Related Research Articles

Metabolomics is the scientific study of chemical processes involving metabolites, the small molecule substrates, intermediates, and products of cell metabolism. Specifically, metabolomics is the "systematic study of the unique chemical fingerprints that specific cellular processes leave behind", the study of their small-molecule metabolite profiles. The metabolome represents the complete set of metabolites in a biological cell, tissue, organ, or organism, which are the end products of cellular processes. Messenger RNA (mRNA), gene expression data, and proteomic analyses reveal the set of gene products being produced in the cell, data that represents one aspect of cellular function. Conversely, metabolic profiling can give an instantaneous snapshot of the physiology of that cell, and thus, metabolomics provides a direct "functional readout of the physiological state" of an organism. There are indeed quantifiable correlations between the metabolome and the other cellular ensembles, which can be used to predict metabolite abundances in biological samples from, for example mRNA abundances. One of the ultimate challenges of systems biology is to integrate metabolomics with all other -omics information to provide a better understanding of cellular biology.

The metabolome refers to the complete set of small-molecule chemicals found within a biological sample. The biological sample can be a cell, a cellular organelle, an organ, a tissue, a tissue extract, a biofluid or an entire organism. The small molecule chemicals found in a given metabolome may include both endogenous metabolites that are naturally produced by an organism as well as exogenous chemicals that are not naturally produced by an organism.
The DrugBank database is a comprehensive, freely accessible, online database containing information on drugs and drug targets created and maintained by the University of Alberta and The Metabolomics Innovation Centre located in Alberta, Canada. As both a bioinformatics and a cheminformatics resource, DrugBank combines detailed drug data with comprehensive drug target information. DrugBank has used content from Wikipedia; Wikipedia also often links to Drugbank, posing potential circular reporting issues.

KEGG is a collection of databases dealing with genomes, biological pathways, diseases, drugs, and chemical substances. KEGG is utilized for bioinformatics research and education, including data analysis in genomics, metagenomics, metabolomics and other omics studies, modeling and simulation in systems biology, and translational research in drug development.
TRANSFAC is a manually curated database of eukaryotic transcription factors, their genomic binding sites and DNA binding profiles. The contents of the database can be used to predict potential transcription factor binding sites.

The Human Metabolome Database (HMDB) is a comprehensive, high-quality, freely accessible, online database of small molecule metabolites found in the human body. It has been created by the Human Metabolome Project funded by Genome Canada and is one of the first dedicated metabolomics databases. The HMDB facilitates human metabolomics research, including the identification and characterization of human metabolites using NMR spectroscopy, GC-MS spectrometry and LC/MS spectrometry. To aid in this discovery process, the HMDB contains three kinds of data: 1) chemical data, 2) clinical data, and 3) molecular biology/biochemistry data (Fig. 1–3). The chemical data includes 41,514 metabolite structures with detailed descriptions along with nearly 10,000 NMR, GC-MS and LC/MS spectra.

The Toxin and Toxin-Target Database (T3DB), also known as the Toxic Exposome Database, is a freely accessible online database of common substances that are toxic to humans, along with their protein, DNA or organ targets. The database currently houses nearly 3,700 toxic compounds or poisons described by nearly 42,000 synonyms. This list includes various groups of toxins, including common pollutants, pesticides, drugs, food toxins, household and industrial/workplace toxins, cigarette toxins, and uremic toxins. These toxic substances are linked to 2,086 corresponding protein/DNA target records. In total there are 42,433 toxic substance-toxin target associations. Each toxic compound record (ToxCard) in T3DB contains nearly 100 data fields and holds information such as chemical properties and descriptors, mechanisms of action, toxicity or lethal dose values, molecular and cellular interactions, medical information, NMR an MS spectra, and up- and down-regulated genes. This information has been extracted from over 18,000 sources, which include other databases, government documents, books, and scientific literature.
The Small Molecule Pathway Database (SMPDB) is a comprehensive, high-quality, freely accessible, online database containing more than 600 small molecule (i.e. metabolic) pathways found in humans. SMPDB is designed specifically to support pathway elucidation and pathway discovery in metabolomics, transcriptomics, proteomics and systems biology. It is able to do so, in part, by providing colorful, detailed, fully searchable, hyperlinked diagrams of five types of small molecule pathways: 1) general human metabolic pathways; 2) human metabolic disease pathways; 3) human metabolite signaling pathways; 4) drug-action pathways and 5) drug metabolism pathways. SMPDB pathways may be navigated, viewed and zoomed interactively using a Google Maps-like interface. All SMPDB pathways include information on the relevant organs, subcellular compartments, protein cofactors, protein locations, metabolite locations, chemical structures and protein quaternary structures (Fig. 1). Each small molecule in SMPDB is hyperlinked to detailed descriptions contained in the HMDB or DrugBank and each protein or enzyme complex is hyperlinked to UniProt. Additionally, all SMPDB pathways are accompanied with detailed descriptions and references, providing an overview of the pathway, condition or processes depicted in each diagram. Users can browse the SMPDB (Fig. 2) or search its contents by text searching (Fig. 3), sequence searching, or chemical structure searching. More powerful queries are also possible including searching with lists of gene or protein names, drug names, metabolite names, GenBank IDs, Swiss-Prot IDs, Agilent or Affymetrix microarray IDs. These queries will produce lists of matching pathways and highlight the matching molecules on each of the pathway diagrams. Gene, metabolite and protein concentration data can also be visualized through SMPDB's mapping interface.
MetaboAnalyst is a set of online tools for metabolomic data analysis and interpretation, created by members of the Wishart Research Group at the University of Alberta. It was first released in May 2009 and version 2.0 was released in January 2012. MetaboAnalyst provides a variety of analysis methods that have been tailored for metabolomic data. These methods include metabolomic data processing, normalization, multivariate statistical analysis, and data annotation. The current version is focused on biomarker discovery and classification.
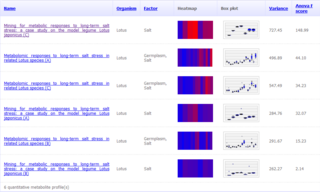
The Golm Metabolome Database (GMD) is a gas chromatography (GC) – mass spectrometry (MS) reference library dedicated to metabolite profiling experiments and comprises mass spectral and retention index (RI) information for non-annotated mass spectral tags together with data of a multitude of already identified metabolites and reference substances. The GMD is hosted at the Max Planck Institute of Molecular Plant Physiology in Golm district of Potsdam, Germany.
The Yeast Metabolome Database (YMDB) is a comprehensive, high-quality, freely accessible, online database of small molecule metabolites found in or produced by Saccharomyces cerevisiae. The YMDB was designed to facilitate yeast metabolomics research, specifically in the areas of general fermentation as well as wine, beer and fermented food analysis. YMDB supports the identification and characterization of yeast metabolites using NMR spectroscopy, GC-MS spectrometry and Liquid chromatography–mass spectrometry. The YMDB contains two kinds of data: 1) chemical data and 2) molecular biology/biochemistry data. The chemical data includes 2027 metabolite structures with detailed metabolite descriptions along with nearly 4000 NMR, GC-MS and LC/MS spectra.
Metabolomic Pathway Analysis, shortened to MetPA, is a freely available, user-friendly web server to assist with the identification analysis and visualization of metabolic pathways using metabolomic data. MetPA makes use of advances originally developed for pathway analysis in microarray experiments and applies those principles and concepts to the analysis of metabolic pathways. For input, MetPA expects either a list of compound names or a metabolite concentration table with phenotypic labels. The list of compounds can include common names, HMDB IDs or KEGG IDs with one compound per row. Compound concentration tables must have samples in rows and compounds in columns. MetPA's output is a series of tables indicating which pathways are significantly enriched as well as a variety of graphs or pathway maps illustrating where and how certain pathways were enriched. MetPA's graphical output uses a colorful Google-Maps visualization system that allows simple, intuitive data exploration that lets users employ a computer mouse or track pad to select, drag and place images and to seamlessly zoom in and out. Users can explore MetPA's output using three different views or levels: 1) a metabolome view; 2) a pathway view; 3) a compound view.
FooDB is a freely available, open-access database containing chemical composition data on common, unprocessed foods. It also contains extensive data on flavour and aroma constituents, food additives as well as positive and negative health effects associated with food constituents. The database contains information on more than 28,000 chemicals found in more than 1000 raw or unprocessed food products. The data in FooDB was collected from many sources including textbooks, scientific journals, on-line food composition or nutrient databases, flavour and aroma databases and various on-line metabolomic databases. This literature-derived information has been combined with experimentally derived data measured on thousands of compounds from more than 40 very common food products through the Alberta Food Metabolome Project which is led by David S. Wishart. Users are able to browse through the FooDB data by food source, name, descriptors or function. Chemical structures and molecular weights for compounds in FooDB may be searched via a specialized chemical structure search utility. Users are able to view the content of FooDB using two different “Viewing” options: FoodView, which lists foods by their chemical compounds, or ChemView, which lists chemicals by their food sources. Knowledge about the precise chemical composition of foods can be used to guide public health policies, assist food companies with improved food labelling, help dieticians prepare better dietary plans, support nutraceutical companies with their submissions of health claims and guide consumer choices with regard to food purchases.
The Serum Metabolome database is a free web database about small molecule metabolites found in human serum and their concentration values. The database includes chemical data, clinical data and molecular/biochemistry data from literature and experiment. This database also references many other databases, such as KEGG, PubChem, MetaCyc, ChEBI, PDB, Swiss-Prot, GenBank, and Human Metabolome Database (HMDB).

MetaboMiner is a tool which can be used to automatically or semi-automatically identify metabolites in complex biofluids from 2D-NMR spectra. MetaboMiner is able to handle both 1H-1H total correlation spectroscopy (TOCSY) and 1H-13C heteronuclear single quantum correlation (HSQC) data. It identifies compounds by comparing 2D spectral patterns in the NMR spectrum of the biofluid mixture with specially constructed libraries containing reference spectra of approximately 500 pure compounds. MetaboMiner protocol is available via MetaboMiner website.
The E. coli Metabolome Database (ECMDB) is a freely accessible, online database of small molecule metabolites found in or produced by Escherichia coli. Escherichia coli is perhaps the best studied bacterium on earth and has served as the "model microbe" in microbiology research for more than 60 years. The ECMDB is essentially an E. coli "omics" encyclopedia containing detailed data on the genome, proteome and metabolome of E. coli. ECMDB is part of a suite of organism-specific metabolomics databases that includes DrugBank, HMDB, YMDB and SMPDB. As a metabolomics resource, the ECMDB is designed to facilitate research in the area gut/microbiome metabolomics and environmental metabolomics. The ECMDB contains two kinds of data: 1) chemical data and 2) molecular biology and/or biochemical data. The chemical data includes more than 2700 metabolite structures with detailed metabolite descriptions along with nearly 5000 NMR, GC-MS and LC-MS spectra corresponding to these metabolites. The biochemical data includes nearly 1600 protein sequences and more than 3100 biochemical reactions that are linked to these metabolite entries. Each metabolite entry in the ECMDB contains more than 80 data fields with approximately 65% of the information being devoted to chemical data and the other 35% of the information devoted to enzymatic or biochemical data. Many data fields are hyperlinked to other databases. The ECMDB also has a variety of structure and pathway viewing applets. The ECMDB database offers a number of text, sequence, spectral, chemical structure and relational query searches. These are described in more detail below.
METAGENassist is a freely available web server for comparative metagenomic analysis. Comparative metagenomic studies involve the large-scale comparison of genomic or taxonomic census data from bacterial samples across different environments. Historically this has required a sound knowledge of statistics, computer programming, genetics and microbiology. As a result, only a small number of researchers are routinely able to perform comparative metagenomic studies. To circumvent these limitations, METAGENassist was developed to allow metagenomic analyses to be performed by non-specialists, easily and intuitively over the web. METAGENassist is particularly notable for its rich graphical output and its extensive database of bacterial phenotypic information.

Gene set enrichment analysis (GSEA) (also called functional enrichment analysis or pathway enrichment analysis) is a method to identify classes of genes or proteins that are over-represented in a large set of genes or proteins, and may have an association with different phenotypes (e.g. different organism growth patterns or diseases). The method uses statistical approaches to identify significantly enriched or depleted groups of genes. Transcriptomics technologies and proteomics results often identify thousands of genes, which are used for the analysis.

MetaboLights is a data repository founded in 2012 for cross-species and cross-platform metabolomic studies that provides primary research data and meta data for metabolomic studies as well as a knowledge base for properties of individual metabolites. The database is maintained by the European Bioinformatics Institute (EMBL-EBI) and the development is funded by Biotechnology and Biological Sciences Research Council (BBSRC). As of July 2018, the MetaboLights browse functionality consists of 383 studies, two analytical platforms, NMR spectroscopy and mass spectrometry.
David S. Wishart is a Canadian researcher in metabolomics and a Distinguished University Professor in the Department of Biological Sciences and the Department of Computing Science at the University of Alberta. Wishart also holds cross appointments in the Faculty of Pharmacy and Pharmaceutical Sciences and the Department of Laboratory Medicine and Pathology in the Faculty of Medicine and Dentistry. Additionally, Wishart holds a joint appointment in metabolomics at the Pacific Northwest National Laboratory in Richland, Washington. Wishart is well known for his pioneering contributions to the fields of protein NMR spectroscopy, bioinformatics, cheminformatics and metabolomics. In 2011, Wishart founded the Metabolomics Innovation Centre (TMIC), which is Canada's national metabolomics laboratory.
References
- 1 2 3 Xia, J; Wishart DS. (Jul 2010). "MSEA: a web-based tool to identify biologically meaningful patterns in quantitative metabolomic data". Nucleic Acids Res. 38 (Web Server issue): W71-7. doi:10.1093/nar/gkq329. PMC 2896187 . PMID 20457745.
- ↑ Subramanian, A; Tamayo P; Mootha VK; Mukherjee S; Ebert BL; Gillette MA; Paulovich A; Pomeroy SL; Golub TR; Lander ES; Mesirov JP. (Oct 2005). "Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles". Proc Natl Acad Sci U S A. 102 (43): 15545–50. doi: 10.1073/pnas.0506580102 . PMC 1239896 . PMID 16199517.
- ↑ Wishart DS, Jewison T, Guo AC, Wilson M, Knox C, Liu Y, Djoumbou Y, Mandal R, Aziat F, Dong E, Bouatra S, Sinelnikov I, Arndt D, Xia J, Liu P, Yallou F, Bjorndahl T, Perez-Pineiro R, Eisner R, Allen F, Neveu V, Greiner R, Scalbert A (Jan 2013). "HMDB 3.0--The Human Metabolome Database in 2013". Nucleic Acids Res. 41 (Database issue): D801-7. doi:10.1093/nar/gks1065. PMC 3531200 . PMID 23161693.
- 1 2 Xia, J; Mandal R; Sinelnikov IV; Broadhurst D; Wishart DS. (Jul 2012). "MetaboAnalyst 2.0--a comprehensive server for metabolomic data analysis". Nucleic Acids Res. 40 (Web Server issue): W127-33. doi:10.1093/nar/gks374. PMC 3394314 . PMID 22553367.
- ↑ Kessler, N; Neuweger H; Bonte A; Langenkämper G; Niehaus K; Nattkemper TW; Goesmann A. (Oct 2013). "MeltDB 2.0-advances of the metabolomics software system". Bioinformatics. 29 (19): 2452–9. doi:10.1093/bioinformatics/btt414. PMC 3777109 . PMID 23918246.