
β-Galactosidase, is a glycoside hydrolase enzyme that catalyzes hydrolysis of terminal non-reducing β-D-galactose residues in β-D-galactosides.

A cloning vector is a small piece of DNA that can be stably maintained in an organism, and into which a foreign DNA fragment can be inserted for cloning purposes. The cloning vector may be DNA taken from a virus, the cell of a higher organism, or it may be the plasmid of a bacterium. The vector contains features that allow for the convenient insertion of a DNA fragment into the vector or its removal from the vector, for example through the presence of restriction sites. The vector and the foreign DNA may be treated with a restriction enzyme that cuts the DNA, and DNA fragments thus generated contain either blunt ends or overhangs known as sticky ends, and vector DNA and foreign DNA with compatible ends can then be joined by molecular ligation. After a DNA fragment has been cloned into a cloning vector, it may be further subcloned into another vector designed for more specific use.

The lactose operon is an operon required for the transport and metabolism of lactose in E. coli and many other enteric bacteria. Although glucose is the preferred carbon source for most enteric bacteria, the lac operon allows for the effective digestion of lactose when glucose is not available through the activity of beta-galactosidase. Gene regulation of the lac operon was the first genetic regulatory mechanism to be understood clearly, so it has become a foremost example of prokaryotic gene regulation. It is often discussed in introductory molecular and cellular biology classes for this reason. This lactose metabolism system was used by François Jacob and Jacques Monod to determine how a biological cell knows which enzyme to synthesize. Their work on the lac operon won them the Nobel Prize in Physiology in 1965.
A cosmid is a type of hybrid plasmid that contains a Lambda phage cos sequence. They are often used as a cloning vector in genetic engineering. Cosmids can be used to build genomic libraries. They were first described by Collins and Hohn in 1978. Cosmids can contain 37 to 52 kb of DNA, limits based on the normal bacteriophage packaging size. They can replicate as plasmids if they have a suitable origin of replication (ori): for example SV40 ori in mammalian cells, ColE1 ori for double-stranded DNA replication, or f1 ori for single-stranded DNA replication in prokaryotes. They frequently also contain a gene for selection such as antibiotic resistance, so that the transformed cells can be identified by plating on a medium containing the antibiotic. Those cells which did not take up the cosmid would be unable to grow.
A DNA construct is an artificially-designed segment of DNA borne on a vector that can be used to incorporate genetic material into a target tissue or cell. A DNA construct contains a DNA insert, called a transgene, delivered via a transformation vector which allows the insert sequence to be replicated and/or expressed in the target cell. This gene can be cloned from a naturally occurring gene, or synthetically constructed. The vector can be delivered using physical, chemical or viral methods. Typically, the vectors used in DNA constructs contain an origin of replication, a multiple cloning site, and a selectable marker. Certain vectors can carry additional regulatory elements based on the expression system involved.

Isopropyl β-d-1-thiogalactopyranoside (IPTG) is a molecular biology reagent. This compound is a molecular mimic of allolactose, a lactose metabolite that triggers transcription of the lac operon, and it is therefore used to induce protein expression where the gene is under the control of the lac operator.

A multiple cloning site (MCS), also called a polylinker, is a short segment of DNA which contains many restriction sites - a standard feature of engineered plasmids. Restriction sites within an MCS are typically unique, occurring only once within a given plasmid. The purpose of an MCS in a plasmid is to allow a piece of DNA to be inserted into that region.

In molecular biology, subcloning is a technique used to move a particular DNA sequence from a parent vector to a destination vector.
In biology, a marker gene may have several meanings. In nuclear biology and molecular biology, a marker gene is a gene used to determine if a nucleic acid sequence has been successfully inserted into an organism's DNA. In particular, there are two sub-types of these marker genes: a selectable marker and a marker for screening. In metagenomics and phylogenetics, a marker gene is an orthologous gene group which can be used to delineate between taxonomic lineages.
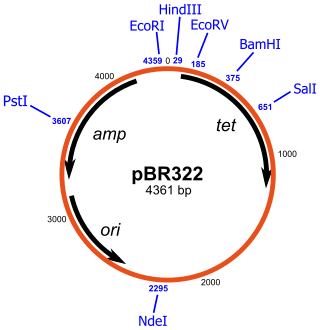
pBR322 is a plasmid and was one of the first widely used E. coli cloning vectors. Created in 1977 in the laboratory of Herbert Boyer at the University of California, San Francisco, it was named after Francisco Bolivar Zapata, the postdoctoral researcher and Raymond L. Rodriguez. The p stands for "plasmid," and BR for "Bolivar" and "Rodriguez."

X-gal is an organic compound consisting of galactose linked to a substituted indole. The compound was synthesized by Jerome Horwitz and collaborators in 1964. The formal chemical name is often shortened to less accurate but also less cumbersome phrases such as bromochloroindoxyl galactoside. The X from indoxyl may be the source of the X in the X-gal contraction. X-gal is often used in molecular biology to test for the presence of an enzyme, β-galactosidase, in the place of its usual target, a β-galactoside. It is also used to detect activity of this enzyme in histochemistry and bacteriology. X-gal is one of many indoxyl glycosides and esters that yield insoluble blue compounds similar to indigo dye as a result of enzyme-catalyzed hydrolysis.
A genomic library is a collection of overlapping DNA fragments that together make up the total genomic DNA of a single organism. The DNA is stored in a population of identical vectors, each containing a different insert of DNA. In order to construct a genomic library, the organism's DNA is extracted from cells and then digested with a restriction enzyme to cut the DNA into fragments of a specific size. The fragments are then inserted into the vector using DNA ligase. Next, the vector DNA can be taken up by a host organism - commonly a population of Escherichia coli or yeast - with each cell containing only one vector molecule. Using a host cell to carry the vector allows for easy amplification and retrieval of specific clones from the library for analysis.
Galactosidases are enzymes that catalyze the hydrolysis of galactosides into monosaccharides.
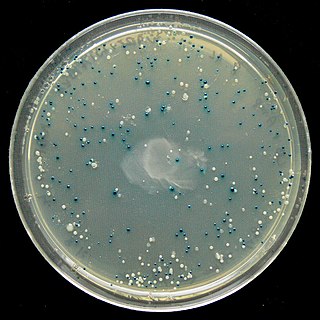
The blue–white screen is a screening technique that allows for the rapid and convenient detection of recombinant bacteria in vector-based molecular cloning experiments. This method of screening is usually performed using a suitable bacterial strain, but other organisms such as yeast may also be used. DNA of transformation is ligated into a vector. The vector is then inserted into a competent host cell viable for transformation, which are then grown in the presence of X-gal. Cells transformed with vectors containing recombinant DNA will produce white colonies; cells transformed with non-recombinant plasmids grow into blue colonies.
The pGreen plasmids are vectors for plant transformation. They were first described in 2000 as components of a novel T-DNA binary system. The supporting web page provides supplementary information and ongoing support to researchers to request their plasmid resources. As these plasmids have been taken up by the research community, the plasmids have been developed, expanding the resources available to the community.
In molecular cloning, a vector is any particle used as a vehicle to artificially carry a foreign nucleic sequence – usually DNA – into another cell, where it can be replicated and/or expressed. A vector containing foreign DNA is termed recombinant DNA. The four major types of vectors are plasmids, viral vectors, cosmids, and artificial chromosomes. Of these, the most commonly used vectors are plasmids. Common to all engineered vectors are an origin of replication, a multicloning site, and a selectable marker.
The Gateway cloning method, invented and commercialized by Invitrogen since the late 1990s, is the cloning method of the integration and excision recombination reactions that take place when bacteriophage lambda infects bacteria. This technology provides a fast and highly efficient way to transport DNA sequences into multi-vector systems for functional analysis and protein expression using Gateway att sites, and two proprietary enzyme mixes called BP Clonase and LR Clonase. In vivo, these recombination reactions are facilitated by the recombination of attachment sites from the lambda/phage chromosome (attP) and the bacteria (attB). As a result of recombination between the attP and attB sites, the phage integrates into the bacterial genome flanked by two new recombination sites. The removal of the phage from the bacterial chromosome and the regeneration of attP and attB sites can both result from the attL and attR sites recombining under specific circumstances.

Molecular cloning is a set of experimental methods in molecular biology that are used to assemble recombinant DNA molecules and to direct their replication within host organisms. The use of the word cloning refers to the fact that the method involves the replication of one molecule to produce a population of cells with identical DNA molecules. Molecular cloning generally uses DNA sequences from two different organisms: the species that is the source of the DNA to be cloned, and the species that will serve as the living host for replication of the recombinant DNA. Molecular cloning methods are central to many contemporary areas of modern biology and medicine.
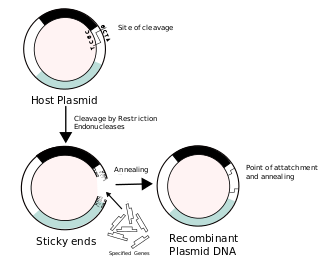
Recombinant DNA (rDNA), or molecular cloning, is the process by which a single gene, or segment of DNA, is isolated and amplified. Recombinant DNA is also known as in vitro recombination. A cloning vector is a DNA molecule that carries foreign DNA into a host cell, where it replicates, producing many copies of itself along with the foreign DNA. There are many types of cloning vectors such as plasmids and phages. In order to carry out recombination between vector and the foreign DNA, it is necessary the vector and DNA to be cloned by digestion, ligase the foreign DNA into the vector with the enzyme DNA ligase. And DNA is inserted by introducing the DNA into bacteria cells by transformation.

Golden Gate Cloning or Golden Gate assembly is a molecular cloning method that allows a researcher to simultaneously and directionally assemble multiple DNA fragments into a single piece using Type IIS restriction enzymes and T4 DNA ligase. This assembly is performed in vitro. Most commonly used Type IIS enzymes include BsaI, BsmBI, and BbsI.













