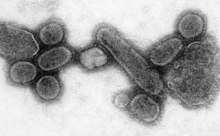
Influenza A virus (IAV) is a pathogen that causes the flu in birds and some mammals, including humans. It is an RNA virus whose subtypes have been isolated from wild birds. Occasionally, it is transmitted from wild to domestic birds, and this may cause severe disease, outbreaks, or human influenza pandemics.

Avian influenza, also known as avian flu, is a bird flu caused by the influenza A virus, which can infect people. It is similar to other types of animal flu in that it is caused by a virus strain that has adapted to a specific host. The type with the greatest risk is highly pathogenic avian influenza (HPAI).

Antigenic shift is the process by which two or more different strains of a virus, or strains of two or more different viruses, combine to form a new subtype having a mixture of the surface antigens of the two or more original strains. The term is often applied specifically to influenza, as that is the best-known example, but the process is also known to occur with other viruses, such as visna virus in sheep. Antigenic shift is a specific case of reassortment or viral shift that confers a phenotypic change.

Influenza hemagglutinin (HA) or haemagglutinin[p] is a homotrimeric glycoprotein found on the surface of influenza viruses and is integral to its infectivity.

Orthomyxoviridae is a family of negative-sense RNA viruses. It includes seven genera: Alphainfluenzavirus, Betainfluenzavirus, Gammainfluenzavirus, Deltainfluenzavirus, Isavirus, Thogotovirus, and Quaranjavirus. The first four genera contain viruses that cause influenza in birds and mammals, including humans. Isaviruses infect salmon; the thogotoviruses are arboviruses, infecting vertebrates and invertebrates. The Quaranjaviruses are also arboviruses, infecting vertebrates (birds) and invertebrates (arthropods).
Antigenic drift is a kind of genetic variation in viruses, arising from the accumulation of mutations in the virus genes that code for virus-surface proteins that host antibodies recognize. This results in a new strain of virus particles that is not effectively inhibited by the antibodies that prevented infection by previous strains. This makes it easier for the changed virus to spread throughout a partially immune population. Antigenic drift occurs in both influenza A and influenza B viruses.

Influenza A virus subtype H5N1 (A/H5N1) is a subtype of the influenza A virus which can cause illness in humans and many other species. A bird-adapted strain of H5N1, called HPAI A(H5N1) for highly pathogenic avian influenza virus of type A of subtype H5N1, is the highly pathogenic causative agent of H5N1 flu, commonly known as avian influenza. It is enzootic in many bird populations, especially in Southeast Asia. One strain of HPAI A(H5N1) is spreading globally after first appearing in Asia. It is epizootic and panzootic, killing tens of millions of birds and spurring the culling of hundreds of millions of others to stem its spread. Many references to "bird flu" and H5N1 in the popular media refer to this strain.

Swine influenza is an infection caused by any of several types of swine influenza viruses. Swine influenza virus (SIV) or swine-origin influenza virus (S-OIV) refers to any strain of the influenza family of viruses that is endemic in pigs. As of 2009, identified SIV strains include influenza C and the subtypes of influenza A known as H1N1, H1N2, H2N1, H3N1, H3N2, and H2N3.

In virology, influenza A virus subtype H1N1 (A/H1N1) is a subtype of influenza A virus. Major outbreaks of H1N1 strains in humans include the 1918 Spanish flu pandemic, the 1977 Russian flu pandemic and the 2009 swine flu pandemic. It is an orthomyxovirus that contains the glycoproteins hemagglutinin (H) and neuraminidase (N), antigens whose subtypes are used to classify the strains of the virus as H1N1, H1N2 etc. Hemagglutinin causes red blood cells to clump together and binds the virus to the infected cell. Neuraminidase is a type of glycoside hydrolase enzyme which helps to move the virus particles through the infected cell and assist in budding from the host cells.

An influenza pandemic is an epidemic of an influenza virus that spreads across a large region and infects a large proportion of the population. There have been six major influenza epidemics in the last 140 years, with the 1918 flu pandemic being the most severe; this is estimated to have been responsible for the deaths of 50–100 million people. The 2009 swine flu pandemic, resulted in under 300,000 deaths and is considered relatively mild. These pandemics occur irregularly.

Influenza A virus subtype H2N2 (A/H2N2) is a subtype of Influenza A virus. H2N2 has mutated into various strains including the "Asian flu" strain, H3N2, and various strains found in birds. It is also suspected of causing a human pandemic in 1889. The geographic spreading of the 1889 Russian flu has been studied and published.

Transmission and infection of H5N1 from infected avian sources to humans has been a concern since the first documented case of human infection in 1997, due to the global spread of H5N1 that constitutes a pandemic threat.

H5N1 genetic structure is the molecular structure of the H5N1 virus's RNA.

Fujian flu refers to flu caused by either a Fujian human flu strain of the H3N2 subtype of the Influenza A virus or a Fujian bird flu strain of the H5N1 subtype of the Influenza A virus. These strains are named after Fujian, a coastal province in Southeast China.

Human mortality from H5N1 or the human fatality ratio from H5N1 or the case-fatality rate of H5N1 is the ratio of the number of confirmed human deaths resulting from confirmed cases of transmission and infection of H5N1 to the number of those confirmed cases. For example, if there are 100 confirmed cases of humans infected with H5N1 and 10 die, then there is a 10% human fatality ratio. H5N1 flu is a concern due to the global spread of H5N1 that constitutes a pandemic threat. The majority of H5N1 flu cases have been reported in southeast and east Asia. The case-fatality rate is central to pandemic planning. Estimates of case-fatality (CF) rates for past influenza pandemics have ranged from to 2-3% for the 1918 pandemic to about 0.6% for the 1957 pandemic to 0.2% for the 1968 pandemic. As of 2008, the official World Health Organization estimate for the case-fatality rate for the outbreak of H5N1 avian influenza was approximately 60%. Public health officials in Ontario, Canada argue that the true case-fatality rate could be lower, pointing to studies suggesting it could be 14-33%, and warned that it was unlikely to be as low as the 0.1–0.4% rate that was built into many pandemic plans.
Jeffery K. Taubenberger is an American virologist. With Ann Reid, he was the first to sequence the genome of the influenza virus which caused the 1918 pandemic of Spanish flu. He is Chief of the Viral Pathogenesis and Evolution Section, Laboratory of Infectious Diseases, National Institute of Allergy and Infectious Diseases, National Institutes of Health. Taubenberger's laboratory studies a number of viruses, including influenza A viruses (IAVs), which are the pathogens that cause yearly flu epidemics and have caused periodic pandemics, such as the 1968 outbreak that killed an estimated one million people. His research aims to inform public health strategies on several important aspects of flu: seasonal flu; avian flu, which circulates among birds and has infected humans in the past; swine flu, which circulates among pigs and has infected humans in the past; and pandemic flu, which can arise from numerous sources and spread quickly because humans have little to no immunity to it.

Influenza, commonly known as "the flu", is an infectious disease caused by influenza viruses. Symptoms range from mild to severe and often include fever, runny nose, sore throat, muscle pain, headache, coughing, and fatigue. These symptoms begin from one to four days after exposure to the virus and last for about 2–8 days. Diarrhea and vomiting can occur, particularly in children. Influenza may progress to pneumonia, which can be caused by the virus or by a subsequent bacterial infection. Other complications of infection include acute respiratory distress syndrome, meningitis, encephalitis, and worsening of pre-existing health problems such as asthma and cardiovascular disease.
Johan Hultin was a Swedish-born American pathologist known for recovering tissues containing traces of the 1918 influenza virus that killed millions worldwide.

The pandemic H1N1/09 virus is a swine origin influenza A virus subtype H1N1 strain that was responsible for the 2009 swine flu pandemic. This strain is often called swine flu by the public media. For other names, see the Nomenclature section below.

A H5N1 vaccine is an influenza vaccine intended to provide immunization to influenza A virus subtype H5N1.