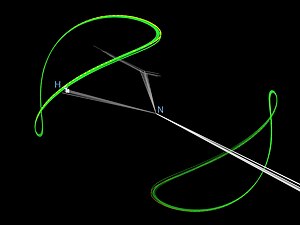
The nuclear Overhauser effect (NOE) is the transfer of nuclear spin polarization from one population of spin-active nuclei to another via cross-relaxation. A phenomenological definition of the NOE in nuclear magnetic resonance spectroscopy (NMR) is the change in the integrated intensity of one NMR resonance that occurs when another is saturated by irradiation with an RF field. The change in resonance intensity of a nucleus is a consequence of the nucleus being close in space to those directly affected by the RF perturbation.
In nuclear magnetic resonance (NMR) spectroscopy, the chemical shift is the resonant frequency of an atomic nucleus relative to a standard in a magnetic field. Often the position and number of chemical shifts are diagnostic of the structure of a molecule. Chemical shifts are also used to describe signals in other forms of spectroscopy such as photoemission spectroscopy.

Nuclear magnetic resonance spectroscopy, most commonly known as NMR spectroscopy or magnetic resonance spectroscopy (MRS), is a spectroscopic technique based on re-orientation of atomic nuclei with non-zero nuclear spins in an external magnetic field. This re-orientation occurs with absorption of electromagnetic radiation in the radio frequency region from roughly 4 to 900 MHz, which depends on the isotopic nature of the nucleus and increased proportionally to the strength of the external magnetic field. Notably, the resonance frequency of each NMR-active nucleus depends on its chemical environment. As a result, NMR spectra provide information about individual functional groups present in the sample, as well as about connections between nearby nuclei in the same molecule. As the NMR spectra are unique or highly characteristic to individual compounds and functional groups, NMR spectroscopy is one of the most important methods to identify molecular structures, particularly of organic compounds.
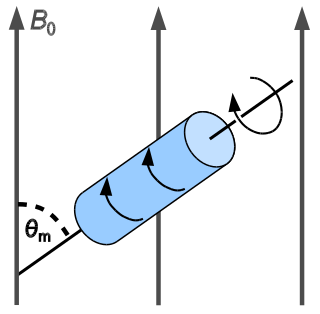
In solid-state NMR spectroscopy, magic-angle spinning (MAS) is a technique routinely used to produce better resolution NMR spectra. MAS NMR consists in spinning the sample at the magic angle θm with respect to the direction of the magnetic field.

Electron paramagnetic resonance (EPR) or electron spin resonance (ESR) spectroscopy is a method for studying materials that have unpaired electrons. The basic concepts of EPR are analogous to those of nuclear magnetic resonance (NMR), but the spins excited are those of the electrons instead of the atomic nuclei. EPR spectroscopy is particularly useful for studying metal complexes and organic radicals. EPR was first observed in Kazan State University by Soviet physicist Yevgeny Zavoisky in 1944, and was developed independently at the same time by Brebis Bleaney at the University of Oxford.

Solid-state NMR (ssNMR) spectroscopy is a technique for characterizing atomic level structure in solid materials e.g. powders, single crystals and amorphous samples and tissues using nuclear magnetic resonance (NMR) spectroscopy. The anisotropic part of many spin interactions are present in solid-state NMR, unlike in solution-state NMR where rapid tumbling motion averages out many of the spin interactions. As a result, solid-state NMR spectra are characterised by larger linewidths than in solution state NMR, which can be utilized to give quantitative information on the molecular structure, conformation and dynamics of the material. Solid-state NMR is often combined with magic angle spinning to remove anisotropic interactions and improve the resolution as well as the sensitivity of the technique.
The magic angle is a precisely defined angle, the value of which is approximately 54.7356°. The magic angle is a root of a second-order Legendre polynomial, P2(cos θ) = 0, and so any interaction which depends on this second-order Legendre polynomial vanishes at the magic angle. This property makes the magic angle of particular importance in magic angle spinning solid-state NMR spectroscopy. In magnetic resonance imaging, structures with ordered collagen, such as tendons and ligaments, oriented at the magic angle may appear hyperintense in some sequences; this is called the magic angle artifact or effect.
Nuclear magnetic resonance spectroscopy of proteins is a field of structural biology in which NMR spectroscopy is used to obtain information about the structure and dynamics of proteins, and also nucleic acids, and their complexes. The field was pioneered by Richard R. Ernst and Kurt Wüthrich at the ETH, and by Ad Bax, Marius Clore, Angela Gronenborn at the NIH, and Gerhard Wagner at Harvard University, among others. Structure determination by NMR spectroscopy usually consists of several phases, each using a separate set of highly specialized techniques. The sample is prepared, measurements are made, interpretive approaches are applied, and a structure is calculated and validated.
In nuclear chemistry and nuclear physics, J-couplings are mediated through chemical bonds connecting two spins. It is an indirect interaction between two nuclear spins that arises from hyperfine interactions between the nuclei and local electrons. In NMR spectroscopy, J-coupling contains information about relative bond distances and angles. Most importantly, J-coupling provides information on the connectivity of chemical bonds. It is responsible for the often complex splitting of resonance lines in the NMR spectra of fairly simple molecules.
Magnetic dipole–dipole interaction, also called dipolar coupling, refers to the direct interaction between two magnetic dipoles. Roughly speaking, the magnetic field of a dipole goes as the inverse cube of the distance, and the force of its magnetic field on another dipole goes as the first derivative of the magnetic field. It follows that the dipole-dipole interaction goes as the inverse fourth power of the distance.
Residual chemical shift anisotropy (RCSA) is the difference between the chemical shift anisotropy (CSA) of aligned and non-aligned molecules. It is normally three orders of magnitude smaller than the static CSA, with values on the order of parts-per-billion (ppb). RCSA is useful for structural determination and it is among the new developments in NMR spectroscopy.
Adriaan "Ad" Bax is a Dutch-American molecular biophysicist. He was born in the Netherlands and is the Chief of the Section on Biophysical NMR Spectroscopy at the National Institutes of Health. He is known for his work on the methodology of biomolecular NMR spectroscopy. He is a corresponding member of the Royal Netherlands Academy of Arts and Sciences, a member of the National Academy of Sciences, a fellow of the American Academy of Arts and Sciences, and a Foreign Member of the Royal Society.

Nuclear magnetic resonance (NMR) is a physical phenomenon in which nuclei in a strong constant magnetic field are perturbed by a weak oscillating magnetic field and respond by producing an electromagnetic signal with a frequency characteristic of the magnetic field at the nucleus. This process occurs near resonance, when the oscillation frequency matches the intrinsic frequency of the nuclei, which depends on the strength of the static magnetic field, the chemical environment, and the magnetic properties of the isotope involved; in practical applications with static magnetic fields up to ca. 20 tesla, the frequency is similar to VHF and UHF television broadcasts (60–1000 MHz). NMR results from specific magnetic properties of certain atomic nuclei. Nuclear magnetic resonance spectroscopy is widely used to determine the structure of organic molecules in solution and study molecular physics and crystals as well as non-crystalline materials. NMR is also routinely used in advanced medical imaging techniques, such as in magnetic resonance imaging (MRI). The original application of NMR to condensed matter physics is nowadays mostly devoted to strongly correlated electron systems. It reveals large many-body couplings by fast broadband detection and it should not to be confused with solid state NMR, which aims at removing the effect of the same couplings by Magic Angle Spinning techniques.
Nuclear magnetic resonance crystallography is a method which utilizes primarily NMR spectroscopy to determine the structure of solid materials on the atomic scale. Thus, solid-state NMR spectroscopy would be used primarily, possibly supplemented by quantum chemistry calculations, powder diffraction etc. If suitable crystals can be grown, any crystallographic method would generally be preferred to determine the crystal structure comprising in case of organic compounds the molecular structures and molecular packing. The main interest in NMR crystallography is in microcrystalline materials which are amenable to this method but not to X-ray, neutron and electron diffraction. This is largely because interactions of comparably short range are measured in NMR crystallography.
Nucleic acid NMR is the use of nuclear magnetic resonance spectroscopy to obtain information about the structure and dynamics of nucleic acid molecules, such as DNA or RNA. It is useful for molecules of up to 100 nucleotides, and as of 2003, nearly half of all known RNA structures had been determined by NMR spectroscopy.
CS-ROSETTA is a framework for structure calculation of biological macromolecules on the basis of conformational information from NMR, which is built on top of the biomolecular modeling and design software called ROSETTA. The name CS-ROSETTA for this branch of ROSETTA stems from its origin in combining NMR chemical shift (CS) data with ROSETTA structure prediction protocols. The software package was later extended to include additional NMR conformational parameters, such as Residual Dipolar Couplings (RDC), NOE distance restraints, pseudocontact chemical shifts (PCS) and restraints derived from homologous proteins. This software can be used together with other molecular modeling protocols, such as docking to model protein oligomers. In addition, CS-ROSETTA can be combined with chemical shift resonance assignment algorithms to create a fully automated NMR structure determination pipeline. The CS-ROSETTA software is freely available for academic use and can be licensed for commercial use. A software manual and tutorials are provided on the supporting website https://csrosetta.chemistry.ucsc.edu/.

In computational chemistry, conformational ensembles, also known as structural ensembles, are experimentally constrained computational models describing the structure of intrinsically unstructured proteins. Such proteins are flexible in nature, lacking a stable tertiary structure, and therefore cannot be described with a single structural representation. The techniques of ensemble calculation are relatively new on the field of structural biology, and are still facing certain limitations that need to be addressed before it will become comparable to classical structural description methods such as biological macromolecular crystallography.
The Biological Magnetic Resonance Data Bank is an open access repository of nuclear magnetic resonance (NMR) spectroscopic data from peptides, proteins, nucleic acids and other biologically relevant molecules. The database is operated by the University of Wisconsin–Madison and is supported by the National Library of Medicine. The BMRB is part of the Research Collaboratory for Structural Bioinformatics and, since 2006, it is a partner in the Worldwide Protein Data Bank (wwPDB). The repository accepts NMR spectral data from laboratories around the world and, once the data is validated, it is available online at the BMRB website. The database has also an ftp site, where data can be downloaded in the bulk. The BMRB has two mirror sites, one at the Protein Database Japan (PDBj) at Osaka University and one at the Magnetic Resonance Research Center (CERM) at the University of Florence in Italy. The site at Japan accepts and processes data depositions.

James J. Chou (周界文) is a Chinese American scientist and Professor of Biological Chemistry and Molecular Pharmacology at the Harvard Medical School. He is known for pioneering the use of Nuclear Magnetic Resonance (NMR) Spectroscopy to reveal the structural details of the membrane regions of cell surface proteins, particularly those of immune receptors and viral membrane proteins.

Spinach is an open-source magnetic resonance simulation package initially released in 2011 and continuously updated since. The package is written in Matlab and makes use of the built-in parallel computing and GPU interfaces of Matlab.


























