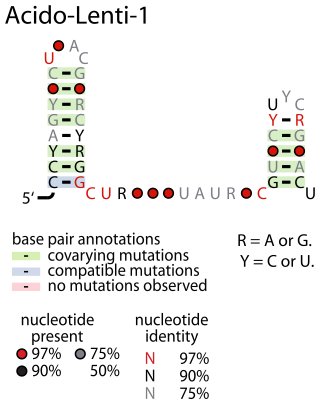
The Acido-Lenti-1 RNA motif describes a predicted non-coding RNA that is found in bacteria within the phyla acidobacteriota and lentisphaerota. It is sometimes found nearby to group II introns, but the reason for this apparent association is unknown.

The Bacillaceae-1 RNA motif is a conserved RNA structure identified by bioinformatics within bacteria in the family bacillaceae. The RNA is presumed to operate as a non-coding RNA, and is sometimes adjacent to operons containing ribosomal RNAs. The most characteristic feature is two terminal loops that have the nucleotide consensus RUCCU, where R is either A or G. The motif might be related to the Desulfotalea-1 RNA motif, as the motifs share some similarity in conserved features, and the Desulfotalea-1 RNA motif is also sometimes adjacent to ribosomal RNA operons.
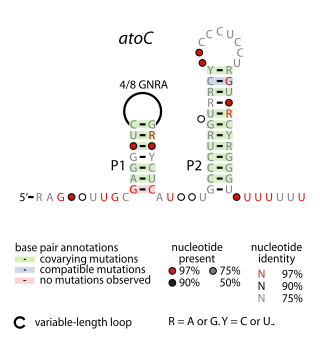
The atoC RNA motif is a conserved RNA-like structure identified by bioinformatics. It consistently appears upstream of protein-coding gene that are predicted to encode oxidoreductase activity, dihydropteroate synthase or DNA-binding response regulators.

The Bacteroidales-1 RNA motif is a conserved RNA structure identified by bioinformatics. It has been identified only in bacteria within the order (biology) Bacteroidales. Its presumed length is marked by a promoter on one end that conforms to an alternate consensus sequence that is common in the phylum Bacteroidota, and its 3′ end is indicated by predicted transcription terminators. It is often located downstream of a gene that encodes the L20 ribosomal subunit, although it is unclear whether there is a functional reason underlying this apparent association.

The Bacteroides-1 RNA motif is a conserved RNA structure identified in bacteria within the genus Bacteroides. The RNAs are typically found downstream of of genes that participate in the synthesis of exopolysaccharides of unknown types. It is possible that Bacteroides-1 RNAs regulate the upstream genes, but since this mode of regulation is unusual in bacteria, it is more likely that the structure functions as a non-coding RNA.
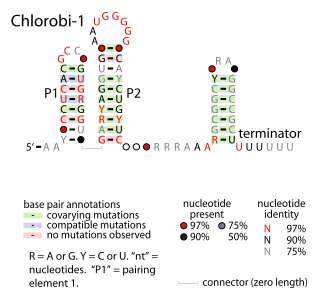
The Chlorobi-1 RNA motif is a conserved RNA secondary structure identified by bioinformatics. It is predicted to be used only by Chlorobiota, a phylum of bacteria. The motif consists of two stem-loops that are followed by an apparent rho-independent transcription terminator. The motif is presumed to function as an independently transcribed non-coding RNA.

The Chloroflexi-1 RNA motif is a conserved RNA structure detected by bioinformatics within the species Chloroflexus aggregans. C. aggregans has three predicted Chloroflexi-1 RNAs, which are located nearby to one another. This arrangement might suggest a repetitive element. C. aggregans is classified as belonging to the bacterial phylum Chloroflexota.

The Collinsella-1 RNA motif denotes a particular conserved RNA structure discovered by bioinformatics. Of the six sequences belonging to this motif that were originally identified, five are from uncultivated bacteria residing in the human gut, while only the sixth is in a cultivated species, Collinsella aerofaciens. The evidence supporting the stem-loops designated as "P1" and "P2" is ambiguous.

The Cyano-2 RNA motif is a conserved RNA structure identified by bioinformatics. Cyano-2 RNAs are found in Cyanobacterial species classified within the genus Synechococcus. Many terminal loops in the two conserved stem-loops contain the nucleotide sequence GCGA, and these sequences might in some cases form stable GNRA tetraloops. Since the two stem-loops are somewhat distant from one another it is possible that they represent two independent non-coding RNAs that are often or always co-transcribed. The region one thousand base pairs upstream of predicted Cyano-2 RNAs is usually devoid of annotated features such as RNA or protein-coding genes. This absence of annotated genes within one thousand base pairs is relatively unusual within bacteria.

The Downstream-peptide motif refers to a conserved RNA structure identified by bioinformatics in the cyanobacterial genera Synechococcus and Prochlorococcus and one phage that infects such bacteria. It was also detected in marine samples of DNA from uncultivated bacteria, which are presumably other species of cyanobacteria.
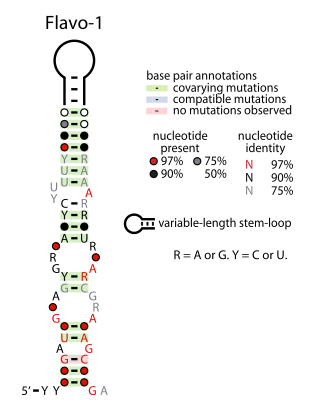
The Flavo-1 RNA motif is a conserved RNA structure that was identified by bioinformatics. The vast majority of Flavo-1 RNAs are found in Flavobacteria, but some were detected in the phylum Bacteroidota, which contains Flavobacteria, or the phylum Spirochaetota, which is evolutionarily related to Bacteroidota. It was presumed that Flavo-1 RNAs function as non-coding RNAs.

The Gut-1 RNA motif is a conserved RNA structure identified by bioinformatics. These RNAs are present in environmental sequences, and as of 2010 are not known to be present in any species that has been grown under laboratory conditions. Gut-1 RNA is exclusively found in DNA from uncultivated bacteria present in samples from the human gut.

The manA RNA motif refers to a conserved RNA structure that was identified by bioinformatics. Instances of the manA RNA motif were detected in bacteria in the genus Photobacterium and phages that infect certain kinds of cyanobacteria. However, most predicted manA RNA sequences are derived from DNA collected from uncultivated marine bacteria. Almost all manA RNAs are positioned such that they might be in the 5' untranslated regions of protein-coding genes, and therefore it was hypothesized that manA RNAs function as cis-regulatory elements. Given the relative complexity of their secondary structure, and their hypothesized cis-regulatory role, they might be riboswitches.

The Methylobacterium-1 RNA motif is a conserved RNA structure discovered using bioinformatics. Almost all known examples of this RNA are found in DNA extracted from marine bacteria. However, one instance is predicted in Methylobacterium sp. 4-46, a species of alphaproteobacteria. The motif is presumed to function as a non-coding RNA.
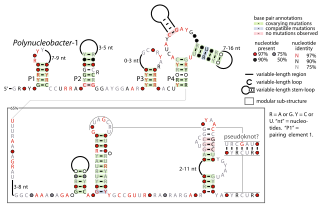
The Polynucleobacter-1 RNA motif is a conserved RNA structure that was identified by bioinformatics. The RNA structure is predominantly located in genome sequences derived from DNA extracted from uncultivated marine samples. However it was also predicted in the genome of Polynucleobacter species QLW-P1DMWA-1, a kind of betaproteobacteria. The RNAs are often located near to a conserved gene that might be homologous to a gene found in a phage that infects cyanobacteria. However, it is unknown if the RNA is used by phages.

The Pyrobac-1 RNA motif is a conserved RNA structure discovered by bioinformatics. RNAs conforming to this motif have been found only in Pyrobaculum, a genus of archaea. Instances of the motif are hypothesized to function as non-coding RNAs. The motif has been shown to be part of sRNA202 and sRNA203 canonical and noncanonical pseudouridine guide RNAs in Pyrobaculum.

The rmf RNA motif is a conserved RNA structure that was originally detected using bioinformatics. rmf RNAs are consistently foundwithin species classified into the genus Pseudomonas, and is located potentially in the 5′ untranslated regions of rmf genes. These genes encodes the ribosome modulation factor protein, which affects the translation of genes by modifying ribosome structure in response to stress such as starvation. This ribosome modulation is a part of the stringent response in bacteria. The likely biological role of rmf RNAs is ambiguous. Since the RNA could be in the 5′ UTRs of protein-coding genes, it was hypothesized that it functions as a cis-regulatory element. This hypothesis is bolstered by the observation that ribosome modulation factor binds ribosomal RNA, and many cis-regulatory RNAs called ribosomal protein leaders participate in a feedback regulation mechanism by binding to proteins that normally bind to ribosomal RNA. However, since rmf RNAs are not very close to the rmf genes, they might function as non-coding RNAs.

The Rhizobiales-2 RNA motif is a set of RNAs found in certain bacteria that are presumed to be homologous because they conserve a common primary and secondary structure. The motif was discovered using bioinformatics, and is found only within bacteria that belong to the order Hyphomicrobiales, in turn a kind of alphaproteobacteria. Because Rhizobiales-2 RNAs are not consistently located in proximity to genes of a consistent class or function, these RNAs are presumed to function as non-coding RNAs.
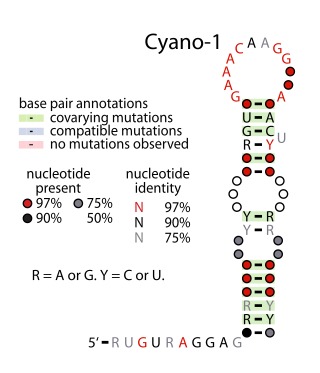
Yfr2 is a family of non-coding RNAs. Members of the Yrf2 family have been identified in almost all studied species of cyanobacteria. The family was identified through a bioinformatics screen of published cyanobacterial genomes, having previously been grouped in a family of Yfr2–5.

Tetrahydrofolate riboswitches are a class of homologous RNAs in certain bacteria that bind tetrahydrofolate (THF). It is almost exclusively located in the probable 5' untranslated regions of protein-coding genes, and most of these genes are known to encode either folate transporters or enzymes involved in folate metabolism. For these reasons it was inferred that the RNAs function as riboswitches. THF riboswitches are found in a variety of Bacillota, specifically the orders Clostridiales and Lactobacillales, and more rarely in other lineages of bacteria. The THF riboswitch was one of many conserved RNA structures found in a project based on comparative genomics. The 3-d structure of the tetrahydrofolate riboswitch has been solved by separate groups using X-ray crystallography. These structures were deposited into the Protein Data Bank under accessions 3SD1 and 3SUX, with other entries containing variants.




















