Antiviral drugs are a class of medication used specifically for treating viral infections rather than bacterial ones. Most antivirals are used for specific viral infections, while a broad-spectrum antiviral is effective against a wide range of viruses. Unlike most antibiotics, antiviral drugs do not destroy their target pathogen; instead they inhibit their development.
Gene silencing is the regulation of gene expression in a cell to prevent the expression of a certain gene. Gene silencing can occur during either transcription or translation and is often used in research. In particular, methods used to silence genes are being increasingly used to produce therapeutics to combat cancer and diseases, such as infectious diseases and neurodegenerative disorders.

A Morpholino, also known as a Morpholino oligomer and as a phosphorodiamidate Morpholino oligomer (PMO), is a type of oligomer molecule used in molecular biology to modify gene expression. Its molecular structure has DNA bases attached to a backbone of methylenemorpholine rings linked through phosphorodiamidate groups. Morpholinos block access of other molecules to small specific sequences of the base-pairing surfaces of ribonucleic acid (RNA). Morpholinos are used as research tools for reverse genetics by knocking down gene function.
In molecular biology and genetics, the sense of nucleic acid molecules is the nature of their roles and their complementary molecules' nucleic acid units' roles in specifying amino acids. Depending on the context within molecular biology, sense may have slightly different meanings.
The fertility factor allows genes to be transferred from one bacterium carrying the factor to another bacterium lacking the factor by conjugation. The F factor is carried on the F episome, the first episome to be discovered. Unlike other plasmids, F factor is constitutive for transfer proteins due to a mutation in the gene finO. The F plasmid belongs to a class of conjugative plasmids that control sexual functions of bacteria with a fertility inhibition (Fin) system.
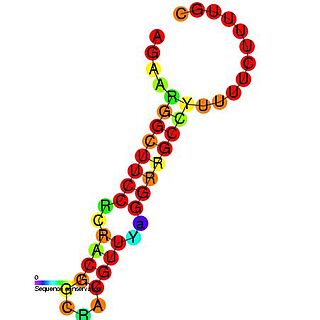
In molecular biology ctRNA is a plasmid encoded noncoding RNA that binds to the mRNA of repB and causes translational inhibition. ctRNA is encoded by plasmids and functions in rolling circle replication to maintain a low copy number. In Corynebacterium glutamicum, it achieves this by antisense pairing with the mRNA of RepB, a replication initiation protein. In Enterococcus faecium the plasmid pJB01 contains three open reading frames, copA, repB, and repC. The pJB01 ctRNA is coded on the opposite strand from the copA/repB intergenic region and partially overlaps an atypical ribosome binding site for repB.
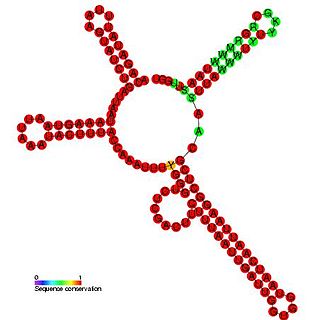
Plasmid RNAIII is a non-coding RNA found in bacterial plasmids including pIP501. RNAIII acts by transcriptional attenuation of the essential repR-mRNA. RNAIII is composed of four stem-loops with loops L3 and L4 that interact with the RNA target.
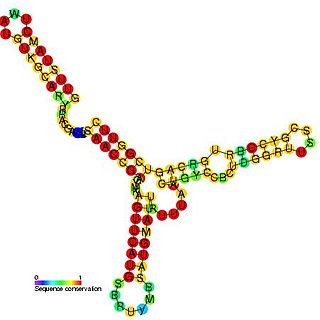
Anti-Q RNA is a small ncRNA from the conjugal plasmid pCF10 of Enterococcus faecalis. It is coded in cis to its regulatory target, prgQ, but can also act in trans. Anti-Q is known to interact with nascent prgQ transcripts to allow formation of an intrinsic terminator, or attenuator, thus preventing transcription of downstream genes. This mode of regulation is essentially the same as that of the countertranscript-driven attenuators that control copy number in pT181, pAMbeta1 and pIP501 and related Staphylococcal plasmids.
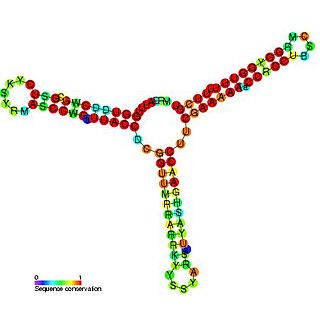
RNAI is a non-coding RNA that is an antisense repressor of the replication of some E. coli plasmids, including ColE1. Plasmid replication is usually initiated by RNAII, which acts as a primer by binding to its template DNA. The complementary RNAI binds RNAII prohibiting it from its initiation role. The rate of degradation of RNAI is therefore a major factor in the control of plasmid replication. This rate of degradation is aided by the pcnB gene product, which polyadenylates the 3' end of RNAI targeting it for degradation by PNPase.
The hok/sok system is a postsegregational killing mechanism employed by the R1 plasmid in Escherichia coli. It was the first type I toxin-antitoxin pair to be identified through characterisation of a plasmid-stabilising locus. It is a type I system because the toxin is neutralised by a complementary RNA, rather than a partnered protein.

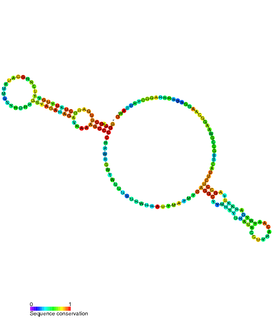
The Bacillus-plasmid RNA motif is a predicted conserved RNA structure usually located in plasmids. It is known in species under the genera Bacillus and Lactobacillus. In Bacillus subtilis, it is found upstream of the hypothetical gene ydcS, whose function is unknown.

The lactis-plasmid RNA motif is a conserved RNA structure identified by bioinformatics. The RNAs are restricted to lactic acid bacteria, and are especially common in Lactococcus lactis. They typically lie near to repB genes, and are almost found in plasmids. This data suggested that lactis-plasmid RNAs participate in the control of plasmid abundance. However, many of the plasmids that carry lactis-plasmid RNAs also carry ctRNA-pND324 RNAs, which are involved in plasmid copy count regulation. Therefore lactis-plasmid RNAs might have a different function.

The traJ-II RNA motif is a conserved RNA structure discovered in bacteria by using bioinformatics. traJ-II RNAs appear to be in the 5' untranslated regions of protein-coding genes called traJ, which functions in the process of bacterial conjugation. A previously identified motif known as TraJ 5' UTR is also found upstream of traJ genes functions as the target of FinP antisense RNAs, so it is possible that traJ-II RNAs play a similar role as targets of an antisense RNA. However, some sequence features within the traJ-II RNA motif suggest that the biological RNA might be transcribed from the reverse-complement strand. Thus is it unclear whether traJ-II function as cis-regulatory elements. traJ-II RNAs are found in a variety of proteobacteria.
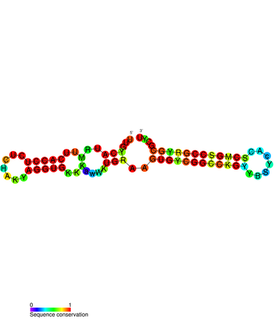
PtaRNA1 is a family of non-coding RNAs. Homologs of PtaRNA1 can be found in the proteobacteria families, Betaproteobacteria and Gammaproteobacteria. In all cases the PtaRNA1 is located anti-sense to a short protein-coding gene. In Xanthomonas campestris pv. vesicatoria, this gene is annotated as XCV2162 and is included in the plasmid toxin family of proteins.

A toxin-antitoxin system is a set of two or more closely linked genes that together encode both a "toxin" protein and a corresponding "antitoxin". When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK). Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies.

The FlmA-FlmB toxin-antitoxin system consists of FlmB RNA, a family of non-coding RNAs and the protein toxin FlmA. The FlmB RNA transcript is 100 nucleotides in length and is homologous to sok RNA from the hok/sok system and fulfills the identical function as a post-segregational killing (PSK) mechanism.
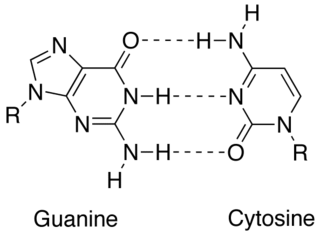
In molecular biology, complementarity describes a relationship between two structures each following the lock-and-key principle. In nature complementarity is the base principle of DNA replication and transcription as it is a property shared between two DNA or RNA sequences, such that when they are aligned antiparallel to each other, the nucleotide bases at each position in the sequences will be complementary, much like looking in the mirror and seeing the reverse of things. This complementary base pairing allows cells to copy information from one generation to another and even find and repair damage to the information stored in the sequences.
Plasmids must regulate their copy number to ensure that they do not excessively burden the host or become lost during cell division. Plasmids may be either high copy number plasmids or low copy number plasmids; the regulation mechanisms between these two types are often significantly different. Biotechnology applications may involve engineering plasmids to allow a very high copy number. For example, pBR322 is a low copy number plasmid from which several very high copy number cloning vectors have been derived.
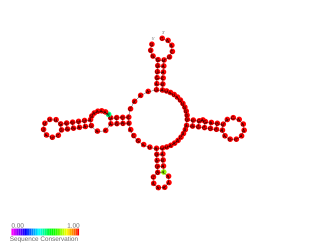
RnaG is a small regulatory non-coding RNA encoded by the virulence plasmid of Shigella flexneri, a Gram-negative pathogenic bacterium that causes human bacillary dysentery. It is a first regulatory RNA characterised in S. flexneri. The RNA is 450 nucleotides long and it contains a region with specific secondary structure that interacts with icsA mRNA and forms a transcription terminator. Acting as antisense, RnaG is transcribed from the complementary strand of its target, icsA mRNA. The activity of the incA protein is crucial for spreading of the bacterial pathogen in the host cells.


















