
A transmembrane protein (TP) is a type of integral membrane protein that spans the entirety of the cell membrane. Many transmembrane proteins function as gateways to permit the transport of specific substances across the membrane. They frequently undergo significant conformational changes to move a substance through the membrane. They are usually highly hydrophobic and aggregate and precipitate in water. They require detergents or nonpolar solvents for extraction, although some of them (beta-barrels) can be also extracted using denaturing agents.

Porins are beta barrel proteins that cross a cellular membrane and act as a pore, through which molecules can diffuse. Unlike other membrane transport proteins, porins are large enough to allow passive diffusion, i.e., they act as channels that are specific to different types of molecules. They are present in the outer membrane of gram-negative bacteria and some gram-positive mycobacteria, the outer membrane of mitochondria, and the outer chloroplast membrane.

The bacterial outer membrane is found in gram-negative bacteria. Its composition is distinct from that of the inner cytoplasmic cell membrane - among other things, the outer leaflet of the outer membrane of many gram-negative bacteria includes a complex lipopolysaccharide whose lipid portion acts as an endotoxin - and in some bacteria such as E. coli it is linked to the cell's peptidoglycan by Braun's lipoprotein.
The Transporter Classification Database is an International Union of Biochemistry and Molecular Biology (IUBMB)-approved classification system for membrane transport proteins, including ion channels.
Bacterial display is a protein engineering technique used for in vitro protein evolution. Libraries of polypeptides displayed on the surface of bacteria can be screened using flow cytometry or iterative selection procedures (biopanning). This protein engineering technique allows us to link the function of a protein with the gene that encodes it. Bacterial display can be used to find target proteins with desired properties and can be used to make affinity ligands which are cell-specific. This system can be used in many applications including the creation of novel vaccines, the identification of enzyme substrates and finding the affinity of a ligand for its target protein.
Protein K is a porin expressed in some pathogenic strains of E. coli bacteria. It has a molecular weight of about 40 kDa and is localized to the outer membrane, through which it allows both inorganic and organic ions to pass.

A colicin is a type of bacteriocin produced by and toxic to some strains of Escherichia coli. Colicins are released into the environment to reduce competition from other bacterial strains. Colicins bind to outer membrane receptors, using them to translocate to the cytoplasm or cytoplasmic membrane, where they exert their cytotoxic effect, including depolarisation of the cytoplasmic membrane, DNase activity, RNase activity, or inhibition of murein synthesis.

The MicC non-coding RNA is located between the ompN and ydbK genes in E. coli. This Hfq-associated RNA is thought to be a regulator of the expression level of the OmpC porin protein, with a 5′ region of 22 nucleotides potentially forming an antisense interaction with the ompC mRNA. Along with MicF RNA this family may act in conjunction with EnvZ-OmpR two-component system to control the OmpF/OmpC protein ratio in response to a variety of environmental stimuli. The expression of micC was shown to be increased in the presence of beta-lactam antibiotics.

The micF RNA is a non-coding RNA stress response gene found in Escherichia coli and related bacteria that post-transcriptionally controls expression of the outer membrane porin gene ompF. The micF gene encodes a non-translated 93 nucleotide antisense RNA that binds its target ompF mRNA and regulates ompF expression by inhibiting translation and inducing degradation of the message. In addition, other factors, such as the RNA chaperone protein StpA also play a role in this regulatory system. Expression of micF is controlled by both environmental and internal stress factors. Four transcriptional regulators are known to bind the micF promoter region and activate micF expression.

General bacterial porins are a family of proteins from the outer membranes of Gram-negative bacteria. The porins act as molecular filters for hydrophilic compounds. They are responsible for the 'molecular sieve' properties of the outer membrane. Porins form large water-filled channels which allow the diffusion of hydrophilic molecules into the periplasmic space. Some porins form general diffusion channels that allow any solute up to a certain size to cross the membrane, while other porins are specific for one particular solute and contain a binding site for that solute inside the pores. As porins are the major outer membrane proteins, they also serve as receptor sites for the binding of phages and bacteriocins.
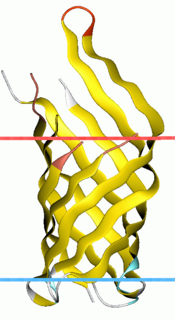
OmpA-like transmembrane domain is an evolutionarily conserved domain of bacterial outer membrane proteins. This domain consists of an eight-stranded beta barrel. OmpA is the predominant cell surface antigen in enterobacteria found in about 100,000 copies per cell. The expression of OmpA is tightly regulated by a variety of mechanisms. One mechanism by which OmpA expression is regulated in Vibrio species is by an antisense non-coding RNA called VrrA.
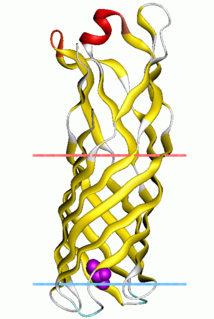
Outer membrane protein W (OmpW) family is a family of evolutionarily related proteins from the bacterial outer membrane.
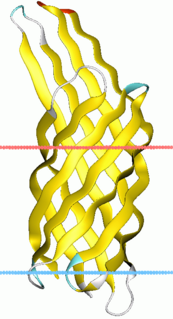
Virulence-related outer membrane proteins are expressed in the outer membrane of gram-negative bacteria and are essential to bacterial survival within macrophages and for eukaryotic cell invasion.

Nucleoside-specific porin is an outer membrane protein, Tsx, which constitutes the receptor for colicin K and Bacteriophage T6, and functions as a substrate-specific channel for nucleosides and deoxy-nucleosides. The protein contains 294 amino acids, the first 22 of which are characteristic of a bacterial signal sequence peptide. Tsx shows no significant similarities to general bacterial porins.

Invasion gene associated RNA is a small non-coding RNA involved in regulating one of the major outer cell membrane porin proteins in Salmonella species.
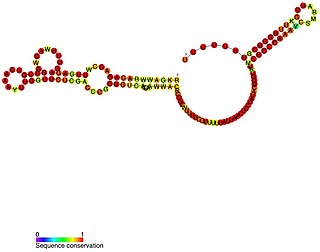

VrrA is a non-coding RNA that is conserved across all Vibrio species of bacteria and acts as a repressor for the synthesis of the outer membrane protein OmpA. This non-coding RNA was initially identified from Tn5 transposon mutant libraries of Vibrio cholerae and its location within the bacterial genome was mapped to the intergenic region between genes VC1741 and VC1743 by RACE analysis.
Bacterial small RNAs (sRNA) are small RNAs produced by bacteria; they are 50- to 500-nucleotide non-coding RNA molecules, highly structured and containing several stem-loops. Numerous sRNAs have been identified using both computational analysis and laboratory-based techniques such as Northern blotting, microarrays and RNA-Seq in a number of bacterial species including Escherichia coli, the model pathogen Salmonella, the nitrogen-fixing alphaproteobacterium Sinorhizobium meliloti, marine cyanobacteria, Francisella tularensis, Streptococcus pyogenes, the pathogen Staphylococcus aureus, and the plant pathogen Xanthomonas oryzae pathovar oryzae. Bacterial sRNAs affect how genes are expressed within bacterial cells via interaction with mRNA or protein, and thus can affect a variety of bacterial functions like metabolism, virulence, environmental stress response, and structure.
EnvZ/OmpR is a two-component regulatory system widely distributed in bacteria and particularly well characterized in Escherichia coli. Its function is in osmoregulation, responding to changes in environmental osmolality by regulating the expression of the outer membrane porins OmpF and OmpC. EnvZ is a histidine kinase which also possesses a cytoplasmic osmosensory domain, and OmpR is its corresponding response regulator protein.

In molecular biology, the OmpA domain is a conserved protein domain with a beta/alpha/beta/alpha-beta(2) structure found in the C-terminal region of many Gram-negative bacterial outer membrane proteins, such as porin-like integral membrane proteins, small lipid-anchored proteins, and MotB proton channels. The N-terminal half of these proteins is variable although some of the proteins in this group have the OmpA-like transmembrane domain at the N terminus. OmpA from Escherichia coli is required for pathogenesis, and can interact with host receptor molecules. MotB serve two functions in E. coli, the MotA(4)-MotB(2) complex attaches to the cell wall via MotB to form the stator of the flagellar motor, and the MotA-MotB complex couples the flow of ions across the cell membrane to movement of the rotor.

OmpT is an aspartyl protease found on the outer membrane of Escherichia coli. OmpT is a subtype of the family of omptin proteases, which are found on some gram-negative species of bacteria.















