along with other genetic elements, into a suitable vector. This vector is then used to insert the gene into the host genome, creating a transgenic or edited organism.
Self-complementary adeno-associated virus (scAAV) is a viral vector engineered from the naturally occurring adeno-associated virus (AAV) to be used as a tool for gene therapy. Use of recombinant AAV (rAAV) has been successful in clinical trials addressing a variety of diseases. This lab-made progeny of rAAV is termed "self-complementary" because the coding region has been designed to form an intra-molecular double-stranded DNA template. A rate-limiting step for the standard AAV genome involves the second-strand synthesis since the typical AAV genome is a single-stranded DNA template. However, this is not the case for scAAV genomes. Upon infection, rather than waiting for cell mediated synthesis of the second strand, the two complementary halves of scAAV will associate to form one double stranded DNA (dsDNA) unit that is ready for immediate replication and transcription. The caveat of this construct is that instead of the full coding capacity found in rAAV (4.7–6kb) scAAV can only hold about half of that amount (≈2.4kb).