Related Research Articles

Vibrio cholerae is a species of Gram-negative, facultative anaerobe and comma-shaped bacteria. The bacteria naturally live in brackish or saltwater where they attach themselves easily to the chitin-containing shells of crabs, shrimp, and other shellfish. Some strains of V. cholerae are pathogenic to humans and cause a deadly disease called cholera, which can be derived from the consumption of undercooked or raw marine life species.

Vibrio is a genus of Gram-negative bacteria, possessing a curved-rod (comma) shape, several species of which can cause foodborne infection, usually associated with eating undercooked seafood. Being highly salt tolerant and unable to survive in fresh water, Vibrio spp. are commonly found in various salt water environments. Vibrio spp. are facultative anaerobes that test positive for oxidase and do not form spores. All members of the genus are motile. They are able to have polar or lateral flagellum with or without sheaths. Vibrio species typically possess two chromosomes, which is unusual for bacteria. Each chromosome has a distinct and independent origin of replication, and are conserved together over time in the genus. Recent phylogenies have been constructed based on a suite of genes.
The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.

The gcvB RNA gene encodes a small non-coding RNA involved in the regulation of a number of amino acid transport systems as well as amino acid biosynthetic genes. The GcvB gene is found in enteric bacteria such as Escherichia coli. GcvB regulates genes by acting as an antisense binding partner of the mRNAs for each regulated gene. This binding is dependent on binding to a protein called Hfq. Transcription of the GcvB RNA is activated by the adjacent GcvA gene and repressed by the GcvR gene. A deletion of GcvB RNA from Y. pestis changed colony shape as well as reducing growth. It has been shown by gene deletion that GcvB is a regulator of acid resistance in E. coli. GcvB enhances the ability of the bacterium to survive low pH by upregulating the levels of the alternate sigma factor RpoS. A polymeric form of GcvB has recently been identified. Interaction of GcvB with small RNA SroC triggers the degradation of GcvB by RNase E, lifting the GcvB-mediated mRNA repression of its target genes.

The OmrA-B RNA gene family is a pair of homologous OmpR-regulated small non-coding RNA that was discovered in E. coli during two large-scale screens. OmrA-B is highly abundant in stationary phase, but low levels could be detected in exponentially growing cells as well. RygB is adjacent to RygA a closely related RNA. These RNAs bind to the Hfq protein and regulate gene expression by antisense binding. They negatively regulate the expression of several genes encoding outer membrane proteins, including cirA, CsgD, fecA, fepA and ompT by binding in the vicinity of the Shine-Dalgarno sequence, suggesting the control of these targets is dependent on Hfq protein and RNase E. Taken together, these data suggest that OmrA-B participates in the regulation of outer membrane composition, responding to environmental conditions.
RNAIII is a stable 514 nt regulatory RNA transcribed by the P3 promoter of the Staphylococcus aureus quorum-sensing agr system ). It is the major effector of the agr regulon, which controls the expression of many S. aureus genes encoding exoproteins and cell wall associated proteins plus others encoding regulatory proteins The RNAIII transcript also encodes the 26 amino acid δ-haemolysin peptide (Hld). RNAIII contains many stem loops, most of which match the Shine-Dalgarno sequence involved in translation initiation of the regulated genes. Some of these interactions are inhibitory, others stimulatory; among the former is the regulatory protein Rot. In vitro, RNAIII is expressed post exponentially, inhibiting translation of the surface proteins, notably protein A, while stimulating that of the exoproteins, many of which are tissue-degrading enzymes or cytolysins. Among the latter is the important virulence factor, α-hemolysin (Hla), whose translation RNAIII activates by preventing the formation of an inhibitory foldback loop in the hla mRNA leader.

RyhB RNA is a 90 nucleotide RNA that down-regulates a set of iron-storage and iron-using proteins when iron is limiting; it is itself negatively regulated by the ferric uptake repressor protein, Fur.

SgrS is a 227 nucleotide small RNA that is activated by SgrR in Escherichia coli during glucose-phosphate stress. The nature of glucose-phosphate stress is not fully understood, but is correlated with intracellular accumulation of glucose-6-phosphate. SgrS helps cells recover from glucose-phosphate stress by base pairing with ptsG mRNA and causing its degradation in an RNase E dependent manner. Base pairing between SgrS and ptsG mRNA also requires Hfq, an RNA chaperone frequently required by small RNAs that affect their targets through base pairing. The inability of cells expressing sgrS to create new glucose transporters leads to less glucose uptake and reduced levels of glucose-6-phosphate. SgrS is an unusual small RNA in that it also encodes a 43 amino acid functional polypeptide, SgrT, which helps cells recover from glucose-phosphate stress by preventing glucose uptake. The activity of SgrT does not affect the levels of ptsG mRNA of PtsG protein. It has been proposed that SgrT exerts its effects through regulation of the glucose transporter, PtsG.

Spot 42 (spf) RNA is a regulatory non-coding bacterial small RNA encoded by the spf gene. Spf is found in gammaproteobacteria and the majority of experimental work on Spot42 has been performed in Escherichia coli and recently in Aliivibrio salmonicida. In the cell Spot42 plays essential roles as a regulator in carbohydrate metabolism and uptake, and its expression is activated by glucose, and inhibited by the cAMP-CRP complex.

In molecular biology the ArcZ RNA is a small non-coding RNA (ncRNA). It is the functional product of a gene which is not translated into protein. ArcZ is an Hfq binding RNA that functions as an antisense regulator of a number of protein coding genes.

The Hfq protein encoded by the hfq gene was discovered in 1968 as an Escherichia coli host factor that was essential for replication of the bacteriophage Qβ. It is now clear that Hfq is an abundant bacterial RNA binding protein which has many important physiological roles that are usually mediated by interacting with Hfq binding sRNA.

An Hfq binding sRNA is an sRNA that binds the bacterial RNA binding protein called Hfq. A number of bacterial small RNAs which have been shown to bind to Hfq have been characterised . Many of these RNAs share a similar structure composed of three stem-loops. Several studies have expanded this list, and experimentally validated a total of 64 Hfq binding sRNA in Salmonella Typhimurium. A transcriptome wide study on Hfq binding sites in Salmonella mapped 126 Hfq binding sites within sRNAs. Genomic SELEX has been used to show that Hfq binding RNAs are enriched in the sequence motif 5′-AAYAAYAA-3′. Genome-wide study identified 40 candidate Hfq-dependent sRNAs in plant pathogen Erwinia amylovora. 12 of them were confirmed by Northern blot.
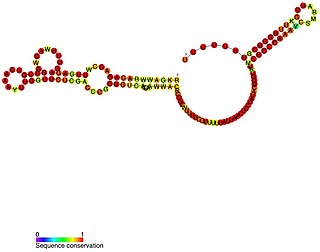

VrrA is a non-coding RNA that is conserved across all Vibrio species of bacteria and acts as a repressor for the synthesis of the outer membrane protein OmpA. This non-coding RNA was initially identified from Tn5 transposon mutant libraries of Vibrio cholerae and its location within the bacterial genome was mapped to the intergenic region between genes VC1741 and VC1743 by RACE analysis.
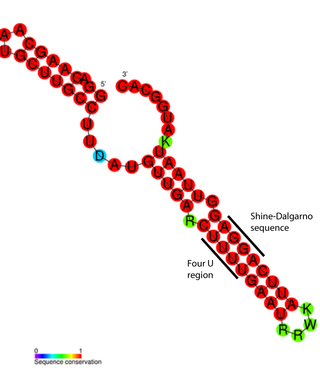
FourU thermometers are a class of non-coding RNA thermometers found in Salmonella. They are named 'FourU' due to the four highly conserved uridine nucleotides found directly opposite the Shine-Dalgarno sequence on hairpin II (pictured). RNA thermometers such as FourU control regulation of temperature via heat shock proteins in many prokaryotes. FourU thermometers are relatively small RNA molecules, only 57 nucleotides in length, and have a simple two-hairpin structure.

MicX sRNA is a small non-coding RNA found in Vibrio cholerae. It was given the name MicX as it has a similar function to MicA, MicC and MicF in E. coli. MicX sRNA negatively regulates an outer membrane protein and also a component of an ABC transporter. These interactions were predicted and then confirmed using a DNA microarray.
Bacterial small RNAs (bsRNA) are small RNAs produced by bacteria; they are 50- to 500-nucleotide non-coding RNA molecules, highly structured and containing several stem-loops. Numerous sRNAs have been identified using both computational analysis and laboratory-based techniques such as Northern blotting, microarrays and RNA-Seq in a number of bacterial species including Escherichia coli, the model pathogen Salmonella, the nitrogen-fixing alphaproteobacterium Sinorhizobium meliloti, marine cyanobacteria, Francisella tularensis, Streptococcus pyogenes, the pathogen Staphylococcus aureus, and the plant pathogen Xanthomonas oryzae pathovar oryzae. Bacterial sRNAs affect how genes are expressed within bacterial cells via interaction with mRNA or protein, and thus can affect a variety of bacterial functions like metabolism, virulence, environmental stress response, and structure.
Rsa RNAs are non-coding RNAs found in the bacterium Staphylococcus aureus. The shared name comes from their discovery, and does not imply homology. Bioinformatics scans identified the 16 Rsa RNA families named RsaA-K and RsaOA-OG. Others, RsaOH-OX, were found thanks to an RNomic approach. Although the RNAs showed varying expression patterns, many of the newly discovered RNAs were shown to be Hfq-independent and most carried a C-rich motif (UCCC).
John Mekalanos is a microbiologist who is primarily known for leading one of the first teams that reported the discovery of the type VI secretion system as well as his work on the pathogenicity of the bacterial species Vibrio cholerae, its toxin, and its secretion systems. Since 1998, he has been a member of the National Academy of Sciences.

Virginia L. Miller is a microbiologist known for her work on studying the factors leading to disease caused by bacteria. Miller is an elected fellow of the American Academy of Microbiology (2003) and a former Pew Charitable Trust Biomedical Scholar (1989).
References
- ↑ Bardill JP, Hammer BK (April 2012). "Non-coding sRNAs regulate virulence in the bacterial pathogen Vibrio cholerae". RNA Biology. 9 (4): 392–401. doi:10.4161/rna.19975. PMC 3384565 . PMID 22546941.
- ↑ Richard AL, Withey JH, Beyhan S, Yildiz F, DiRita VJ (December 2010). "The Vibrio cholerae virulence regulatory cascade controls glucose uptake through activation of TarA, a small regulatory RNA". Molecular Microbiology. 78 (5): 1171–1181. doi:10.1111/j.1365-2958.2010.07397.x. PMC 3064952 . PMID 21091503.
- ↑ Bradley ES, Bodi K, Ismail AM, Camilli A (July 2011). "A genome-wide approach to discovery of small RNAs involved in regulation of virulence in Vibrio cholerae". PLOS Pathogens. 7 (7): e1002126. doi: 10.1371/journal.ppat.1002126 . PMC 3136459 . PMID 21779167.