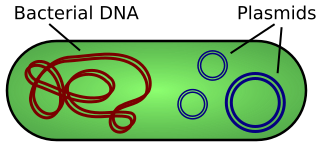
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; however, plasmids are sometimes present in archaea and eukaryotic organisms. In nature, plasmids often carry genes that benefit the survival of the organism and confer selective advantage such as antibiotic resistance. While chromosomes are large and contain all the essential genetic information for living under normal conditions, plasmids are usually very small and contain only additional genes that may be useful in certain situations or conditions. Artificial plasmids are widely used as vectors in molecular cloning, serving to drive the replication of recombinant DNA sequences within host organisms. In the laboratory, plasmids may be introduced into a cell via transformation. Synthetic plasmids are available for procurement over the internet.
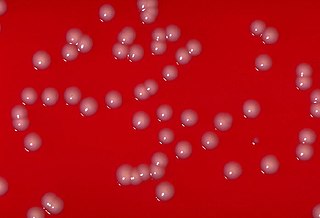
Corynebacterium is a genus of bacteria that are Gram-positive and most are aerobic. They are bacilli (rod-shaped), and in some phases of life they are, more specifically, club-shaped, which inspired the genus name.

A tumour inducing (Ti) plasmid is a plasmid found in pathogenic species of Agrobacterium, including A. tumefaciens, A. rhizogenes, A. rubi and A. vitis.
The fertility factor allows genes to be transferred from one bacterium carrying the factor to another bacterium lacking the factor by conjugation. The F factor was the first plasmid to be discovered. Unlike other plasmids, F factor is constitutive for transfer proteins due to a mutation in the gene finO. The F plasmid belongs to a class of conjugative plasmids that control sexual functions of bacteria with a fertility inhibition (Fin) system.
P1 is a temperate bacteriophage that infects Escherichia coli and some other bacteria. When undergoing a lysogenic cycle the phage genome exists as a plasmid in the bacterium unlike other phages that integrate into the host DNA. P1 has an icosahedral head containing the DNA attached to a contractile tail with six tail fibers. The P1 phage has gained research interest because it can be used to transfer DNA from one bacterial cell to another in a process known as transduction. As it replicates during its lytic cycle it captures fragments of the host chromosome. If the resulting viral particles are used to infect a different host the captured DNA fragments can be integrated into the new host's genome. This method of in vivo genetic engineering was widely used for many years and is still used today, though to a lesser extent. P1 can also be used to create the P1-derived artificial chromosome cloning vector which can carry relatively large fragments of DNA. P1 encodes a site-specific recombinase, Cre, that is widely used to carry out cell-specific or time-specific DNA recombination by flanking the target DNA with loxP sites.
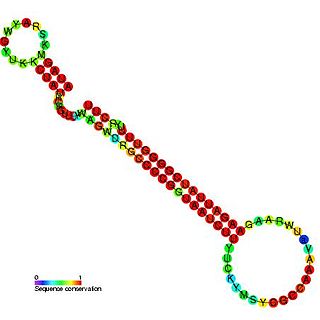
CopA-like RNA is a family of non-coding RNAs found on the R1 plasmid.
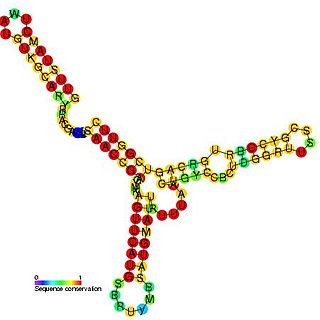
Anti-Q RNA is a small ncRNA from the conjugal plasmid pCF10 of Enterococcus faecalis. It is coded in cis to its regulatory target, prgQ, but can also act in trans. Anti-Q is known to interact with nascent prgQ transcripts to allow formation of an intrinsic terminator, or attenuator, thus preventing transcription of downstream genes. This mode of regulation is essentially the same as that of the countertranscript-driven attenuators that control copy number in pT181, pAMbeta1 and pIP501 and related Staphylococcal plasmids.

R1162-like plasmid antisense RNA is a 75-base RNA molecule which negatively regulates the RepI region of the plasmid. The protein product of this gene region, along with another protein, controls the copy number of the 8.75kB R1162 plasmid.
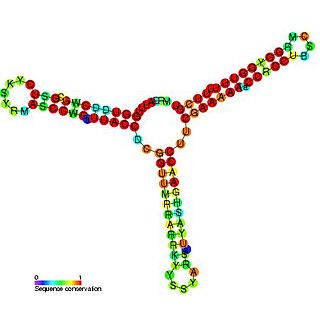
RNAI is a non-coding RNA that is an antisense repressor of the replication of some E. coli plasmids, including ColE1. Plasmid replication is usually initiated by RNAII, which acts as a primer by binding to its template DNA. The complementary RNAI binds RNAII prohibiting it from its initiation role. The rate of degradation of RNAI is therefore a major factor in the control of plasmid replication. This rate of degradation is aided by the pcnB gene product, which polyadenylates the 3' end of RNAI targeting it for degradation by PNPase.
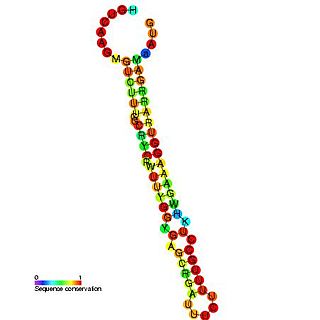
The S-element is an RNA element found in p42d and related plasmids. The S-element has multiple functions and is believed to act as a negative regulator of repC transcription, and be required for efficient replication and act as a translational enhancer of repC.

Rhodococcus equi is a Gram-positive coccobacillus bacterium. The organism is commonly found in dry and dusty soil and can be important for diseases of domesticated animals. The frequency of infection can reach near 60%. R. equi is an important pathogen causing pneumonia in foals. Since 2008, R. equi has been known to infect wild boar and domestic pigs. R. equi can infect humans. At-risk groups are immunocompromised people, such as HIV-AIDS patients or transplant recipients. Rhodococcus infection in these patients resemble clinical and pathological signs of pulmonary tuberculosis. It is facultative intracellular.
PcrA, standing for plasmid copy reduced is a helicase that was originally discovered in a screen for chromosomally encoded genes that are affected in plasmid rolling circle replication in the Gram-positive pathogen Staphylococcus aureus.

The lactis-plasmid RNA motif is a conserved RNA structure identified by bioinformatics. The RNAs are restricted to lactic acid bacteria, and are especially common in Lactococcus lactis. They typically lie near to repB genes, and are almost found in plasmids. This data suggested that lactis-plasmid RNAs participate in the control of plasmid abundance. However, many of the plasmids that carry lactis-plasmid RNAs also carry ctRNA-pND324 RNAs, which are involved in plasmid copy count regulation. Therefore lactis-plasmid RNAs might have a different function.

The regulatory region of the repZ gene, which encodes the replication initiator of plasmid ColIb-P9, contains a pseudoknot. This acts as a molecular switch controlling translation of repZ and repY.
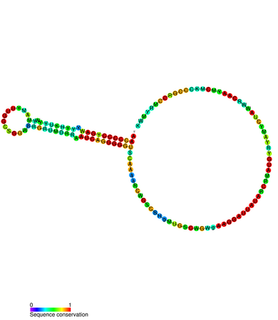
In plasmids, the regulatory region of repBA gene forms a pseudoknot. The repA gene, which encodes a protein likely to function as an initiator for replication, and the repB gene are translationally coupled. The leader sequence of the repA mRNA contains two complementary sequences of 8 bases. Base-pairing between these two sequences forms a pseudoknot which is essential for translation. The first of these complementary sequences is found within a stem-loop, which forms a target for RNAI. Binding of RNAI to this stem-loop inhibits pseudoknot formation and translation of RepA.

A toxin-antitoxin system is a set of two or more closely linked genes that together encode both a "toxin" protein and a corresponding "antitoxin". Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK).
Plasmid-mediated resistance is the transfer of antibiotic resistance genes which are carried on plasmids. The plasmids can be transferred between bacteria within the same species or between different species via conjugation. Plasmids often carry multiple antibiotic resistance genes, contributing to the spread of multidrug-resistance (MDR). Antibiotic resistance mediated by MDR plasmids severely limits the treatment options for the infections caused by Gram-negative bacteria, especially family Enterobacteriaceae. The global spread of MDR plasmids has been enhanced by selective pressure from antibiotic usage in human and veterinary medicine.
In cellular biology, the plasmid copy number is the number of copies of a given plasmid in a cell. To ensure survival and thus the continued propagation of the plasmid, they must regulate their copy number. If a plasmid has too high of a copy number, they may excessively burden their host by occupying too much cellular machinery and using too much energy. On the other hand, too low of a copy number may result in the plasmid not being present in all of their host's progeny. Plasmids may be either low, medium or high copy number plasmids; the regulation mechanisms between low and medium copy number plasmids are different. Low copy plasmids require either a partitioning system or a toxin-antitoxin pair such as CcdA/CcdB to ensure that each daughter receives the plasmid. For example, the F plasmid, which is the origin of BACs is a single copy plasmid with a partitioning system encoded in an operon right next to the plasmid origin. The partitioning system interacts with the septation apparatus to ensure that each daughter receives a copy of the plasmid. Many biotechnology applications utilize mutated plasmids that replicate to high copy number. For example, pBR322 is a medium copy number plasmid from which several high copy number cloning vectors have been derived by mutagenesis, such as the well known pUC series. This delivers the convenience of high plasmid DNA yields but the additional burden of the high copy number restricts the plasmid size. Larger high copy plasmids (>30kb) are disfavoured and also prone to size reduction through deletional mutagenesis.

Corynebacterium glutamicum is a Gram-positive, rod-shaped bacterium that is used industrially for large-scale production of amino acids. While originally identified in a screen for organisms secreting L-glutamate, mutants of C. glutamicum have also been identified that produce various other amino acids.

The bglG cis regulatory RNA was identified by RNA deep sequencing of Clostridioides difficile 630 where it is located immediately upstream of gene bglG2. BglG2 is an antiterminator protein involved in the regulation of genes of the Beta-glucoside phosphotransferase system in. Homologues of the C. difficile element were found across Clostridiales and other firmicutes. The element is consistently located 5’ of bglG2 or LicT, another antiterminator of the Beta-Glucoside Phosphotransferase System [2], consistent with a cis-regularly function at these operons. Only Clostridiales sequences were included in the seed alignment. Sequence is named S0591 in ref.
















