
Coronaviruses are a group of related RNA viruses that cause diseases in mammals and birds. In humans and birds, they cause respiratory tract infections that can range from mild to lethal. Mild illnesses in humans include some cases of the common cold, while more lethal varieties can cause SARS, MERS and COVID-19, which is causing the ongoing pandemic. In cows and pigs they cause diarrhea, while in mice they cause hepatitis and encephalomyelitis.

Severe acute respiratory syndrome–related coronavirus is a species of virus consisting of many known strains phylogenetically related to severe acute respiratory syndrome coronavirus 1 (SARS-CoV-1) that have been shown to possess the capability to infect humans, bats, and certain other mammals. These enveloped, positive-sense single-stranded RNA viruses enter host cells by binding to the angiotensin-converting enzyme 2 (ACE2) receptor. The SARSr-CoV species is a member of the genus Betacoronavirus and of the subgenus Sarbecovirus.
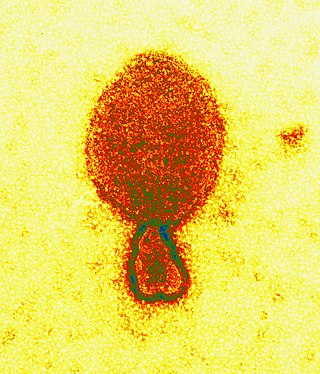
Henipavirus is a genus of negative-strand RNA viruses in the family Paramyxoviridae, order Mononegavirales containing six established species, and numerous others still under study. Henipaviruses are naturally harboured by several species of small mammals, notably pteropid fruit bats, microbats of several species, and shrews. Henipaviruses are characterised by long genomes and a wide host range. Their recent emergence as zoonotic pathogens capable of causing illness and death in domestic animals and humans is a cause of concern.
An NTP binding site is a type of binding site found in nucleoside monophosphate (NMP) kinases, N can be adenosine or guanosine. A P-loop is one of the structural motifs common for nucleoside triphosphate (NTP) binding sites, it interacts with the bound nucleotide's phosphoryl groups. For the binding site to be able to bind a nucleotide, the nucleotide must be complex bound to Mg2+ or Mn2+. Nucleotide binding will cause conformational changes in the protein because the P-loop will bend.

Nidovirales is an order of enveloped, positive-strand RNA viruses which infect vertebrates and invertebrates. Host organisms include mammals, birds, reptiles, amphibians, fish, arthropods, molluscs, and helminths. The order includes the families Coronaviridae, Arteriviridae, Roniviridae, and Mesoniviridae.

RNA-dependent RNA polymerase (RdRp) or RNA replicase is an enzyme that catalyzes the replication of RNA from an RNA template. Specifically, it catalyzes synthesis of the RNA strand complementary to a given RNA template. This is in contrast to typical DNA-dependent RNA polymerases, which all organisms use to catalyze the transcription of RNA from a DNA template.

The Coronavirus packaging signal is a conserved cis-regulatory element found in Betacoronavirus. It has an important role in regulating the packaging of the viral genome into the capsid. As part of the viral life cycle, within the infected cell, the viral genome becomes associated with viral proteins and assembles into new infective progeny viruses. This process is called packaging and is vital for viral replication.
Putative transmembrane domain more commonly known as Non-structural Protein 6 (NSP6) is one of the two non-structural proteins that gene 11 in rotavirus encodes for alongside NSP5. NSP6 is composed of six transmembrane domains and a C terminal tail. In contrast to the other rotavirus non-structural proteins, NSP6 was found to have a high rate of turnover, being completely degraded within 2 hours of synthesis. NSP6 was found to be a sequence-independent nucleic acid binding protein, with similar affinities for ssRNA and dsRNA

Hepatitis B virus DNA polymerase is a hepatitis B viral protein. It is a DNA polymerase that can use either DNA or RNA templates and a ribonuclease H that cuts RNA in the duplex. Both functions are supplied by the reverse transcriptase (RT) domain.
The term proofreading is used in genetics to refer to the error-correcting processes, first proposed by John Hopfield and Jacques Ninio, involved in DNA replication, immune system specificity, enzyme-substrate recognition among many other processes that require enhanced specificity. The proofreading mechanisms of Hopfield and Ninio are non-equilibrium active processes that consume ATP to enhance specificity of various biochemical reactions.

Deepak Thankappan Nair is an Indian Structural Biologist and a scientist at Regional Centre for Biotechnology. He is known for his studies on DNA and RNA polymerases. Deepak was a Ramanujan fellow of the Science and Engineering Research Board (2008–2013) and a recipient of the National BioScience Award for Career Development. The Council of Scientific and Industrial Research, the apex agency of the Government of India for scientific research, awarded him the Shanti Swarup Bhatnagar Prize for Science and Technology, one of the highest Indian science awards, for his contributions to biological sciences in 2017.

The first step of transcription for some negative, single-stranded RNA viruses is cap snatching, in which the first 10 to 20 residues of a host cell RNA are removed (snatched) and used as the 5′ cap and primer to initiate the synthesis of the nascent viral mRNA. The viral RNA-dependent RNA polymerase (RdRp) can then proceed to replicate the negative-sense genome from the positive-sense template. Cap-snatching also explains why some viral mRNA have 5’ terminal extensions of 10-20 nucleotides that are not encoded for in the genome. Examples of viruses that engage in cap-snatching include influenza viruses (Orthomyxoviridae), Lassa virus (Arenaviridae), hantaan virus (Hantaviridae) and rift valley fever virus (Phenuiviridae). Most viruses snatch 15-20 nucleotides except for the families Arenaviridae and Nairoviridae and the genus Thogotovirus (Orthomyxoviridae) which use a shorter strand.
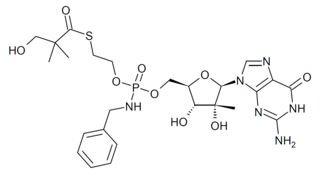
IDX-184 is an antiviral drug which was developed as a treatment for hepatitis C, acting as a NS5B RNA polymerase inhibitor. While it showed reasonable effectiveness in early clinical trials it did not progress past Phase IIb. However research using this drug has continued as it shows potentially useful activity against other emerging viral diseases such as Zika virus, and coronaviruses including MERS, and SARS-CoV-2.
Coronavirus genomes are positive-sense single-stranded RNA molecules with an untranslated region (UTR) at the 5′ end which is called the 5′ UTR. The 5′ UTR is responsible for important biological functions, such as viral replication, transcription and packaging. The 5′ UTR has a conserved RNA secondary structure but different Coronavirus genera have different structural features described below.

The envelope (E) protein is the smallest and least well-characterized of the four major structural proteins found in coronavirus virions. It is an integral membrane protein less than 110 amino acid residues long; in SARS-CoV-2, the causative agent of Covid-19, the E protein is 75 residues long. Although it is not necessarily essential for viral replication, absence of the E protein may produce abnormally assembled viral capsids or reduced replication. E is a multifunctional protein and, in addition to its role as a structural protein in the viral capsid, it is thought to be involved in viral assembly, likely functions as a viroporin, and is involved in viral pathogenesis.

The nucleocapsid (N) protein is a protein that packages the positive-sense RNA genome of coronaviruses to form ribonucleoprotein structures enclosed within the viral capsid. The N protein is the most highly expressed of the four major coronavirus structural proteins. In addition to its interactions with RNA, N forms protein-protein interactions with the coronavirus membrane protein (M) during the process of viral assembly. N also has additional functions in manipulating the cell cycle of the host cell. The N protein is highly immunogenic and antibodies to N are found in patients recovered from SARS and COVID-19.
ORF7b is a gene found in coronaviruses of the genus Betacoronavirus, which expresses the accessory protein Betacoronavirus NS7b protein. It is a short, highly hydrophobic transmembrane protein of unknown function.

ORF8 is a gene that encodes a viral accessory protein, Betacoronavirus NS8 protein, in coronaviruses of the subgenus Sarbecovirus. It is one of the least well conserved and most variable parts of the genome. In some viruses, a deletion splits the region into two smaller open reading frames, called ORF8a and ORF8b - a feature present in many SARS-CoV viral isolates from later in the SARS epidemic, as well as in some bat coronaviruses. For this reason the full-length gene and its protein are sometimes called ORF8ab. The full-length gene, exemplified in SARS-CoV-2, encodes a protein with an immunoglobulin domain of unknown function, possibly involving interactions with the host immune system. It is similar in structure to the ORF7a protein, suggesting it may have originated through gene duplication.
ORF1ab refers collectively to two open reading frames (ORFs), ORF1a and ORF1b, that are conserved in the genomes of nidoviruses, a group of viruses that includes coronaviruses. The genes express large polyproteins that undergo proteolysis to form several nonstructural proteins with various functions in the viral life cycle, including proteases and the components of the replicase-transcriptase complex (RTC). Together the two ORFs are sometimes referred to as the replicase gene. They are related by a programmed ribosomal frameshift that allows the ribosome to continue translating past the stop codon at the end of ORF1a, in a -1 reading frame. The resulting polyproteins are known as pp1a and pp1ab.
Planarian secretory cell nidovirus (PSCNV) is a virus of the species Planidovirus 1, a nidovirus notable for its extremely large genome. At 41.1 kilobases, it is the largest known genome of an RNA virus. It was discovered by inspecting the transcriptomes of the planarian flatworm Schmidtea mediterranea and is the first known RNA virus infecting planarians. It was first described in 2018.











