Related Research Articles

Histone deacetylases (EC 3.5.1.98, HDAC) are a class of enzymes that remove acetyl groups (O=C-CH3) from an ε-N-acetyl lysine amino acid on both histone and non-histone proteins. HDACs allow histones to wrap the DNA more tightly. This is important because DNA is wrapped around histones, and DNA expression is regulated by acetylation and de-acetylation. HDAC's action is opposite to that of histone acetyltransferase. HDAC proteins are now also called lysine deacetylases (KDAC), to describe their function rather than their target, which also includes non-histone proteins.

Trichostatin A (TSA) is an organic compound that serves as an antifungal antibiotic and selectively inhibits the class I and II mammalian histone deacetylase (HDAC) families of enzymes, but not class III HDACs. However, there are recent reports of the interactions of this molecule with Sirt 6 protein. TSA inhibits the eukaryotic cell cycle during the beginning of the growth stage. TSA can be used to alter gene expression by interfering with the removal of acetyl groups from histones and therefore altering the ability of DNA transcription factors to access the DNA molecules inside chromatin. It is a member of a larger class of histone deacetylase inhibitors that have a broad spectrum of epigenetic activities. Thus, TSA has some potential as an anti-cancer drug. One suggested mechanism is that TSA promotes the expression of apoptosis-related genes, leading to cancerous cells surviving at lower rates, thus slowing the progression of cancer. Other mechanisms may include the activity of HDIs to induce cell differentiation, thus acting to "mature" some of the de-differentiated cells found in tumors. HDIs have multiple effects on non-histone effector molecules, so the anti-cancer mechanisms are truly not understood at this time.
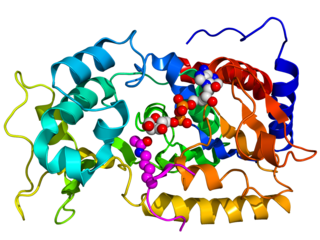
Sirtuins are a family of signaling proteins involved in metabolic regulation. They are ancient in animal evolution and appear to possess a highly conserved structure throughout all kingdoms of life. Chemically, sirtuins are a class of proteins that possess either mono-ADP-ribosyltransferase or deacylase activity, including deacetylase, desuccinylase, demalonylase, demyristoylase and depalmitoylase activity. The name Sir2 comes from the yeast gene 'silent mating-type information regulation 2', the gene responsible for cellular regulation in yeast.

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation.

Histone deacetylase 1 (HDAC1) is an enzyme that in humans is encoded by the HDAC1 gene.
Histone deacetylase inhibitors are chemical compounds that inhibit histone deacetylases.

Histone deacetylase 2 (HDAC2) is an enzyme that in humans is encoded by the HDAC2 gene. It belongs to the histone deacetylase class of enzymes responsible for the removal of acetyl groups from lysine residues at the N-terminal region of the core histones. As such, it plays an important role in gene expression by facilitating the formation of transcription repressor complexes and for this reason is often considered an important target for cancer therapy.

Histone deacetylase 3 is an enzyme encoded by the HDAC3 gene in both humans and mice.

Histone deacetylase 4, also known as HDAC4, is a protein that in humans is encoded by the HDAC4 gene.

Histone deacetylase 6 is an enzyme that in humans is encoded by the HDAC6 gene. HDAC6 has emerged as a highly promising candidate to selectively inhibit as a therapeutic strategy to combat several types of cancer and neurodegenerative disorders.

Sirtuin 1, also known as NAD-dependent deacetylase sirtuin-1, is a protein that in humans is encoded by the SIRT1 gene.

Histone deacetylase 5 is an enzyme that in humans is encoded by the HDAC5 gene.

Histone deacetylase 9 is an enzyme that in humans is encoded by the HDAC9 gene.

NAD-dependent deacetylase sirtuin 2 is an enzyme that in humans is encoded by the SIRT2 gene. SIRT2 is an NAD+ -dependent deacetylase. Studies of this protein have often been divergent, highlighting the dependence of pleiotropic effects of SIRT2 on cellular context. The natural polyphenol resveratrol is known to exert opposite actions on neural cells according to their normal or cancerous status. Similar to other sirtuin family members, SIRT2 displays a ubiquitous distribution. SIRT2 is expressed in a wide range of tissues and organs and has been detected particularly in metabolically relevant tissues, including the brain, muscle, liver, testes, pancreas, kidney, and adipose tissue of mice. Of note, SIRT2 expression is much higher in the brain than all other organs studied, particularly in the cortex, striatum, hippocampus, and spinal cord.
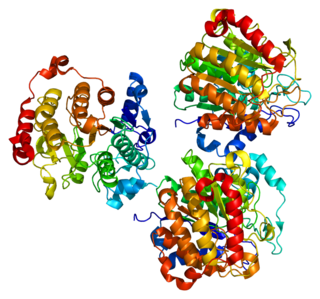
Histone deacetylase 7 is an enzyme that in humans is encoded by the HDAC7 gene.

NAD-dependent deacetylase sirtuin-3, mitochondrial also known as SIRT3 is a protein that in humans is encoded by the SIRT3 gene [sirtuin 3 ]. SIRT3 is member of the mammalian sirtuin family of proteins, which are homologs to the yeast Sir2 protein. SIRT3 exhibits NAD+-dependent deacetylase activity.

Histone deacetylase 11 is a 39kDa histone deacetylase enzyme that in humans is encoded by the HDAC11 gene on chromosome 3 in humans and chromosome 6 in mice.
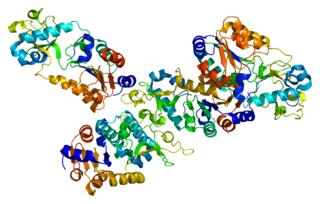
Sirtuin 5 , also known as SIRT5 is a protein which in humans in encoded by the SIRT5 gene and in other species by the orthologous Sirt5 gene.

Sirtuin 6 is a stress responsive protein deacetylase and mono-ADP ribosyltransferase enzyme encoded by the SIRT6 gene. In laboratory research, SIRT6 appears to function in multiple molecular pathways related to aging, including DNA repair, telomere maintenance, glycolysis and inflammation. SIRT6 is member of the mammalian sirtuin family of proteins, which are homologs to the yeast Sir2 protein.
Cocaine addiction is the compulsive use of cocaine despite adverse consequences. It arises through epigenetic modification and transcriptional regulation of genes in the nucleus accumbens.
References
- ↑ "Protein Deacetylase - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 2023-03-16.
- ↑ Gupta, Rohan; Ambasta, Rashmi K.; Kumar, Pravir (2022-01-01). "Multifaced role of protein deacetylase sirtuins in neurodegenerative disease". Neuroscience & Biobehavioral Reviews. 132: 976–997. doi:10.1016/j.neubiorev.2021.10.047. ISSN 0149-7634. PMID 34742724. S2CID 241905442.