Helicobacter pylori, previously known as Campylobacter pylori, is a gram-negative, microaerophilic, spiral (helical) bacterium usually found in the stomach. Its helical shape is thought to have evolved in order to penetrate the mucoid lining of the stomach and thereby establish infection. The bacterium was first identified in 1982 by the Australian doctors Barry Marshall and Robin Warren. H. pylori has been associated with cancer of the mucosa-associated lymphoid tissue in the stomach, esophagus, colon, rectum, or tissues around the eye, and of lymphoid tissue in the stomach.

Antisense RNA (asRNA), also referred to as antisense transcript, natural antisense transcript (NAT) or antisense oligonucleotide, is a single stranded RNA that is complementary to a protein coding messenger RNA (mRNA) with which it hybridizes, and thereby blocks its translation into protein. asRNAs have been found in both prokaryotes and eukaryotes, and can be classified into short and long non-coding RNAs (ncRNAs). The primary function of asRNA is regulating gene expression. asRNAs may also be produced synthetically and have found wide spread use as research tools for gene knockdown. They may also have therapeutic applications.
The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.

DicF RNA is a non-coding RNA that is an antisense inhibitor of cell division gene ftsZ. DicF is bound by the Hfq protein which enhances its interaction with its targets. Pathogenic E. coli strains possess multiple copies of sRNA DicF in their genomes, while non-pathogenic strains do not. DicF and Hfq are both necessary to reduce FtsZ protein levels, leading to cell filamentation under anaerobic conditions.
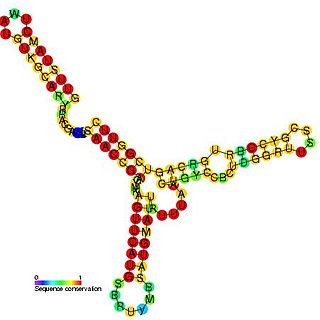
Anti-Q RNA is a small ncRNA from the conjugal plasmid pCF10 of Enterococcus faecalis. It is coded in cis to its regulatory target, prgQ, but can also act in trans. Anti-Q is known to interact with nascent prgQ transcripts to allow formation of an intrinsic terminator, or attenuator, thus preventing transcription of downstream genes. This mode of regulation is essentially the same as that of the countertranscript-driven attenuators that control copy number in pT181, pAMbeta1 and pIP501 and related Staphylococcal plasmids.
RNAIII is a stable 514 nt regulatory RNA transcribed by the P3 promoter of the Staphylococcus aureus quorum-sensing agr system ). It is the major effector of the agr regulon, which controls the expression of many S. aureus genes encoding exoproteins and cell wall associated proteins plus others encoding regulatory proteins The RNAIII transcript also encodes the 26 amino acid δ-haemolysin peptide (Hld). RNAIII contains many stem loops, most of which match the Shine-Dalgarno sequence involved in translation initiation of the regulated genes. Some of these interactions are inhibitory, others stimulatory; among the former is the regulatory protein Rot. In vitro, RNAIII is expressed post exponentially, inhibiting translation of the surface proteins, notably protein A, while stimulating that of the exoproteins, many of which are tissue-degrading enzymes or cytolysins. Among the latter is the important virulence factor, α-hemolysin (Hla), whose translation RNAIII activates by preventing the formation of an inhibitory foldback loop in the hla mRNA leader.

Spot 42 (spf) RNA is a regulatory non-coding bacterial small RNA encoded by the spf gene. Spf is found in gammaproteobacteria and the majority of experimental work on Spot42 has been performed in Escherichia coli and recently in Aliivibrio salmonicida. In the cell Spot42 plays essential roles as a regulator in carbohydrate metabolism and uptake, and its expression is activated by glucose, and inhibited by the cAMP-CRP complex.

The MicA RNA is a small non-coding RNA that was discovered in E. coli during a large scale screen. Expression of SraD is highly abundant in stationary phase, but low levels could be detected in exponentially growing cells as well.

The hok/sok system is a postsegregational killing mechanism employed by the R1 plasmid in Escherichia coli. It was the first type I toxin-antitoxin pair to be identified through characterisation of a plasmid-stabilising locus. It is a type I system because the toxin is neutralised by a complementary RNA, rather than a partnered protein.
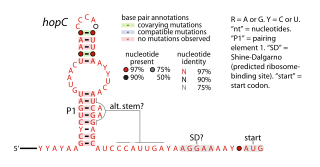
The hopC RNA motif is a predicted cis-regulatory element identified by a bioinformatic screen for conserved RNA secondary structures. hopC RNAs are exclusively found within bacteria classified within the genus Helicobacter, some of which are human pathogens that infect the stomach and can cause ulcers.

The psaA RNA motif describes a class of RNAs with a common secondary structure. psaA RNAs are exclusively found in locations that presumably correspond to the 5' untranslated regions of operons formed of psaA and psaB genes. For this reason, it was hypothesized that psaA RNAs function as cis-regulatory elements of these genes. The psaAB genes encode proteins that form subunits in the photosystem I structure used for photosynthesis. psaA RNAs have been detected only in cyanobacteria, which is consistent with their association with photosynthesis.
Bacterial small RNAs (bsRNA) are small RNAs produced by bacteria; they are 50- to 500-nucleotide non-coding RNA molecules, highly structured and containing several stem-loops. Numerous sRNAs have been identified using both computational analysis and laboratory-based techniques such as Northern blotting, microarrays and RNA-Seq in a number of bacterial species including Escherichia coli, the model pathogen Salmonella, the nitrogen-fixing alphaproteobacterium Sinorhizobium meliloti, marine cyanobacteria, Francisella tularensis, Streptococcus pyogenes, the pathogen Staphylococcus aureus, and the plant pathogen Xanthomonas oryzae pathovar oryzae. Bacterial sRNAs affect how genes are expressed within bacterial cells via interaction with mRNA or protein, and thus can affect a variety of bacterial functions like metabolism, virulence, environmental stress response, and structure.

A toxin-antitoxin system consists of a "toxin" and a corresponding "antitoxin", usually encoded by closely linked genes. The toxin is usually a protein while the antitoxin can be a protein or an RNA. Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK).
Rsa RNAs are non-coding RNAs found in the bacterium Staphylococcus aureus. The shared name comes from their discovery, and does not imply homology. Bioinformatics scans identified the 16 Rsa RNA families named RsaA-K and RsaOA-OG. Others, RsaOH-OX, were found thanks to an RNomic approach. Although the RNAs showed varying expression patterns, many of the newly discovered RNAs were shown to be Hfq-independent and most carried a C-rich motif (UCCC).
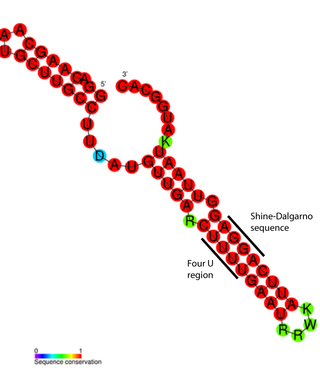
An RNA thermometer is a temperature-sensitive non-coding RNA molecule which regulates gene expression. RNA thermometers often regulate genes required during either a heat shock or cold shock response, but have been implicated in other regulatory roles such as in pathogenicity and starvation.
In molecular biology, the PyrC leader is a cis-regulatory RNA element found at the 5' of the PyrC mRNA in Enterobacteria. The PyrC gene encodes Dihydroorotase, an enzyme involved in pyrimidine biosynthesis. The PyrC leader regulates expression of PyrC. Translation initiation can occur at four different sites within this leader sequence, under high CTP conditions the translation initiation site is upstream of that used under low CTP conditions, additional cytosine residues are incorporated into the mRNA resulting in the formation of an RNA hairpin. This hairpin blocks ribosome-binding at the Shine-Dalgarno sequence, and therefore blocks expression of PyrC. Under low CTP conditions the initiation site is further downstream and does not result in hairpin formation, so the ribosome can bind to the Shine-Dalgarno sequence and PyrC is expressed.

cII or transcriptional activator II is a DNA-binding protein and important transcription factor in the life cycle of lambda phage. It is encoded in the lambda phage genome by the 291 base pair cII gene. cII plays a key role in determining whether the bacteriophage will incorporate its genome into its host and lie dormant (lysogeny), or replicate and kill the host (lysis).
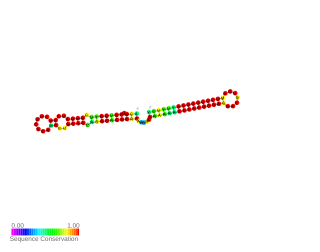
s-SodF RNA is a non-coding RNA (ncRNA) molecule identified in Streptomyces coelicolor. It is produced from sodF mRNA by cleavage of about 90 nucleotides from its 3′UTR. However it does not affect the function of sodF mRNA, but It acts on another mRNA called sodN. s-SodF RNA has a sequence complementary to sodN mRNA from the 5′-end up to the ribosome binding site. It pairs with sodN mRNA, blocks its translation and facilitates sodN mRNA decay. In Streptomyces sodF and sodN genes produce FeSOD and NiSOD superoxide dismutases containing Fe and Ni respectively. Their expression is inversely regulated by nickel-specific Fur-family regulator called Nur. When Ni is present Nur directly represses sodF transcription, and indirectly induces sodN.
Accessory gene regulator (agr) is a complex 5 gene locus that is a global regulator of virulence in Staphylococcus aureus. It encodes a two-component transcriptional quorum-sensing (QS) system activated by an autoinducing, thiolactone-containing cyclic peptide (AIP).
The Bacteroides thetaiotaomicron genome contains hundreds of small RNAs (sRNAs), discovered through RNA sequencing. These include canonical housekeeping RNA species such as the 6S RNA (SsrS), tmRNA (SsrA), M1 RNA (RnpB) and 4.5S RNA (Ffs) as well as several hundred cis and trans encoded small RNAs. More than 20 candidates have been validated with northern blots and the structures of several members have been characterized through in silico analyses and chemical probing experiments.












