Heat shock proteins (HSP) are a family of proteins that are produced by cells in response to exposure to stressful conditions. They were first described in relation to heat shock, but are now known to also be expressed during other stresses including exposure to cold, UV light and during wound healing or tissue remodeling. Many members of this group perform chaperone functions by stabilizing new proteins to ensure correct folding or by helping to refold proteins that were damaged by the cell stress. This increase in expression is transcriptionally regulated. The dramatic upregulation of the heat shock proteins is a key part of the heat shock response and is induced primarily by heat shock factor (HSF). HSPs are found in virtually all living organisms, from bacteria to humans.
The 70 kilodalton heat shock proteins are a family of conserved ubiquitously expressed heat shock proteins. Proteins with similar structure exist in virtually all living organisms. The Hsp70s are an important part of the cell's machinery for protein folding, and help to protect cells from stress.
An internal ribosome entry site, abbreviated IRES, is an RNA element that allows for translation initiation in cap-independent manner, as part of the greater process of protein synthesis. In eukaryotic translation, initiation typically occurs at the 5' end of mRNA molecules, since 5' cap recognition is required for the assembly of the initiation complex. The location for IRES elements is often in the 5'UTR, but can also occur elsewhere in mRNAs.
Ribosome shunting is a mechanism of translation initiation in which ribosomes bypass, or "shunt over", parts of the 5' untranslated region to reach the start codon, enabling viruses to have more information than usual in an mRNA molecule. Some viral RNAs have been shown to use ribosome shunting as a more efficient form of translation during certain stages of viral life cycle or when translation initiation factors are scarce. Some viruses known to use this mechanism include adenovirus, Sendai virus, human papillomavirus, duck hepatitis B pararetrovirus, rice tungro bacilliform viruses, and cauliflower mosaic virus. In these viruses the ribosome is directly translocated from the upstream initiation complex to the start codon (AUG) without the need to unwind RNA secondary structures.

The bag-1 internal ribosome entry site (IRES) is a cis-acting element located in the 5 ' untranslated region of the BAG-1 protein mRNA. Its effects apoptosis through IRES mediated translation of the BAG-1 protein.
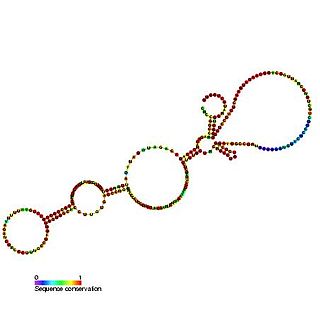

The BiP internal ribosome entry site (IRES) is an RNA element present in the 5' UTR of the mRNA of BiP protein and allows cap-independent translation. BiP protein expression has been found to be significantly enhanced by the heat shock response due to internal ribosome entry site (IRES)-dependent translation. It is thought that this translational mechanism is essential for the survival of cells under stress.

The Cripavirus internal ribosome entry site is an RNA element required for the production of capsid proteins through ribosome recruitment to an intergenic region IRES.
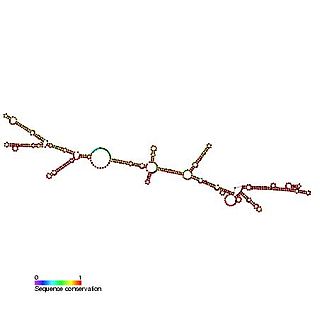
The c-sis internal ribosome entry site (IRES) is a RNA element found in the 5' UTR of the PDGF beta chain gene. The internal ribosome entry site contains three modules that can individually mediate internal ribosome entry. However, the full length sequence is required for maximal IRES activity. It is thought that the three IRES elements are somehow responsive to cellular changes and act to regulate the level of translation.
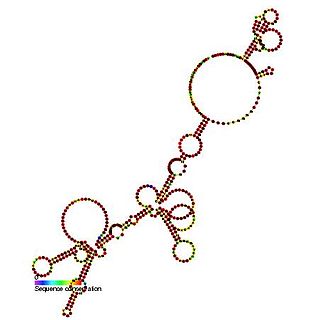
The FGF-2 internal ribosome entry site is an RNA element present in the 5' UTR of the mRNA of fibroblast growth factor-2. It has been found that the FGF-2 internal ribosome entry site (IRES) activity is strictly controlled and highly tissue specific. It is thought that translational IRES dependent activation of FGF-2 plays a vital role in embryogenesis and in the adult brain [1]. When expressed the fibroblast growth factor 2 FGF-2 protein plays a pivotal role in cell proliferation, differentiation and survival as well as being involved in wound-healing [1,2].

This family represents the internal ribosome entry site (IRES) of the hepatitis A virus. HAV IRES is a 450 nucleotide long sequence located in the 735 nt long 5’ UTR of Hepatitis A viral RNA genome. IRES elements allow cap and end-independent translation of mRNA in the host cell. The IRES achieves this by mediating the internal initiation of translation by recruiting a ribosomal 40S pre-initiation complex directly to the initiation codon and eliminates the requirement for eukaryotic initiation factor, eIF4F.
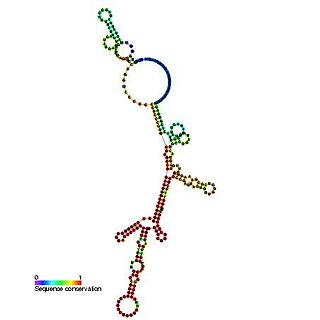
The Hepatitis C virus internal ribosome entry site, or HCV IRES, is an RNA structure within the 5'UTR of the HCV genome that mediates cap-independent translation initiation.

The L-myc internal ribosome entry site (IRES) is an RNA element present in the 5' UTR of the mRNA of L-myc that allows cap-independent translation. L-myc undergoes translation via the internal ribosome entry site and bypasses the typical eukaryotic cap-dependent translation pathway [1]. The myc family of genes when expressed are known to be involved in the control of cell growth, differentiation and apoptosis.
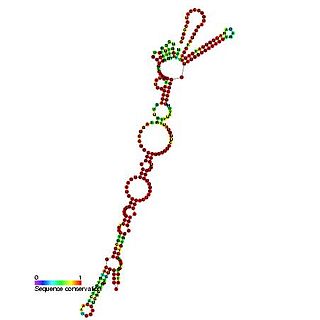

This family represents the Picornavirus internal ribosome entry site (IRES) element present in their 5' untranslated region. These elements were discovered in picornaviruses. They are cis-acting RNA sequences that adopt diverse three-dimensional structures, recruit the translation machinery and that often operate in association with specific RNA-binding proteins. IRES elements allow cap and end-independent translation of mRNA in the host cell. It has been found that La autoantigen (La) is required for Coxsackievirus B3 (CVB3) IRES-mediated translation, and it has been suggested that La may be required for the efficient translation of the viral RNA in the pancreas. Based on their secondary structure, picornavirus IRES are grouped into four main types. Type I comprises enteroand rhinovirus IRES and type II, those of cardio- and aphthovirus, among others. Type III is used to name hepatitis A IRES.

The repression of heat shock gene expression (ROSE) element is an RNA element found in the 5' UTR of some heat shock protein's mRNAs. The ROSE element is an RNA thermometer that negatively regulates heat shock gene expression. The secondary structure is thought to be altered by temperature, thus it is an RNA thermometer. This structure blocks access to the ribosome binding site at normal temperatures. During heat shock however, the structure changes freeing the ribosome binding site and allowing expression to occur.

The HIF-1α internal ribosome entry site (IRES) is an RNA element present in the 5' UTR of the mRNA of HIF-1α that allows cap-independent translation. The HIF-1α internal ribosome entry site (IRES) allows translation to be maintained under hypoxic cell conditions that inhibit cap-dependent translation [1]. The hypoxia-inducible factor-1α protein (HIF-1α) is a subunit of the HIF-1 transcription factor, which induces transcription of several genes involved in the cellular response to hypoxia.
A ribosome binding site, or ribosomal binding site (RBS), is a sequence of nucleotides upstream of the start codon of an mRNA transcript that is responsible for the recruitment of a ribosome during the initiation of translation. Mostly, RBS refers to bacterial sequences, although internal ribosome entry sites (IRES) have been described in mRNAs of eukaryotic cells or viruses that infect eukaryotes. Ribosome recruitment in eukaryotes is generally mediated by the 5' cap present on eukaryotic mRNAs.

FourU thermometers are a class of non-coding RNA thermometers found in Salmonella. They are named 'FourU' due to the four highly conserved uridine nucleotides found directly opposite the Shine-Dalgarno sequence on hairpin II (pictured). RNA thermometers such as FourU control regulation of temperature via heat shock proteins in many prokaryotes. FourU thermometers are relatively small RNA molecules, only 57 nucleotides in length, and have a simple two-hairpin structure.

An RNA thermometer is a temperature-sensitive non-coding RNA molecule which regulates gene expression. RNA thermometers often regulate genes required during either a heat shock or cold shock response, but have been implicated in other regulatory roles such as in pathogenicity and starvation.
In molecular biology, the ODC internal ribosome entry site (IRES) is an RNA element present in the 5′ UTR of the mRNA encoding ornithine decarboxylase. It has been suggested that this IRES allows cap-independent translation of ornithine decarboxylase at the G2/M phase of the cell cycle, however there is some doubt about this. Translation from this IRES is activated by the zinc finger protein ZNF9 and by Poly(rC)-binding protein 2 (PCBP2). It is also activated in Ras-transformed cells.













