A transposable element is a nucleic acid sequence in DNA that can change its position within a genome, sometimes creating or reversing mutations and altering the cell's genetic identity and genome size. Transposition often results in duplication of the same genetic material. In the human genome, L1 and Alu elements are two examples. Barbara McClintock's discovery of them earned her a Nobel Prize in 1983. Its importance in personalized medicine is becoming increasingly relevant, as well as gaining more attention in data analytics given the difficulty of analysis in very high dimensional spaces.

A frameshift mutation is a genetic mutation caused by indels of a number of nucleotides in a DNA sequence that is not divisible by three. Due to the triplet nature of gene expression by codons, the insertion or deletion can change the reading frame, resulting in a completely different translation from the original. The earlier in the sequence the deletion or insertion occurs, the more altered the protein. A frameshift mutation is not the same as a single-nucleotide polymorphism in which a nucleotide is replaced, rather than inserted or deleted. A frameshift mutation will in general cause the reading of the codons after the mutation to code for different amino acids. The frameshift mutation will also alter the first stop codon encountered in the sequence. The polypeptide being created could be abnormally short or abnormally long, and will most likely not be functional.

Retrotransposons are a type of genetic component that copy and paste themselves into different genomic locations (transposon) by converting RNA back into DNA through the reverse transcription process using an RNA transposition intermediate.
A transposase is any of a class of enzymes capable of binding to the end of a transposon and catalysing its movement to another part of a genome, typically by a cut-and-paste mechanism or a replicative mechanism, in a process known as transposition. The word "transposase" was first coined by the individuals who cloned the enzyme required for transposition of the Tn3 transposon. The existence of transposons was postulated in the late 1940s by Barbara McClintock, who was studying the inheritance of maize, but the actual molecular basis for transposition was described by later groups. McClintock discovered that some segments of chromosomes changed their position, jumping between different loci or from one chromosome to another. The repositioning of these transposons allowed other genes for pigment to be expressed. Transposition in maize causes changes in color; however, in other organisms, such as bacteria, it can cause antibiotic resistance. Transposition is also important in creating genetic diversity within species and generating adaptability to changing living conditions.
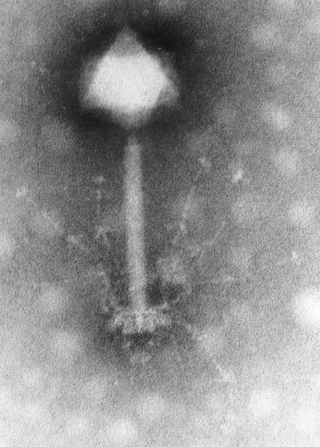
Myoviridae is a family of bacteriophages in the order Caudovirales. Bacteria and archaea serve as natural hosts. There are 625 species in this family, assigned to eight subfamilies and 217 genera.

In genetics, an insertion is the addition of one or more nucleotide base pairs into a DNA sequence. This can often happen in microsatellite regions due to the DNA polymerase slipping. Insertions can be anywhere in size from one base pair incorrectly inserted into a DNA sequence to a section of one chromosome inserted into another. The mechanism of the smallest single base insertion mutations is believed to be through base-pair separation between the template and primer strands followed by non-neighbor base stacking, which can occur locally within the DNA polymerase active site. On a chromosome level, an insertion refers to the insertion of a larger sequence into a chromosome. This can happen due to unequal crossover during meiosis.
Indel (insertion-deletion) is a molecular biology term for an insertion or deletion of bases in the genome of an organism. Indels ≥ 50 bases in length are classified as structural variants.
P elements are transposable elements that were discovered in Drosophila as the causative agents of genetic traits called hybrid dysgenesis. The transposon is responsible for the P trait of the P element and it is found only in wild flies. They are also found in many other eukaryotes.

Insertion element is a short DNA sequence that acts as a simple transposable element. Insertion sequences have two major characteristics: they are small relative to other transposable elements and only code for proteins implicated in the transposition activity. These proteins are usually the transposase which catalyses the enzymatic reaction allowing the IS to move, and also one regulatory protein which either stimulates or inhibits the transposition activity. The coding region in an insertion sequence is usually flanked by inverted repeats. For example, the well-known IS911 is flanked by two 36bp inverted repeat extremities and the coding region has two genes partially overlapping orfA and orfAB, coding the transposase (OrfAB) and a regulatory protein (OrfA). A particular insertion sequence may be named according to the form ISn, where n is a number ; this is not the only naming scheme used, however. Although insertion sequences are usually discussed in the context of prokaryotic genomes, certain eukaryotic DNA sequences belonging to the family of Tc1/mariner transposable elements may be considered to be, insertion sequences.

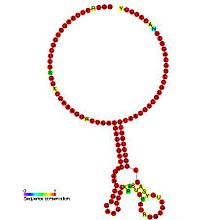
In molecular biology, the coronavirus frameshifting stimulation element is a conserved stem-loop of RNA found in coronaviruses that can promote ribosomal frameshifting. Such RNA molecules interact with a downstream region to form a pseudoknot structure; the region varies according to the virus but pseudoknot formation is known to stimulate frameshifting. In the classical situation, a sequence 32 nucleotides downstream of the stem is complementary to part of the loop. In other coronaviruses, however, another stem-loop structure around 150 nucleotides downstream can interact with members of this family to form kissing stem-loops and stimulate frameshifting.

The DnaX ribosomal frameshifting element is a RNA element found in the mRNA of the dnaX gene in E. coli. The dnaX gene has two encoded products, tau and gamma, which are produced in a 1:1 ratio. The gamma protein is synthesised due to programmed frameshifting and is shorter than tau. The two products of the dnaX gene are DNA polymerase III subunits.

HIV ribosomal frameshift signal is a ribosomal frameshift (PRF) that human immunodeficiency virus (HIV) uses to translate several different proteins from the same sequence.
Ribosomal frameshifting, also known as translational frameshifting or translational recoding, is a biological phenomenon that occurs during translation that results in the production of multiple, unique proteins from a single mRNA. The process can be programmed by the nucleotide sequence of the mRNA and is sometimes affected by the secondary, 3-dimensional mRNA structure. It has been described mainly in viruses, retrotransposons and bacterial insertion elements, and also in some cellular genes.

ALIL pseudoknot is an RNA element that induces frameshifting in bacteria. The expression of a minority of genes requires frameshifting to occur where the frequency of frameshifting is increased by a RNA secondary structure located on the 3' side of the shift site. This structure can be either a pseudoknot or a stem-loop and acts as a physical barrier to mRNA translocation so therefore causes ribosome pausing.
Transposons are semi-parasitic DNA sequences which can replicate and spread through the host's genome. They can be harnessed as a genetic tool for analysis of gene and protein function. The use of transposons is well-developed in Drosophila and in Thale cress and bacteria such as Escherichia coli.

Ribosomal pause refers to the queueing or stacking of ribosomes during translation of the nucleotide sequence of mRNA transcripts. These transcripts are decoded and converted into an amino acid sequence during protein synthesis by ribosomes. Due to the pause sites of some mRNA's, there is a disturbance caused in translation. Ribosomal pausing occurs in both eukaryotes and prokaryotes. A more severe pause is known as a ribosomal stall.

A slippery sequence is a small section of codon nucleotide sequences that controls the rate and chance of ribosomal frameshifting. A slippery sequence causes a faster ribosomal transfer which in turn can cause the reading ribosome to "slip." This allows a tRNA to shift by 1 base (−1) after it has paired with its anticodon, changing the reading frame. A −1 frameshift triggered by such a sequence is a Programmed −1 Ribosomal Frameshift. It is followed by a spacer region, and an RNA secondary structure. Such sequences are common in virus polyproteins.
Ac/Ds transposable controlling elements was the first transposable element system recognized in maize. The Ac Activator element is autonomous, whereas the Ds Dissociation element requires an Activator element to transpose. Ac was initially discovered as enabling a Ds element to break chromosomes. Both Ac and Ds can also insert into genes, causing mutants that may revert to normal on excision of the element. The phenotypic consequence of Ac/Ds transposable element includes mosaic colors in kernels and leaves in maize.
DNA transposons are DNA sequences, sometimes referred to "jumping genes", that can move and integrate to different locations within the genome. They are class II transposable elements (TEs) that move through a DNA intermediate, as opposed to class I TEs, retrotransposons, that move through an RNA intermediate. DNA transposons can move in the DNA of an organism via a single-or double-stranded DNA intermediate. DNA transposons have been found in both prokaryotic and eukaryotic organisms. They can make up a significant portion of an organism's genome, particularly in eukaryotes. In prokaryotes, TE's can facilitate the horizontal transfer of antibiotic resistance or other genes associated with virulence. After replicating and propagating in a host, all transposon copies become inactivated and are lost unless the transposon passes to a genome by starting a new life cycle with horizontal transfer. It is important to note that DNA transposons do not randomly insert themselves into the genome, but rather show preference for specific sites.
Metavirus is a genus of viruses in the family Metaviridae. They are retrotransposons that invade a eukaryotic host genome and may only replicate once the virus has infected the host. These genetic elements exist to infect and replicate in their host genome and are derived from ancestral elements unrelated from their host. Metavirus may use several different hosts for transmission, and has been found to be transmissible through ovule and pollen of some plants.