
A biological membrane, biomembrane or cell membrane is a selectively permeable membrane that separates the interior of a cell from the external environment or creates intracellular compartments by serving as a boundary between one part of the cell and another. Biological membranes, in the form of eukaryotic cell membranes, consist of a phospholipid bilayer with embedded, integral and peripheral proteins used in communication and transportation of chemicals and ions. The bulk of lipids in a cell membrane provides a fluid matrix for proteins to rotate and laterally diffuse for physiological functioning. Proteins are adapted to high membrane fluidity environment of the lipid bilayer with the presence of an annular lipid shell, consisting of lipid molecules bound tightly to the surface of integral membrane proteins. The cell membranes are different from the isolating tissues formed by layers of cells, such as mucous membranes, basement membranes, and serous membranes.

Phospholipids are a class of lipids whose molecule has a hydrophilic "head" containing a phosphate group and two hydrophobic "tails" derived from fatty acids, joined by an alcohol residue. Marine phospholipids typically have omega-3 fatty acids EPA and DHA integrated as part of the phospholipid molecule. The phosphate group can be modified with simple organic molecules such as choline, ethanolamine or serine.

ATPases (EC 3.6.1.3, Adenosine 5'-TriPhosphatase, adenylpyrophosphatase, ATP monophosphatase, triphosphatase, SV40 T-antigen, ATP hydrolase, complex V (mitochondrial electron transport), (Ca2+ + Mg2+)-ATPase, HCO3−-ATPase, adenosine triphosphatase) are a class of enzymes that catalyze the decomposition of ATP into ADP and a free phosphate ion or the inverse reaction. This dephosphorylation reaction releases energy, which the enzyme (in most cases) harnesses to drive other chemical reactions that would not otherwise occur. This process is widely used in all known forms of life.

The lipid bilayer is a thin polar membrane made of two layers of lipid molecules. These membranes are flat sheets that form a continuous barrier around all cells. The cell membranes of almost all organisms and many viruses are made of a lipid bilayer, as are the nuclear membrane surrounding the cell nucleus, and membranes of the membrane-bound organelles in the cell. The lipid bilayer is the barrier that keeps ions, proteins and other molecules where they are needed and prevents them from diffusing into areas where they should not be. Lipid bilayers are ideally suited to this role, even though they are only a few nanometers in width, because they are impermeable to most water-soluble (hydrophilic) molecules. Bilayers are particularly impermeable to ions, which allows cells to regulate salt concentrations and pH by transporting ions across their membranes using proteins called ion pumps.

Peripheral membrane proteins, or extrinsic membrane proteins, are membrane proteins that adhere only temporarily to the biological membrane with which they are associated. These proteins attach to integral membrane proteins, or penetrate the peripheral regions of the lipid bilayer. The regulatory protein subunits of many ion channels and transmembrane receptors, for example, may be defined as peripheral membrane proteins. In contrast to integral membrane proteins, peripheral membrane proteins tend to collect in the water-soluble component, or fraction, of all the proteins extracted during a protein purification procedure. Proteins with GPI anchors are an exception to this rule and can have purification properties similar to those of integral membrane proteins.

ATP synthase is an enzyme that catalyzes the formation of the energy storage molecule adenosine triphosphate (ATP) using adenosine diphosphate (ADP) and inorganic phosphate (Pi). ATP synthase is a molecular machine. The overall reaction catalyzed by ATP synthase is:

The fluid mosaic model explains various characteristics regarding the structure of functional cell membranes. According to this biological model, there is a lipid bilayer in which protein molecules are embedded. The phospholipid bilayer gives fluidity and elasticity to the membrane. Small amounts of carbohydrates are also found in the cell membrane. The biological model, which was devised by Seymour Jonathan Singer and Garth L. Nicolson in 1972, describes the cell membrane as a two-dimensional liquid that restricts the lateral diffusion of membrane components. Such domains are defined by the existence of regions within the membrane with special lipid and protein cocoon that promote the formation of lipid rafts or protein and glycoprotein complexes. Another way to define membrane domains is the association of the lipid membrane with the cytoskeleton filaments and the extracellular matrix through membrane proteins. The current model describes important features relevant to many cellular processes, including: cell-cell signaling, apoptosis, cell division, membrane budding, and cell fusion. The fluid mosaic model is the most acceptable model of the plasma membrane. In this definition of the cell membrane, its main function is to act as a barrier between the contents inside the cell and the extracellular environment.
A membrane transport protein is a membrane protein involved in the movement of ions, small molecules, and macromolecules, such as another protein, across a biological membrane. Transport proteins are integral transmembrane proteins; that is they exist permanently within and span the membrane across which they transport substances. The proteins may assist in the movement of substances by facilitated diffusion, active transport, osmosis, or reverse diffusion. The two main types of proteins involved in such transport are broadly categorized as either channels or carriers. Examples of channel/carrier proteins include the GLUT 1 uniporter, sodium channels, and potassium channels. The solute carriers and atypical SLCs are secondary active or facilitative transporters in humans. Collectively membrane transporters and channels are known as the transportome. Transportomes govern cellular influx and efflux of not only ions and nutrients but drugs as well.
In cellular biology, membrane transport refers to the collection of mechanisms that regulate the passage of solutes such as ions and small molecules through biological membranes, which are lipid bilayers that contain proteins embedded in them. The regulation of passage through the membrane is due to selective membrane permeability – a characteristic of biological membranes which allows them to separate substances of distinct chemical nature. In other words, they can be permeable to certain substances but not to others.
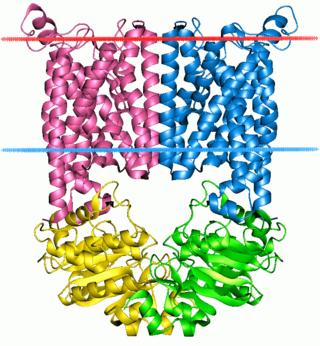
The ABC transporters, ATP synthase (ATP)-binding cassette transporters are a transport system superfamily that is one of the largest and possibly one of the oldest gene families. It is represented in all extant phyla, from prokaryotes to humans. ABC transporters belong to translocases.
Second messengers are intracellular signaling molecules released by the cell in response to exposure to extracellular signaling molecules—the first messengers. Second messengers trigger physiological changes at cellular level such as proliferation, differentiation, migration, survival, apoptosis and depolarization.
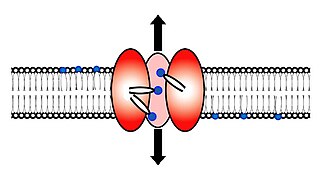
Scramblase is a protein responsible for the translocation of phospholipids between the two monolayers of a lipid bilayer of a cell membrane. In humans, phospholipid scramblases (PLSCRs) constitute a family of five homologous proteins that are named as hPLSCR1–hPLSCR5. Scramblases are members of the general family of transmembrane lipid transporters known as flippases. Scramblases are distinct from flippases and floppases. Scramblases, flippases, and floppases are three different types of enzymatic groups of phospholipid transportation enzymes. The inner-leaflet, facing the inside of the cell, contains negatively charged amino-phospholipids and phosphatidylethanolamine. The outer-leaflet, facing the outside environment, contains phosphatidylcholine and sphingomyelin. Scramblase is an enzyme, present in the cell membrane, that can transport (scramble) the negatively charged phospholipids from the inner-leaflet to the outer-leaflet, and vice versa.

ADP ribosylation factors (ARFs) are members of the ARF family of GTP-binding proteins of the Ras superfamily. ARF family proteins are ubiquitous in eukaryotic cells, and six highly conserved members of the family have been identified in mammalian cells. Although ARFs are soluble, they generally associate with membranes because of N-terminus myristoylation. They function as regulators of vesicular traffic and actin remodelling.
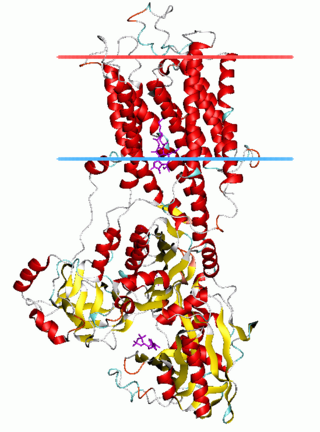
The P-type ATPases, also known as E1-E2 ATPases, are a large group of evolutionarily related ion and lipid pumps that are found in bacteria, archaea, and eukaryotes. P-type ATPases are α-helical bundle primary transporters named based upon their ability to catalyze auto- (or self-) phosphorylation (hence P) of a key conserved aspartate residue within the pump and their energy source, adenosine triphosphate (ATP). In addition, they all appear to interconvert between at least two different conformations, denoted by E1 and E2. P-type ATPases fall under the P-type ATPase (P-ATPase) Superfamily (TC# 3.A.3) which, as of early 2016, includes 20 different protein families.

In membrane biology, fusion is the process by which two initially distinct lipid bilayers merge their hydrophobic cores, resulting in one interconnected structure. If this fusion proceeds completely through both leaflets of both bilayers, an aqueous bridge is formed and the internal contents of the two structures can mix. Alternatively, if only one leaflet from each bilayer is involved in the fusion process, the bilayers are said to be hemifused. In hemifusion, the lipid constituents of the outer leaflet of the two bilayers can mix, but the inner leaflets remain distinct. The aqueous contents enclosed by each bilayer also remain separated.
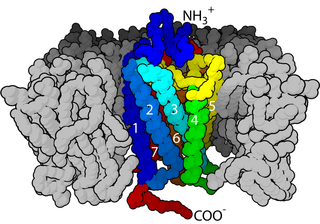
Cell surface receptors are receptors that are embedded in the plasma membrane of cells. They act in cell signaling by receiving extracellular molecules. They are specialized integral membrane proteins that allow communication between the cell and the extracellular space. The extracellular molecules may be hormones, neurotransmitters, cytokines, growth factors, cell adhesion molecules, or nutrients; they react with the receptor to induce changes in the metabolism and activity of a cell. In the process of signal transduction, ligand binding affects a cascading chemical change through the cell membrane.
The thiol-activated Cholesterol-dependent Cytolysin(CDC) family is a member of the MACPF superfamily. Cholesterol dependent cytolysins are a family of β-barrel pore-forming exotoxins that are secreted by gram-positive bacteria. CDCs are secreted as water-soluble monomers of 50-70 kDa, that when bound to the target cell, form a circular homo-oligomeric complex containing as many as 40 monomers. Through multiple conformational changes, the β-barrel transmembrane structure is formed and inserted into the target cell membrane. The presence of cholesterol in the target membrane is required for pore formation, though the presence of cholesterol is not required by all CDCs for binding. For example, intermedilysin secreted by Streptococcus intermedius will bind only to target membranes containing a specific protein receptor, independent of the presence of cholesterol, but cholesterol is required by intermedilysin for pore formation. While the lipid environment of cholesterol in the membrane can affect toxin binding, the exact molecular mechanism that cholesterol regulates the cytolytic activity of the CDC is not fully understood.
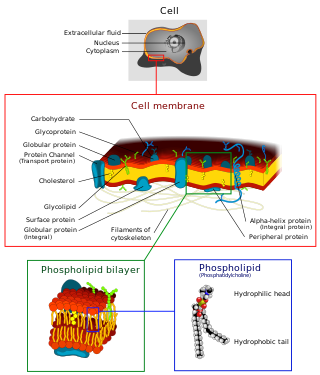
The cell membrane is a biological membrane that separates and protects the interior of a cell from the outside environment. The cell membrane consists of a lipid bilayer, made up of two layers of phospholipids with cholesterols interspersed between them, maintaining appropriate membrane fluidity at various temperatures. The membrane also contains membrane proteins, including integral proteins that span the membrane and serve as membrane transporters, and peripheral proteins that loosely attach to the outer (peripheral) side of the cell membrane, acting as enzymes to facilitate interaction with the cell's environment. Glycolipids embedded in the outer lipid layer serve a similar purpose. The cell membrane controls the movement of substances in and out of a cell, being selectively permeable to ions and organic molecules. In addition, cell membranes are involved in a variety of cellular processes such as cell adhesion, ion conductivity, and cell signalling and serve as the attachment surface for several extracellular structures, including the cell wall and the carbohydrate layer called the glycocalyx, as well as the intracellular network of protein fibers called the cytoskeleton. In the field of synthetic biology, cell membranes can be artificially reassembled.
The SecA protein is a cell membrane associated subunit of the bacterial Sec or Type II secretory pathway, a system which is responsible for the secretion of proteins through the cell membrane. Within this system the SecA ATPase forms a translocase complex with the SecYEG channel, thereby driving the movement of the protein substrate across the membrane.

The CDP-choline pathway, first identified by Eugene P. Kennedy in 1956, is the predominant mechanism by which mammalian cells synthesize phosphatidylcholine (PC) for incorporation into membranes or lipid-derived signalling molecules. The CDP-choline pathway represents one half of what is known as the Kennedy pathway. The other half is the CDP-ethanolamine pathway which is responsible for the biosynthesis of the phospholipid phosphatidylethanolamine (PE).



















