
Escherichia coli, also known as E. coli, is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus Escherichia that is commonly found in the lower intestine of warm-blooded organisms (endotherms). Most E. coli strains are harmless, but some serotypes can cause serious food poisoning in their hosts, and are occasionally responsible for product recalls due to food contamination. The harmless strains are part of the normal microbiota of the gut, and can benefit their hosts by producing vitamin K2, and preventing colonization of the intestine with pathogenic bacteria, having a symbiotic relationship. E. coli is expelled into the environment within fecal matter. The bacterium grows massively in fresh fecal matter under aerobic conditions for 3 days, but its numbers decline slowly afterwards.
A sigma factor is a protein needed for initiation of transcription in bacteria. It is a bacterial transcription initiation factor that enables specific binding of RNA polymerase (RNAP) to gene promoters. It is homologous to archaeal transcription factor B and to eukaryotic factor TFIIB. The specific sigma factor used to initiate transcription of a given gene will vary, depending on the gene and on the environmental signals needed to initiate transcription of that gene. Selection of promoters by RNA polymerase is dependent on the sigma factor that associates with it.
The start codon is the first codon of a messenger RNA (mRNA) transcript translated by a ribosome. The start codon always codes for methionine in eukaryotes and a modified Met (fMet) in prokaryotes. The most common start codon is AUG.
Prokaryotic translation is the process by which messenger RNA is translated into proteins in prokaryotes.
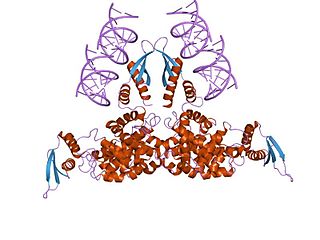
Ribonuclease III (BRENDA 3.1.26.3) is a type of ribonuclease that recognizes dsRNA and cleaves it at specific targeted locations to transform them into mature RNAs. These enzymes are a group of endoribonucleases that are characterized by their ribonuclease domain, which is labelled the RNase III domain. They are ubiquitous compounds in the cell and play a major role in pathways such as RNA precursor synthesis, RNA Silencing, and the pnp autoregulatory mechanism.
The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.
fis is an E. coli gene encoding the Fis protein. The regulation of this gene is more complex than most other genes in the E. coli genome, as Fis is an important protein which regulates expression of other genes. It is supposed that fis is regulated by H-NS, IHF and CRP. It also regulates its own expression (autoregulation). Fis is one of the most abundant DNA binding proteins in Escherichia coli under nutrient-rich growth conditions.

The gcvB RNA gene encodes a small non-coding RNA involved in the regulation of a number of amino acid transport systems as well as amino acid biosynthetic genes. The GcvB gene is found in enteric bacteria such as Escherichia coli. GcvB regulates genes by acting as an antisense binding partner of the mRNAs for each regulated gene. This binding is dependent on binding to a protein called Hfq. Transcription of the GcvB RNA is activated by the adjacent GcvA gene and repressed by the GcvR gene. A deletion of GcvB RNA from Y. pestis changed colony shape as well as reducing growth. It has been shown by gene deletion that GcvB is a regulator of acid resistance in E. coli. GcvB enhances the ability of the bacterium to survive low pH by upregulating the levels of the alternate sigma factor RpoS. A polymeric form of GcvB has recently been identified. Interaction of GcvB with small RNA SroC triggers the degradation of GcvB by RNase E, lifting the GcvB-mediated mRNA repression of its target genes.

The SraC/RyeA RNA is a non-coding RNA that was discovered in E. coli during two large scale screens for RNAs. The function of this RNA is currently unknown. This RNA overlaps the SdsR/RyeB RNA on the opposite strand suggesting that the two RNAs may act in a concerted manner.
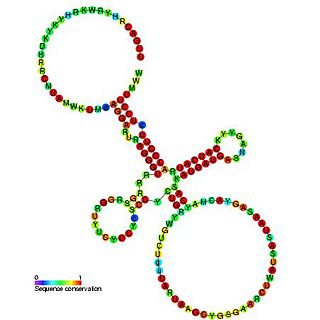
Sib RNA refers to a group of related non-coding RNA. They were originally named QUAD RNA after they were discovered as four repeat elements in Escherichia coli intergenic regions. The family was later renamed Sib when it was discovered that the number of repeats is variable in other species and in other E. coli strains.

RybB is a small non-coding RNA was identified in a large scale screen of Escherichia coli. The function of this short RNA has been studied using a transcriptomic approach and kinetic analyses of target mRNA decay in vivo. RybB was identified as a factor that selectively accelerates the decay of multiple major omp mRNAs upon induction of the envelope stress response. This RNA has been shown to bind to the Hfq protein.
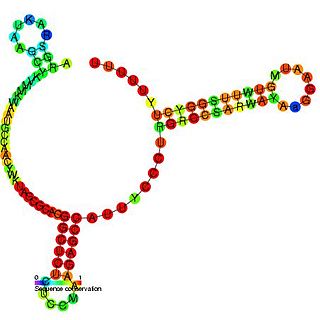
The SdsR/RyeB RNA is a non-coding RNA that was identified in a large scale screen of E. coli. The exact 5' and 3' ends of this RNA are uncertain. This RNA overlaps the SraC/RyeA RNA on the opposite strand suggesting that the two may act in a concerted manner. It is transcribed by general stress factor σs and is most highly expressed in stationary phase. SdsR/RyeB RNA interacts with Hfq.
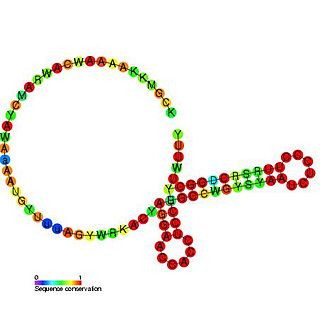
The CyaR RNA non-coding RNA was identified in a large scale screen of Escherichia coli and was called candidate 14. The exact 5' and 3' ends of this RNA are uncertain. This gene lies between yegQ and orgK in E. coli. This small RNA was shown to be bound by the Hfq protein. This RNA has been renamed as CyaR for. It has been shown that the CyaR RNA acts as a repressor of the porin OmpX. It has also been shown that cyaR expression is tightly controlled by the cyclic AMP receptor protein, CRP.
The ryfA RNA gene is a non-coding RNA present in E. coli, Shigella flexneri and Salmonella species where it is found between the ydaN and dbpA genes. These RNA genes are about 300 nucleotides in length. The function of this RNA is unknown.

RyhB RNA is a 90 nucleotide RNA that down-regulates a set of iron-storage and iron-using proteins when iron is limiting; it is itself negatively regulated by the ferric uptake repressor protein, Fur.

Spot 42 RNA is a regulatory non-coding bacterial small RNA encoded by the spf gene. Spf is found in gammaproteobacteria and the majority of experimental work on Spot42 has been performed in Escherichia coli and recently in Aliivibrio salmonicida. In the cell Spot42 plays essential roles as a regulator in carbohydrate metabolism and uptake, and its expression is activated by glucose, and inhibited by the cAMP-CRP complex.
The T44 RNA family consists of a number of bacterial RNA genes of between 135 and 170 bases in length. The t44 gene has been identified in several species of enteric bacteria but homologs have also been identified in Pseudomonas and Coxiella species. The t44 gene is found between the map and rpsB genes in all species in the full alignment apart from Shigella flexneri. The function of this RNA is unknown.
The hok/sok system is a postsegregational killing mechanism employed by the R1 plasmid in Escherichia coli. It was the first type I toxin-antitoxin pair to be identified through characterisation of a plasmid-stabilising locus. It is a type I system because the toxin is neutralised by a complementary RNA, rather than a partnered protein.

The 23S rRNA is a 2904 nt long component of the large subunit (50S) of the bacterial/archean ribosome. The ribosomal peptidyl transferase activity resides in domain V of this rRNA, and this domain is the most common binding site for antibiotics that inhibit translation. A well-known member of this antibiotic class, chloramphenicol, acts by inhibiting peptide bond formation, with recent 3D-structural studies showing two different binding sites depending on the species of ribosome. Linezolid and quinupristin-dalfopristin also bind to the 23S rRNA, and cross-resistance has been demonstrated between these antibiotics. Compared to 16S rRNA genes, 23S rRNA genes typically have higher sequence variations including insertions and/or deletions.

An Hfq binding sRNA is an sRNA that binds the bacterial RNA binding protein called Hfq. A number of bacterial small RNAs which have been shown to bind to Hfq have been characterised . Many of these RNAs share a similar structure composed of three stem-loops. Several studies have expanded this list, and experimentally validated a total of 64 Hfq binding sRNA in Salmonella Typhimurium. A transcriptome wide study on Hfq binding sites in Salmonella mapped 126 Hfq binding sites within sRNAs. Genomic SELEX has been used to show that Hfq binding RNAs are enriched in the sequence motif 5'-AAYAAYAA-3'. Genome-wide study identified 40 candidate Hfq-dependent sRNAs in plant pathogen Erwinia amylovora. 12 of them were confirmed by Northern blot.


















