
Adenosine triphosphate (ATP) is a nucleotide that provides energy to drive and support many processes in living cells, such as muscle contraction, nerve impulse propagation, condensate dissolution, and chemical synthesis. Found in all known forms of life, it is often referred to as the "molecular unit of currency" of intracellular energy transfer.

The citric acid cycle—also known as the Krebs cycle, Szent–Györgyi–Krebs cycle or the TCA cycle (tricarboxylic acid cycle)—is a series of biochemical reactions to release the energy stored in nutrients through the oxidation of acetyl-CoA derived from carbohydrates, fats, and proteins. The chemical energy released is available under the form of ATP. The Krebs cycle is used by organisms that respire (as opposed to organisms that ferment) to generate energy, either by anaerobic respiration or aerobic respiration. In addition, the cycle provides precursors of certain amino acids, as well as the reducing agent NADH, that are used in numerous other reactions. Its central importance to many biochemical pathways suggests that it was one of the earliest components of metabolism. Even though it is branded as a "cycle", it is not necessary for metabolites to follow only one specific route; at least three alternative segments of the citric acid cycle have been recognized.

Oxidative phosphorylation or electron transport-linked phosphorylation or terminal oxidation is the metabolic pathway in which cells use enzymes to oxidize nutrients, thereby releasing chemical energy in order to produce adenosine triphosphate (ATP). In eukaryotes, this takes place inside mitochondria. Almost all aerobic organisms carry out oxidative phosphorylation. This pathway is so pervasive because it releases more energy than alternative fermentation processes such as anaerobic glycolysis.
An electron transport chain (ETC) is a series of protein complexes and other molecules that transfer electrons from electron donors to electron acceptors via redox reactions (both reduction and oxidation occurring simultaneously) and couples this electron transfer with the transfer of protons (H+ ions) across a membrane. Many of the enzymes in the electron transport chain are embedded within the membrane.
A proton pump is an integral membrane protein pump that builds up a proton gradient across a biological membrane. Proton pumps catalyze the following reaction:

The enzyme cytochrome c oxidase or Complex IV, is a large transmembrane protein complex found in bacteria, archaea, and the mitochondria of eukaryotes.

The coenzyme Q : cytochrome c – oxidoreductase, sometimes called the cytochrome bc1 complex, and at other times complex III, is the third complex in the electron transport chain, playing a critical role in biochemical generation of ATP. Complex III is a multisubunit transmembrane protein encoded by both the mitochondrial and the nuclear genomes. Complex III is present in the mitochondria of all animals and all aerobic eukaryotes and the inner membranes of most eubacteria. Mutations in Complex III cause exercise intolerance as well as multisystem disorders. The bc1 complex contains 11 subunits, 3 respiratory subunits, 2 core proteins and 6 low-molecular weight proteins.

Cellular respiration is the process by which biological fuels are oxidized in the presence of an inorganic electron acceptor, such as oxygen, to drive the bulk production of adenosine triphosphate (ATP), which contains energy. Cellular respiration may be described as a set of metabolic reactions and processes that take place in the cells of organisms to convert chemical energy from nutrients into ATP, and then release waste products.

ATP synthase is an enzyme that catalyzes the formation of the energy storage molecule adenosine triphosphate (ATP) using adenosine diphosphate (ADP) and inorganic phosphate (Pi). ATP synthase is a molecular machine. The overall reaction catalyzed by ATP synthase is:
In molecular biology, housekeeping genes are typically constitutive genes that are required for the maintenance of basic cellular function, and are expressed in all cells of an organism under normal and patho-physiological conditions. Although some housekeeping genes are expressed at relatively constant rates in most non-pathological situations, the expression of other housekeeping genes may vary depending on experimental conditions.
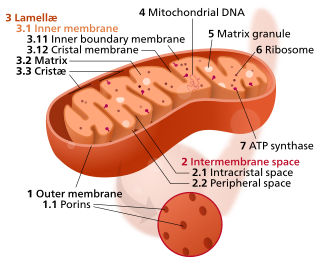
In the mitochondrion, the matrix is the space within the inner membrane. The word "matrix" stems from the fact that this space is viscous, compared to the relatively aqueous cytoplasm. The mitochondrial matrix contains the mitochondrial DNA, ribosomes, soluble enzymes, small organic molecules, nucleotide cofactors, and inorganic ions.[1] The enzymes in the matrix facilitate reactions responsible for the production of ATP, such as the citric acid cycle, oxidative phosphorylation, oxidation of pyruvate, and the beta oxidation of fatty acids.
Substrate-level phosphorylation is a metabolism reaction that results in the production of ATP or GTP supported by the energy released from another high-energy bond that leads to phosphorylation of ADP or GDP to ATP or GTP (note that the reaction catalyzed by creatine kinase is not considered as "substrate-level phosphorylation"). This process uses some of the released chemical energy, the Gibbs free energy, to transfer a phosphoryl (PO3) group to ADP or GDP. Occurs in glycolysis and in the citric acid cycle.

The enzyme citrate synthase E.C. 2.3.3.1 ] exists in nearly all living cells and stands as a pace-making enzyme in the first step of the citric acid cycle. Citrate synthase is localized within eukaryotic cells in the mitochondrial matrix, but is encoded by nuclear DNA rather than mitochondrial. It is synthesized using cytoplasmic ribosomes, then transported into the mitochondrial matrix.

Bongkrek acid is a respiratory toxin produced in fermented coconut or corn contaminated by the bacterium Burkholderia gladioli pathovar cocovenenans. It is a highly toxic, heat-stable, colorless, odorless, and highly unsaturated tricarboxylic acid that inhibits the ADP/ATP translocase, also called the mitochondrial ADP/ATP carrier, preventing ATP from leaving the mitochondria to provide metabolic energy to the rest of the cell. Bongkrek acid, when consumed through contaminated foods, mainly targets the liver, brain, and kidneys along with symptoms that include vomiting, diarrhea, urinary retention, abdominal pain, and excessive sweating. Most of the outbreaks are found in Indonesia and China where fermented coconut and corn-based foods are consumed.

Succinyl coenzyme A synthetase is an enzyme that catalyzes the reversible reaction of succinyl-CoA to succinate. The enzyme facilitates the coupling of this reaction to the formation of a nucleoside triphosphate molecule from an inorganic phosphate molecule and a nucleoside diphosphate molecule. It plays a key role as one of the catalysts involved in the citric acid cycle, a central pathway in cellular metabolism, and it is located within the mitochondrial matrix of a cell.
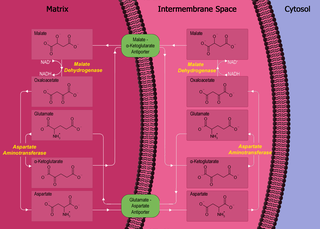
The malate–aspartate shuttle is a biochemical system for translocating electrons produced during glycolysis across the semipermeable inner membrane of the mitochondrion for oxidative phosphorylation in eukaryotes. These electrons enter the electron transport chain of the mitochondria via reduction equivalents to generate ATP. The shuttle system is required because the mitochondrial inner membrane is impermeable to NADH, the primary reducing equivalent of the electron transport chain. To circumvent this, malate carries the reducing equivalents across the membrane.

Adenine nucleotide translocator (ANT), also known as the ADP/ATP translocase (ANT), ADP/ATP carrier protein (AAC) or mitochondrial ADP/ATP carrier, exchanges free ATP with free ADP across the inner mitochondrial membrane. ANT is the most abundant protein in the inner mitochondrial membrane and belongs to mitochondrial carrier family.
Acetyl-CoA synthetase (ACS) or Acetate—CoA ligase is an enzyme involved in metabolism of acetate. It is in the ligase class of enzymes, meaning that it catalyzes the formation of a new chemical bond between two large molecules.
In enzymology, a threonine-tRNA ligase is an enzyme that catalyzes the chemical reaction

NAD-dependent deacetylase sirtuin-3, mitochondrial also known as SIRT3 is a protein that in humans is encoded by the SIRT3 gene [sirtuin 3 ]. SIRT3 is member of the mammalian sirtuin family of proteins, which are homologs to the yeast Sir2 protein. SIRT3 exhibits NAD+-dependent deacetylase activity.













