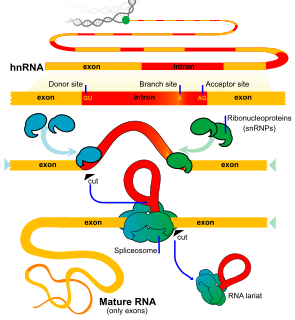
An exon is any part of a gene that will encode a part of the final mature RNA produced by that gene after introns have been removed by RNA splicing. The term exon refers to both the DNA sequence within a gene and to the corresponding sequence in RNA transcripts. In RNA splicing, introns are removed and exons are covalently joined to one another as part of generating the mature messenger RNA. Just as the entire set of genes for a species constitutes the genome, the entire set of exons constitutes the exome.

RNA splicing, in molecular biology, is a form of RNA processing in which a newly made precursor messenger RNA (pre-mRNA) transcript is transformed into a mature messenger RNA (mRNA). During splicing, introns are removed and exons are joined together. For nuclear-encoded genes, splicing takes place within the nucleus either during or immediately after transcription. For those eukaryotic genes that contain introns, splicing is usually required in order to create an mRNA molecule that can be translated into protein. For many eukaryotic introns, splicing is carried out in a series of reactions which are catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs). Self-splicing introns, or ribozymes capable of catalyzing their own excision from their parent RNA molecule, also exist.
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splicing to create monocistronic mRNAs that are translated separately, i.e. several strands of mRNA that each encode a single gene product. The result of this is that the genes contained in the operon are either expressed together or not at all. Several genes must be co-transcribed to define an operon.

A non-coding RNA (ncRNA) is an RNA molecule that is not translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally important types of non-coding RNAs include transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs), as well as small RNAs such as microRNAs, siRNAs, piRNAs, snoRNAs, snRNAs, exRNAs, scaRNAs and the long ncRNAs such as Xist and HOTAIR.

Alternative splicing, or alternative RNA splicing, or differential splicing, is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. Consequently, the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions. Notably, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes.

A spliceosome is a large and complex molecular machine found primarily within the nucleus of eukaryotic cells. The spliceosome is assembled from small nuclear RNAs (snRNA) and approximately 80 proteins. The spliceosome removes introns from a transcribed pre-mRNA, a type of primary transcript. This process is generally referred to as splicing. An analogy is a film editor, who selectively cuts out irrelevant or incorrect material from the initial film and sends the cleaned-up version to the director for the final cut.
Trans-splicing is a special form of RNA processing where exons from two different primary RNA transcripts are joined end to end and ligated. It is usually found in eukaryotes and mediated by the spliceosome, although some bacteria and archaea also have "half-genes" for tRNAs.
An internal ribosome entry site, abbreviated IRES, is an RNA element that allows for translation initiation in poly-A tail dependent manner, as part of the greater process of protein synthesis. In eukaryotic translation, initiation typically occurs at the 5' end of mRNA molecules, since tail is required for the assembly of the initiation complex. The location for IRES elements is often in the 5'UTR, but can also occur elsewhere in mRNAs.

This family represents the SL1 RNA. The gene encoding SL1 RNA is commonly, but not always, located in the spacer region between 5S-rRNA genes. The SL1 RNA is involved in trans-splicing, which is a form of RNA processing. The acquisition of a spliced leader from an SL RNA is an inter-molecular reaction that precisely joins exons derived from separately transcribed RNAs. This mechanism of mRNA maturation has been shown to occur in a number of lower eukaryotes and is commonplace in nematodes.

Small nucleolar RNA TBR17 is a non-coding RNA (ncRNA) molecule identified in Trypanosoma brucei which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.
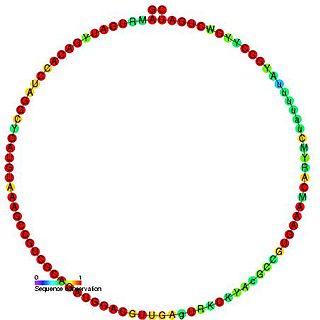
TBR5 is a member of the C/D class of snoRNA which contain the C (UGAUGA) and D (CUGA) box motifs. Most of the members of the box C/D family function in directing site-specific 2'-O-methylation of substrate RNAs.
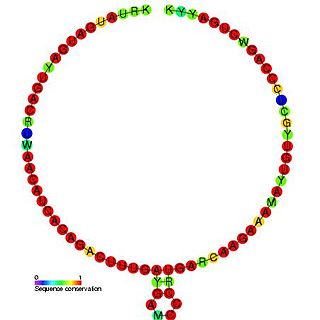
Small nucleolar RNA TBR7 is a non-coding RNA (ncRNA) molecule identified in Trypanosoma brucei which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.

The U4 small nuclear Ribo-Nucleic Acid is a non-coding RNA component of the major or U2-dependent spliceosome – a eukaryotic molecular machine involved in the splicing of pre-messenger RNA (pre-mRNA). It forms a duplex with U6, and with each splicing round, it is displaced from the U6 snRNA in an ATP-dependent manner, allowing U6 to re-fold and create the active site for splicing catalysis. A recycling process involving protein Brr2 releases U4 from U6, while protein Prp24 re-anneals U4 and U6. The crystal structure of a 5′ stem-loop of U4 in complex with a binding protein has been solved.

U6 snRNA is the non-coding small nuclear RNA (snRNA) component of U6 snRNP, an RNA-protein complex that combines with other snRNPs, unmodified pre-mRNA, and various other proteins to assemble a spliceosome, a large RNA-protein molecular complex that catalyzes the excision of introns from pre-mRNA. Splicing, or the removal of introns, is a major aspect of post-transcriptional modification and takes place only in the nucleus of eukaryotes.
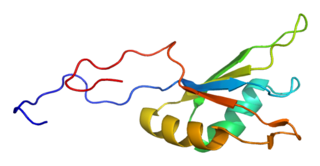
Fox-1 homolog A, also known as ataxin 2-binding protein 1 (A2BP1) or hexaribonucleotide-binding protein 1 (HRNBP1) or RNA binding protein, fox-1 homolog (Rbfox1), is a protein that in humans is encoded by the RBFOX1 gene.
Trans-Spliced Exon Coupled RNA End Determination (TEC-RED) is a transcriptomic technique that, like SAGE, allows for the digital detection of messenger RNA sequences. Unlike SAGE, detection and purification of transcripts from the 5’ end of the messenger RNA require the presence of a trans-spliced leader sequence.
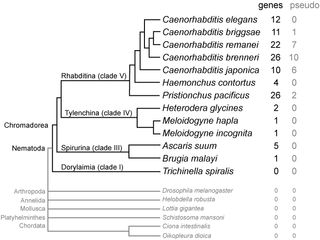
SmY ribonucleic acids are a family of small nuclear RNAs found in some species of nematode worms. They are thought to be involved in mRNA trans-splicing.

Prp8 refers to both the Prp8 protein and Prp8 gene. Prp8's name originates from its involvement in pre-mRNA processing. The Prp8 protein is a large, highly conserved, and unique protein that resides in the catalytic core of the spliceosome and has been found to have a central role in molecular rearrangements that occur there. Prp8 protein is a major central component of the catalytic core in the spliceosome, and the spliceosome is responsible for splicing of precursor mRNA that contains introns and exons. Unexpressed introns are removed by the spliceosome complex in order to create a more concise mRNA transcript. Splicing is just one of many different post-transcriptional modifications that mRNA must undergo before translation. Prp8 has also been hypothesized to be a cofactor in RNA catalysis.
Alice Barkan is an American molecular biologist and a professor of biology at the University of Oregon. She is known for her work on chloroplast gene regulation and protein synthesis.
An outron is a nucleotide sequence at the 5' end of the primary transcript of a gene that is removed by a special form of RNA splicing during maturation of the final RNA product. Whereas intron sequences are located inside the gene, outron sequences lie outside the gene.














