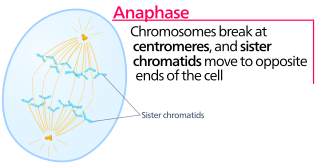
Anaphase is the stage of mitosis after the process of metaphase, when replicated chromosomes are split and the newly-copied chromosomes are moved to opposite poles of the cell. Chromosomes also reach their overall maximum condensation in late anaphase, to help chromosome segregation and the re-formation of the nucleus.

In cell biology, the spindle apparatus is the cytoskeletal structure of eukaryotic cells that forms during cell division to separate sister chromatids between daughter cells. It is referred to as the mitotic spindle during mitosis, a process that produces genetically identical daughter cells, or the meiotic spindle during meiosis, a process that produces gametes with half the number of chromosomes of the parent cell.
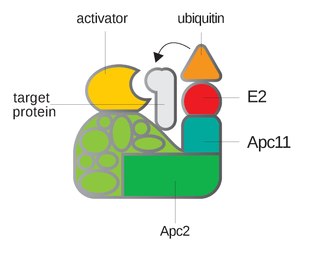
Anaphase-promoting complex is an E3 ubiquitin ligase that marks target cell cycle proteins for degradation by the 26S proteasome. The APC/C is a large complex of 11–13 subunit proteins, including a cullin (Apc2) and RING (Apc11) subunit much like SCF. Other parts of the APC/C have unknown functions but are highly conserved.

Metaphase is a stage of mitosis in the eukaryotic cell cycle in which chromosomes are at their second-most condensed and coiled stage. These chromosomes, carrying genetic information, align in the equator of the cell between the spindle poles at the metaphase plate, before being separated into each of the two daughter nuclei. This alignment marks the beginning of metaphase. Metaphase accounts for approximately 4% of the cell cycle's duration.

The spindle checkpoint, also known as the metaphase-to-anaphase transition, the spindle assembly checkpoint (SAC), the metaphase checkpoint, or the mitotic checkpoint, is a cell cycle checkpoint during metaphase of mitosis or meiosis that prevents the separation of the duplicated chromosomes (anaphase) until each chromosome is properly attached to the spindle. To achieve proper segregation, the two kinetochores on the sister chromatids must be attached to opposite spindle poles. Only this pattern of attachment will ensure that each daughter cell receives one copy of the chromosome. The defining biochemical feature of this checkpoint is the stimulation of the anaphase-promoting complex by M-phase cyclin-CDK complexes, which in turn causes the proteolytic destruction of cyclins and proteins that hold the sister chromatids together.

A kinetochore is a disc-shaped protein structure associated with duplicated chromatids in eukaryotic cells where the spindle fibers attach during cell division to pull sister chromatids apart. The kinetochore assembles on the centromere and links the chromosome to microtubule polymers from the mitotic spindle during mitosis and meiosis. The term kinetochore was first used in a footnote in a 1934 Cytology book by Lester W. Sharp and commonly accepted in 1936. Sharp's footnote reads: "The convenient term kinetochore has been suggested to the author by J. A. Moore", likely referring to John Alexander Moore who had joined Columbia University as a freshman in 1932.

Separase, also known as separin, is a cysteine protease responsible for triggering anaphase by hydrolysing cohesin, which is the protein responsible for binding sister chromatids during the early stage of anaphase. In humans, separin is encoded by the ESPL1 gene.
Securin is a protein involved in control of the metaphase-anaphase transition and anaphase onset. Following bi-orientation of chromosome pairs and inactivation of the spindle checkpoint system, the underlying regulatory system, which includes securin, produces an abrupt stimulus that induces highly synchronous chromosome separation in anaphase.

Cohesin is a protein complex that mediates sister chromatid cohesion, homologous recombination, and DNA looping. Cohesin is formed of SMC3, SMC1, SCC1 and SCC3. Cohesin holds sister chromatids together after DNA replication until anaphase when removal of cohesin leads to separation of sister chromatids. The complex forms a ring-like structure and it is believed that sister chromatids are held together by entrapment inside the cohesin ring. Cohesin is a member of the SMC family of protein complexes which includes Condensin, MukBEF and SMC-ScpAB.

The cell division cycle protein 20 homolog is an essential regulator of cell division that is encoded by the CDC20 gene in humans. To the best of current knowledge its most important function is to activate the anaphase promoting complex (APC/C), a large 11-13 subunit complex that initiates chromatid separation and entrance into anaphase. The APC/CCdc20 protein complex has two main downstream targets. Firstly, it targets securin for destruction, enabling the eventual destruction of cohesin and thus sister chromatid separation. It also targets S and M-phase (S/M) cyclins for destruction, which inactivates S/M cyclin-dependent kinases (Cdks) and allows the cell to exit from mitosis. A closely related protein, Cdc20homologue-1 (Cdh1) plays a complementary role in the cell cycle.

Mitotic checkpoint serine/threonine-protein kinase BUB1 also known as BUB1 is an enzyme that in humans is encoded by the BUB1 gene.
Mitotic checkpoint serine/threonine-protein kinase BUB1 beta is an enzyme that in humans is encoded by the BUB1B gene. Also known as BubR1, this protein is recognized for its mitotic roles in the spindle assembly checkpoint (SAC) and kinetochore-microtubule interactions that facilitate chromosome migration and alignment. BubR1 promotes mitotic fidelity and protects against aneuploidy by ensuring proper chromosome segregation between daughter cells. BubR1 is proposed to prevent tumorigenesis.

Mitotic spindle assembly checkpoint protein MAD2A is a protein that in humans is encoded by the MAD2L1 gene.

Cell division cycle protein 27 homolog is a protein that in humans is encoded by the CDC27 gene.

Cell division cycle protein 16 homolog is a protein that in humans is encoded by the CDC16 gene.

Mitotic checkpoint protein BUB3 is a protein that in humans is encoded by the BUB3 gene.

TRIP13 is a mammalian gene that encodes the thyroid receptor-interacting protein 13. In budding yeast, the analog for TRIP13 is PCH2. TRIP13 is a member of the AAA+ ATPase family, a family known for mechanical forces derived from ATP hydrolase reactions. The TRIP13 gene has been shown to interact with a variety of proteins and implicated in a few diseases, notably interacting with the ligand binding domain of thyroid hormone receptors, and may play a role in early-stage non-small cell lung cancer. However, recent evidence implicates TRIP13 in various cell cycle phases, including meiosis G2/Prophase and during the Spindle Assembly checkpoint (SAC). Evidence shows regulation to occur through the HORMA domains, including Hop1, Rev7, and Mad2. Of note, Mad2's involvement in the SAC is shown to be affected by TRIP13 Due to TRIP13's role in cell cycle arrest and progression, it may present opportunity as a therapeutic candidate for cancers.
A series of biochemical switches control transitions between and within the various phases of the cell cycle. The cell cycle is a series of complex, ordered, sequential events that control how a single cell divides into two cells, and involves several different phases. The phases include the G1 and G2 phases, DNA replication or S phase, and the actual process of cell division, mitosis or M phase. During the M phase, the chromosomes separate and cytokinesis occurs.

Mad1 is a non-essential protein which in yeast has a function in the spindle assembly checkpoint (SAC). This checkpoint monitors chromosome attachment to spindle microtubules and prevents cells from starting anaphase until the spindle is built up. The name Mad refers to the observation that mutant cells are mitotic arrest deficient (MAD) during microtubule depolymerization. Mad1 recruits the anaphase inhibitor Mad2 to unattached kinetochores and is essential for Mad2-Cdc20 complex formation in vivo but not in vitro. In vivo, Mad1 acts as a competitive inhibitor of the Mad2-Cdc20 complex. Mad1 is phosphorylated by Mps1 which then leads together with other activities to the formation of the mitotic checkpoint complex (MCC). Thereby it inhibits the activity of the anaphase-promoting complex/cyclosome (APC/C). Homologues of Mad1 are conserved in eukaryotes from yeast to mammals.
Mitotic exit is an important transition point that signifies the end of mitosis and the onset of new G1 phase for a cell, and the cell needs to rely on specific control mechanisms to ensure that once it exits mitosis, it never returns to mitosis until it has gone through G1, S, and G2 phases and passed all the necessary checkpoints. Many factors including cyclins, cyclin-dependent kinases (CDKs), ubiquitin ligases, inhibitors of cyclin-dependent kinases, and reversible phosphorylations regulate mitotic exit to ensure that cell cycle events occur in correct order with fewest errors. The end of mitosis is characterized by spindle breakdown, shortened kinetochore microtubules, and pronounced outgrowth of astral (non-kinetochore) microtubules. For a normal eukaryotic cell, mitotic exit is irreversible.


















