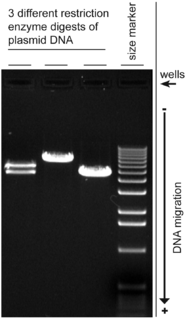
Agarose gel electrophoresis is a method of gel electrophoresis used in biochemistry, molecular biology, genetics, and clinical chemistry to separate a mixed population of macromolecules such as DNA or proteins in a matrix of agarose, one of the two main components of agar. The proteins may be separated by charge and/or size, and the DNA and RNA fragments by length. Biomolecules are separated by applying an electric field to move the charged molecules through an agarose matrix, and the biomolecules are separated by size in the agarose gel matrix.

Agarose is a polysaccharide, generally extracted from certain red seaweed. It is a linear polymer made up of the repeating unit of agarobiose, which is a disaccharide made up of D-galactose and 3,6-anhydro-L-galactopyranose. Agarose is one of the two principal components of agar, and is purified from agar by removing agar's other component, agaropectin.

Gel electrophoresis is a method for separation and analysis of macromolecules and their fragments, based on their size and charge. It is used in clinical chemistry to separate proteins by charge or size and in biochemistry and molecular biology to separate a mixed population of DNA and RNA fragments by length, to estimate the size of DNA and RNA fragments or to separate proteins by charge.
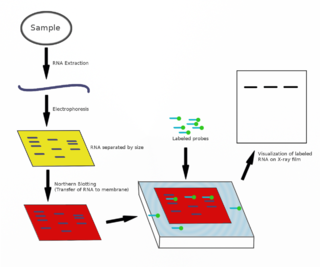
The northern blot, or RNA blot, is a technique used in molecular biology research to study gene expression by detection of RNA in a sample.
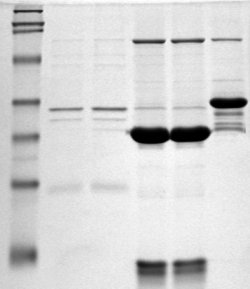
Polyacrylamide gel electrophoresis (PAGE) is a technique widely used in biochemistry, forensic chemistry, genetics, molecular biology and biotechnology to separate biological macromolecules, usually proteins or nucleic acids, according to their electrophoretic mobility. Electrophoretic mobility is a function of the length, conformation and charge of the molecule. Polyacrylamide gel electrophoresis is a powerful tool used to analyze RNA samples. When polyacrylamide gel is denatured after electrophoresis, it provides information on the sample composition of the RNA species.

Nucleic acid electrophoresis is an analytical technique used to separate DNA or RNA fragments by size and reactivity. Nucleic acid molecules which are to be analyzed are set upon a viscous medium, the gel, where an electric field induces the nucleic acids to migrate toward the anode. The separation of these fragments is accomplished by exploiting the mobilities with which different sized molecules are able to pass through the gel. Longer molecules migrate more slowly because they experience more resistance within the gel. Because the size of the molecule affects its mobility, smaller fragments end up nearer to the anode than longer ones in a given period. After some time, the voltage is removed and the fragmentation gradient is analyzed. For larger separations between similar sized fragments, either the voltage or run time can be increased. Extended runs across a low voltage gel yield the most accurate resolution. Voltage is, however, not the sole factor in determining electrophoresis of nucleic acids.

Ethidium bromide is an intercalating agent commonly used as a fluorescent tag in molecular biology laboratories for techniques such as agarose gel electrophoresis. It is commonly abbreviated as EtBr, which is also an abbreviation for bromoethane. To avoid confusion, some laboratories have used the abbreviation EthBr for this salt. When exposed to ultraviolet light, it will fluoresce with an orange colour, intensifying almost 20-fold after binding to DNA. Under the name homidium, it has been commonly used since the 1950s in veterinary medicine to treat trypanosomiasis in cattle. The high incidence of antimicrobial resistance makes this treatment impractical in some areas, where the related isometamidium chloride is used instead. Although ethidium bromide is said to be mutagenic on some circumstances of exposure, evidence shows that it is relatively safe and its alleged high toxicity is largely a myth.
Protein purification is a series of processes intended to isolate one or a few proteins from a complex mixture, usually cells, tissues or whole organisms. Protein purification is vital for the specification of the function, structure and interactions of the protein of interest. The purification process may separate the protein and non-protein parts of the mixture, and finally separate the desired protein from all other proteins. Separation of one protein from all others is typically the most laborious aspect of protein purification. Separation steps usually exploit differences in protein size, physico-chemical properties, binding affinity and biological activity. The pure result may be termed protein isolate.
The first isolation of deoxyribonucleic acid (DNA) was done in 1869 by Friedrich Miescher. Currently it is a routine procedure in molecular biology or forensic analyses. For the chemical method, there are many different kits used for extraction, and selecting the correct one will save time on kit optimization and extraction procedures. PCR sensitivity detection is considered to show the variation between the commercial kits.

Nile blue is a stain used in biology and histology. It may be used with live or fixed cells, and imparts a blue colour to cell nuclei.
A restriction digest is a procedure used in molecular biology to prepare DNA for analysis or other processing. It is sometimes termed DNA fragmentation. Hartl and Jones describe it this way:
This enzymatic technique can be used for cleaving DNA molecules at specific sites, ensuring that all DNA fragments that contain a particular sequence at a particular location have the same size; furthermore, each fragment that contains the desired sequence has the sequence located at exactly the same position within the fragment. The cleavage method makes use of an important class of DNA-cleaving enzymes isolated primarily from bacteria. These enzymes are called restriction endonucleases or restriction enzymes, and they are able to cleave DNA molecules at the positions at which particular short sequences of bases are present.
Immunoprecipitation (IP) is the technique of precipitating a protein antigen out of solution using an antibody that specifically binds to that particular protein. This process can be used to isolate and concentrate a particular protein from a sample containing many thousands of different proteins. Immunoprecipitation requires that the antibody be coupled to a solid substrate at some point in the procedure.

A molecular-weight size marker, also referred to as a protein ladder, DNA ladder, or RNA ladder, is a set of standards that are used to identify the approximate size of a molecule run on a gel during electrophoresis, using the principle that molecular weight is inversely proportional to migration rate through a gel matrix. Therefore, when used in gel electrophoresis, markers effectively provide a logarithmic scale by which to estimate the size of the other fragments.

SYBR Green I (SG) is an asymmetrical cyanine dye used as a nucleic acid stain in molecular biology. The SYBR family of dyes is produced by Molecular Probes Inc., now owned by Thermo Fisher Scientific. SYBR Green I binds to DNA. The resulting DNA-dye-complex best absorbs 497 nanometer blue light and emits green light. The stain preferentially binds to double-stranded DNA, but will stain single-stranded (ss) DNA with lower performance. SYBR Green can also stain RNA with a lower performance than ssDNA.

A gel doc, also known as a gel documentation system, gel image system or gel imager, refers to equipment widely used in molecular biology laboratories for the imaging and documentation of nucleic acid and protein suspended within polyacrylamide or agarose gels. These gels are typically stained with ethidium bromide or other nucleic acid stains such as GelGreen. Generally, a gel doc includes an ultraviolet (UV) light transilluminator, a hood or a darkroom to shield external light sources and protect the user from UV exposure, and a CCD camera for image capturing. With advancement in CMOS camera sensors like Sony's Pregius and Exmor series, low light capable cameras made of these sensors are also being incorporated in gel documentation systems

GelRed is an intercalating nucleic acid stain used in molecular genetics for agarose gel DNA electrophoresis. GelRed structurally consists of two ethidium subunits that are bridged by a linear oxygenated spacer.

GelGreen is an intercalating nucleic acid stain used in molecular genetics for agarose gel DNA electrophoresis. GelGreen consists of two acridine orange subunits that are bridged by a linear oxygenated spacer.
Boom method is a solid phase extraction method for isolating nucleic acid from a biological sample. This method is characterized by "absorbing the nucleic acids (NA) to the silica beads".

SYBR Safe is a cyanine dye used as a nucleic acid stain in molecular biology. SYBR Safe is one of a number of SYBR dyes made by the Life Technologies Corporation. SYBR Safe binds to DNA. The resulting DNA-dye-complex absorbs blue light and emits green light.

SYBR Gold is an asymmetrical cyanine dye. It can be used as a stain for double-stranded DNA, single-stranded DNA, and RNA. SYBR Gold is the most sensitive fluorescent stain of the SYBR family of dyes for the detection of nucleic acids. The SYBR family of dyes is produced by Molecular Probes Inc., now owned by Thermo Fisher Scientific













