Related Research Articles

A reverse transcriptase (RT) is an enzyme used to generate complementary DNA (cDNA) from an RNA template, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryotic cells to extend the telomeres at the ends of their linear chromosomes. Contrary to a widely held belief, the process does not violate the flows of genetic information as described by the classical central dogma, as transfers of information from RNA to DNA are explicitly held possible.
DNA topoisomerases are enzymes that catalyze changes in the topological state of DNA, interconverting relaxed and supercoiled forms, linked (catenated) and unlinked species, and knotted and unknotted DNA. Topological issues in DNA arise due to the intertwined nature of its double-helical structure, which, for example, can lead to overwinding of the DNA duplex during DNA replication and transcription. If left unchanged, this torsion would eventually stop the DNA or RNA polymerases involved in these processes from continuing along the DNA helix. A second topological challenge results from the linking or tangling of DNA during replication. Left unresolved, links between replicated DNA will impede cell division. The DNA topoisomerases prevent and correct these types of topological problems. They do this by binding to DNA and cutting the sugar-phosphate backbone of either one or both of the DNA strands. This transient break allows the DNA to be untangled or unwound, and, at the end of these processes, the DNA backbone is resealed. Since the overall chemical composition and connectivity of the DNA do not change, the DNA substrate and product are chemical isomers, differing only in their topology.

Ribonuclease is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within the EC 2.7 and 3.1 classes of enzymes.

Ribonuclease H is a family of non-sequence-specific endonuclease enzymes that catalyze the cleavage of RNA in an RNA/DNA substrate via a hydrolytic mechanism. Members of the RNase H family can be found in nearly all organisms, from bacteria to archaea to eukaryotes.

The nucleoid is an irregularly shaped region within the prokaryotic cell that contains all or most of the genetic material. The chromosome of a prokaryote is circular, and its length is very large compared to the cell dimensions, so it needs to be compacted in order to fit. In contrast to the nucleus of a eukaryotic cell, it is not surrounded by a nuclear membrane. Instead, the nucleoid forms by condensation and functional arrangement with the help of chromosomal architectural proteins and RNA molecules as well as DNA supercoiling. The length of a genome widely varies and a cell may contain multiple copies of it.
A nick is a discontinuity in a double stranded DNA molecule where there is no phosphodiester bond between adjacent nucleotides of one strand typically through damage or enzyme action. Nicks allow DNA strands to untwist during replication, and are also thought to play a role in the DNA mismatch repair mechanisms that fix errors on both the leading and lagging daughter strands.

Topotecan, sold under the brand name Hycamtin among others, is a chemotherapeutic agent medication that is a topoisomerase inhibitor. It is a synthetic, water-soluble analog of the natural chemical compound camptothecin. It is used in the form of its hydrochloride salt to treat ovarian cancer, lung cancer and other cancer types.
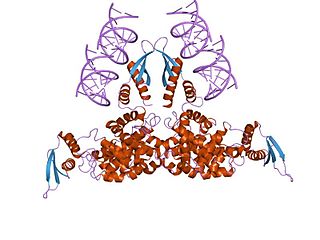
Ribonuclease III (BRENDA 3.1.26.3) is a type of ribonuclease that recognizes dsRNA and cleaves it at specific targeted locations to transform them into mature RNAs. These enzymes are a group of endoribonucleases that are characterized by their ribonuclease domain, which is labelled the RNase III domain. They are ubiquitous compounds in the cell and play a major role in pathways such as RNA precursor synthesis, RNA Silencing, and the pnp autoregulatory mechanism.
Topoisomerase inhibitors are chemical compounds that block the action of topoisomerases, which are broken into two broad subtypes: type I topoisomerases (TopI) and type II topoisomerases (TopII). Topoisomerase plays important roles in cellular reproduction and DNA organization, as they mediate the cleavage of single and double stranded DNA to relax supercoils, untangle catenanes, and condense chromosomes in eukaryotic cells. Topoisomerase inhibitors influence these essential cellular processes. Some topoisomerase inhibitors prevent topoisomerases from performing DNA strand breaks while others, deemed topoisomerase poisons, associate with topoisomerase-DNA complexes and prevent the re-ligation step of the topoisomerase mechanism. These topoisomerase-DNA-inhibitor complexes are cytotoxic agents, as the un-repaired single- and double stranded DNA breaks they cause can lead to apoptosis and cell death. Because of this ability to induce apoptosis, topoisomerase inhibitors have gained interest as therapeutics against infectious and cancerous cells.
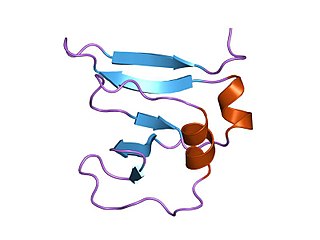
In molecular biology Type I topoisomerases are enzymes that cut one of the two strands of double-stranded DNA, relax the strand, and reanneal the strand. They are further subdivided into two structurally and mechanistically distinct topoisomerases: type IA and type IB.

Type II topoisomerases are topoisomerases that cut both strands of the DNA helix simultaneously in order to manage DNA tangles and supercoils. They use the hydrolysis of ATP, unlike Type I topoisomerase. In this process, these enzymes change the linking number of circular DNA by ±2. Topoisomerases are ubiquitous enzymes, found in all living organisms.
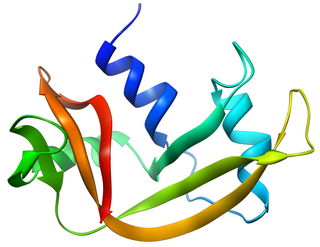
Pancreatic ribonuclease family is a superfamily of pyrimidine-specific endonucleases found in high quantity in the pancreas of certain mammals and of some reptiles.

DNA topoisomerase 1 is an enzyme that in humans is encoded by the TOP1 gene. It is a DNA topoisomerase, an enzyme that catalyzes the transient breaking and rejoining of a single strand of DNA.
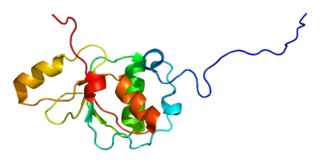
DNA topoisomerase 2-binding protein 1 is an enzyme that in humans is encoded by the TOPBP1 gene.

Tyrosyl-DNA phosphodiesterase 1 is an enzyme that in humans is encoded by the TDP1 gene.
Stephen Levene is an American biophysicist and professor of bioengineering, molecular biology, and physics at the University of Texas at Dallas.
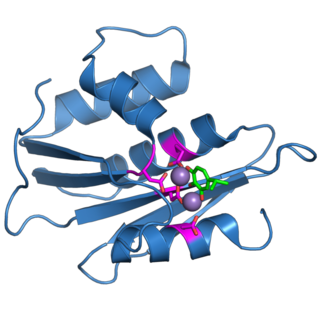
The retroviral ribonuclease H is a catalytic domain of the retroviral reverse transcriptase (RT) enzyme. The RT enzyme is used to generate complementary DNA (cDNA) from the retroviral RNA genome. This process is called reverse transcription. To complete this complex process, the retroviral RT enzymes need to adopt a multifunctional nature. They therefore possess 3 of the following biochemical activities: RNA-dependent DNA polymerase, ribonuclease H, and DNA-dependent DNA polymerase activities. Like all RNase H enzymes, the retroviral RNase H domain cleaves DNA/RNA duplexes and will not degrade DNA or unhybridized RNA.

A topologically associating domain (TAD) is a self-interacting genomic region, meaning that DNA sequences within a TAD physically interact with each other more frequently than with sequences outside the TAD. The median size of a TAD in mouse cells is 880 kb, and they have similar sizes in non-mammalian species. Boundaries at both side of these domains are conserved between different mammalian cell types and even across species and are highly enriched with CCCTC-binding factor (CTCF) and cohesin binding sites. In addition, some types of genes appear near TAD boundaries more often than would be expected by chance.
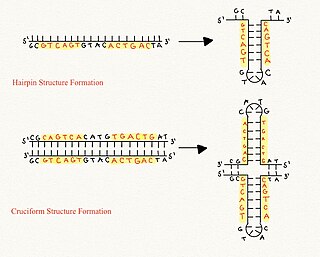
Cruciform DNA is a form of non-B DNA, or an alternative DNA structure. The formation of cruciform DNA requires the presence of palindromes called inverted repeat sequences. These inverted repeats contain a sequence of DNA in one strand that is repeated in the opposite direction on the other strand. As a result, inverted repeats are self-complementary and can give rise to structures such as hairpins and cruciforms. Cruciform DNA structures require at least a six nucleotide sequence of inverted repeats to form a structure consisting of a stem, branch point and loop in the shape of a cruciform, stabilized by negative DNA supercoiling.
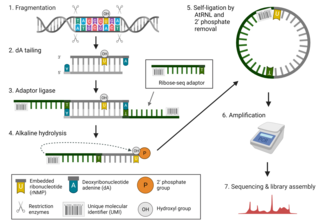
Ribose-seq is a mapping technique used in genetics research to determine the full profile of embedded ribonucleotides, specifically ribonucleoside monophosphates (rNMPs), in genomic DNA. Embedded ribonucleotides are thought to be the most common alteration to DNA in cells, and their presence in genomic DNA can affect genome stability. As recent studies have suggested that ribonucleotides in mouse DNA may affect disease pathology, ribonucleotide incorporation in genomic DNA has become an important target of medical genetics research. Ribose-seq allows scientists to determine the precise location and type of ribonucleotides that have been incorporated into eukaryotic or prokaryotic DNA.
References
- ↑ "Remembering Dr. James Champoux". University of Washington Honors Program. 22 May 2019. Retrieved 26 May 2019.
- ↑ "James Champoux Lab". washington.edu. Retrieved December 22, 2017.
- ↑ "AAAS Names 8 UW Researchers as Fellows in 2017". washington.edu. November 22, 2017. Retrieved December 22, 2017.
- 1 2 "James J. Champoux". washington.edu. Retrieved December 22, 2017.
- ↑ James J. Champoux. DNA Topoisomerases: Structure, Function, and Mechanism, Vol. 70:369-413 (Volume publication date July 2001). Annual review of Biochemistry
- ↑ "James Champoux". scholar.google.com. Retrieved February 24, 2018.