
Thiamine, also known as thiamin and vitamin B1, is a vitamin, an essential micronutrient, which cannot be made in the body. It is found in food and commercially synthesized to be a dietary supplement or medication. Food sources of thiamine include whole grains, legumes, and some meats and fish. Grain processing removes much of the thiamine content, so in many countries cereals and flours are enriched with thiamine. Supplements and medications are available to treat and prevent thiamine deficiency and disorders that result from it, including beriberi and Wernicke encephalopathy. Other uses include the treatment of maple syrup urine disease and Leigh syndrome. They are typically taken by mouth, but may also be given by intravenous or intramuscular injection.

In molecular biology, a riboswitch is a regulatory segment of a messenger RNA molecule that binds a small molecule, resulting in a change in production of the proteins encoded by the mRNA. Thus, an mRNA that contains a riboswitch is directly involved in regulating its own activity, in response to the concentrations of its effector molecule. The discovery that modern organisms use RNA to bind small molecules, and discriminate against closely related analogs, expanded the known natural capabilities of RNA beyond its ability to code for proteins, catalyze reactions, or to bind other RNA or protein macromolecules.

Cobalamin riboswitch is a cis-regulatory element which is widely distributed in 5' untranslated regions of vitamin B12 (Cobalamin) related genes in bacteria.
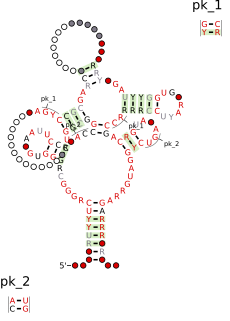
The FMN riboswitch is a highly conserved RNA element that is found frequently in the 5'-untranslated regions of prokaryotic mRNAs that encode for flavin mononucleotide (FMN) biosynthesis and transport proteins. This element is a metabolite-dependent riboswitch that directly binds FMN in the absence of proteins. In Bacillus subtilis, the riboswitch controls gene expression by causing premature transcription termination within the 5' untranslated region of the ribDEAHT operon and precluding access to the ribosome-binding site of ypaA mRNA.

The Lysine riboswitch is a metabolite binding RNA element found within certain messenger RNAs that serve as a precision sensor for the amino acid lysine. Allosteric rearrangement of mRNA structure is mediated by ligand binding, and this results in modulation of gene expression. Lysine riboswitch are most abundant in Firmicutes and Gammaproteobacteria where they are found upstream of a number of genes involved in lysine biosynthesis, transport and catabolism. The lysine riboswitch has also been identified independently and called the L box.

The PreQ1-I riboswitch is a cis-acting element identified in bacteria which regulates expression of genes involved in biosynthesis of the nucleoside queuosine (Q) from GTP. PreQ1 (pre-queuosine1) is an intermediate in the queuosine pathway, and preQ1 riboswitch, as a type of riboswitch, is an RNA element that binds preQ1. The preQ1 riboswitch is distinguished by its unusually small aptamer, compared to other riboswitches. Its atomic-resolution three-dimensional structure has been determined, with the PDB ID 2L1V.

A purine riboswitch is a sequence of ribonucleotides in certain messenger RNA (mRNA) that selectively binds to purine ligands via a natural aptamer domain. This binding causes a conformational change in the mRNA that can affect translation by revealing an expression platform for a downstream gene, or by forming a translation-terminating stem-loop. The ultimate effects of such translational regulation often take action to manage an abundance of the instigating purine, and might produce proteins that facilitate purine metabolism or purine membrane uptake.

The SAM-II riboswitch is a RNA element found predominantly in alpha-proteobacteria that binds S-adenosyl methionine (SAM). Its structure and sequence appear to be unrelated to the SAM riboswitch found in Gram-positive bacteria. This SAM riboswitch is located upstream of the metA and metC genes in Agrobacterium tumefaciens, and other methionine and SAM biosynthesis genes in other alpha-proteobacteria. Like the other SAM riboswitch, it probably functions to turn off expression of these genes in response to elevated SAM levels. A significant variant of SAM-II riboswitches was found in Pelagibacter ubique and related marine bacteria and called SAM-V. Also, like many structured RNAs, SAM-II riboswitches can tolerate long loops between their stems.
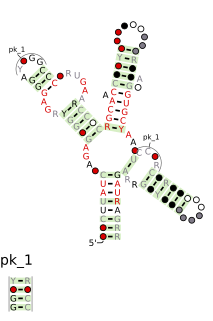
The SAM riboswitch is found upstream of a number of genes which code for proteins involved in methionine or cysteine biosynthesis in Gram-positive bacteria. Two SAM riboswitches in Bacillus subtilis that were experimentally studied act at the level of transcription termination control. The predicted secondary structure consists of a complex stem-loop region followed by a single stem-loop terminator region. An alternative and mutually exclusive form involves bases in the 3' segment of helix 1 with those in the 5' region of helix 5 to form a structure termed the anti-terminator form. When SAM is unbound, the anti-terminator sequence sequesters the terminator sequence so the terminator is unable to form, allowing the polymerase read-through the downstream gene. When S-Adenosyl methionine (SAM) is bound to the aptamer, the anti-terminator is sequestered by an anti-anti-terminator; the terminator forms and terminates the transcription. However, many SAM riboswitches are likely to regulate gene expression at the level of translation.

The ykkC/yxkD leader is a conserved RNA structure found upstream of the ykkC and yxkD genes in Bacillus subtilis and related genes in other bacteria. The function of this family is unclear for many years although it has been suggested that it may function to switch on efflux pumps and detoxification systems in response to harmful environmental molecules. The Thermoanaerobacter tengcongensis sequence AE013027 overlaps with that of purine riboswitch suggesting that the two riboswitches may work in conjunction to regulate the upstream gene which codes for TTE0584 (Q8RC62), a member of the permease family.
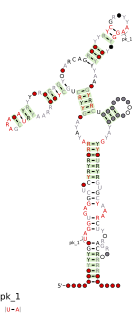
The Ykok leader or M-box is a Mg2+-sensing RNA structure that controls the expression of Magnesium ion transport proteins in bacteria. It is a distinct structure to the Magnesium responsive RNA element.

In enzymology, a hydroxyethylthiazole kinase is an enzyme that catalyzes the chemical reaction

The fluoride riboswitch is a conserved RNA structure identified by bioinformatics in a wide variety of bacteria and archaea. These RNAs were later shown to function as riboswitches that sense fluoride ions. These "fluoride riboswitches" increase expression of downstream genes when fluoride levels are elevated, and the genes are proposed to help mitigate the toxic effects of very high levels of fluoride.

The Downstream-peptide motif refers to a conserved RNA structure identified by bioinformatics in the cyanobacterial genera Synechococcus and Prochlorococcus and one phage that infects such bacteria. It was also detected in marine samples of DNA from uncultivated bacteria, which are presumably other species of cyanobacteria.

The glutamine riboswitch is a conserved RNA structure that was predicted by bioinformatics. It is present in a variety of lineages of cyanobacteria, as well as some phages that infect cyanobacteria. It is also found in DNA extracted from uncultivated bacteria living in the ocean that are presumably species of cyanobacteria.
SAM-V riboswitch is the fifth known riboswitch to bind S-adenosyl methionine (SAM). It was first discovered in the marine bacterium Candidatus Pelagibacter ubique and can also be found in marine metagenomes. SAM-V features a similar consensus sequence and secondary structure as the binding site of SAM-II riboswitch, but bioinformatics scans cluster the two aptamers independently. These similar binding pockets suggest that the two riboswitches have undergone convergent evolution.
Members of this protein family have been assigned as thiamine transporters by a phylogenomic analysis of families of genes regulated by the THI element, a broadly conserved RNA secondary structure element through which thiamine pyrophosphate (TPP) levels can regulate transcription of many genes related to thiamine transport, salvage, and de novo biosynthesis. Species with this protein always lack the ThiBPQ ABC transporter. In some species, yuaJ is the only THI-regulated gene. Evidence from Bacillus cereus indicates thiamine uptake is coupled to proton translocation.
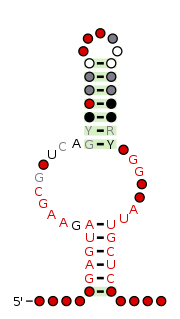
The NMT1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. NMT1 motif RNAs are found in Pseudomonadota. There is also one NMT1 RNA in each of Bacteroidota and Actinomycetota, but these appear to be the result of recent horizontal gene transfer or sequence contamination before or during genome sequencing

The uup RNA motif is a conserved RNA structure that was discovered by bioinformatics. uup motif RNAs are found in Firmicutes and Gammaproteobacteria.

Within the field of biochemistry, 4-amino-5-hydroxymethyl-2-methylpyrimidine (HMP) also known as toxopyrimidine together with its mono phosphate (HMP-P) and pyrophosphate (HMP-PP) esters are biogenetic precursors to the important biochemical cofactor thiamine pyrophosphate (TPP), a derivative of thiamine (vitamin B1).



















