Related Research Articles
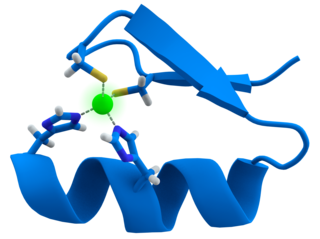
A zinc finger is a small protein structural motif that is characterized by the coordination of one or more zinc ions (Zn2+) in order to stabilize the fold. It was originally coined to describe the finger-like appearance of a hypothesized structure from the African clawed frog (Xenopus laevis) transcription factor IIIA. However, it has been found to encompass a wide variety of differing protein structures in eukaryotic cells. Xenopus laevis TFIIIA was originally demonstrated to contain zinc and require the metal for function in 1983, the first such reported zinc requirement for a gene regulatory protein followed soon thereafter by the Krüppel factor in Drosophila. It often appears as a metal-binding domain in multi-domain proteins.
In biology, a sequence motif is a nucleotide or amino-acid sequence pattern that is widespread and usually assumed to be related to biological function of the macromolecule. For example, an N-glycosylation site motif can be defined as Asn, followed by anything but Pro, followed by either Ser or Thr, followed by anything but Pro residue.
DnaG is a bacterial DNA primase and is encoded by the dnaG gene. The enzyme DnaG, and any other DNA primase, synthesizes short strands of RNA known as oligonucleotides during DNA replication. These oligonucleotides are known as primers because they act as a starting point for DNA synthesis. DnaG catalyzes the synthesis of oligonucleotides that are 10 to 60 nucleotides long, however most of the oligonucleotides synthesized are 11 nucleotides. These RNA oligonucleotides serve as primers, or starting points, for DNA synthesis by bacterial DNA polymerase III. DnaG is important in bacterial DNA replication because DNA polymerase cannot initiate the synthesis of a DNA strand, but can only add nucleotides to a preexisting strand. DnaG synthesizes a single RNA primer at the origin of replication. This primer serves to prime leading strand DNA synthesis. For the other parental strand, the lagging strand, DnaG synthesizes an RNA primer every few kilobases (kb). These primers serve as substrates for the synthesis of Okazaki fragments.

EGR-1 also known as ZNF268 or NGFI-A is a protein that in humans is encoded by the EGR1 gene.

Two-hybrid screening is a molecular biology technique used to discover protein–protein interactions (PPIs) and protein–DNA interactions by testing for physical interactions between two proteins or a single protein and a DNA molecule, respectively.
A DNA-binding domain (DBD) is an independently folded protein domain that contains at least one structural motif that recognizes double- or single-stranded DNA. A DBD can recognize a specific DNA sequence or have a general affinity to DNA. Some DNA-binding domains may also include nucleic acids in their folded structure.
C7 protein is an engineered zinc finger protein based on the murine ZFP, Zif268 and discovered by Wu et al. in 1994. It shares the same zinc finger 2 and zinc finger 3 of Zif268, but differs in the sequence of finger 1. It also shares the same DNA target, 5'-GCGTGGGCG-3'.
The C7.GAT protein is a zinc finger protein based on the C7 protein. It features an alternative zinc finger 3 alpha helix sequence, preventing the target site overlap caused by the aspartic acid residue of the finger 3 of C7. The sequence of this third finger is TSG-N-LVR according to the single letter amino acid code. As the name suggest, the target site of finger 3 is altered to 5'-GAT-3', giving the overall protein a target of 5'-GCGTGGGAT-3'.
Therapeutic gene modulation refers to the practice of altering the expression of a gene at one of various stages, with a view to alleviate some form of ailment. It differs from gene therapy in that gene modulation seeks to alter the expression of an endogenous gene whereas gene therapy concerns the introduction of a gene whose product aids the recipient directly.

Tristetraprolin (TTP), also known as zinc finger protein 36 homolog (ZFP36), is a protein that in humans, mice and rats is encoded by the ZFP36 gene. It is a member of the TIS11 family, along with butyrate response factors 1 and 2.
Zinc finger protein chimera are chimeric proteins composed of a DNA-binding zinc finger protein domain and another domain through which the protein exerts its effect. The effector domain may be a transcriptional activator (A) or repressor (R), a methylation domain (M) or a nuclease (N).

Zinc finger protein 804A is a protein that in humans is encoded by the ZNF804A gene. The human gene maps to chromosome 2 q32.1 and consists of 4 exons that code for a protein of 1210 amino acids.
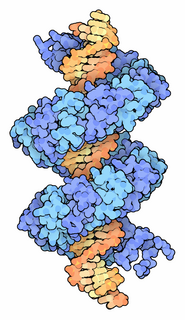
TALeffectors are proteins secreted by some β- and γ-proteobacteria. Most of these are Xanthomonads. Plant pathogenic Xanthomonas bacteria are especially known for TALEs, produced via their type III secretion system. These proteins can bind promoter sequences in the host plant and activate the expression of plant genes that aid bacterial infection. They recognize plant DNA sequences through a central repeat domain consisting of a variable number of ~34 amino acid repeats. There appears to be a one-to-one correspondence between the identity of two critical amino acids in each repeat and each DNA base in the target sequence. These proteins are interesting to researchers both for their role in disease of important crop species and the relative ease of retargeting them to bind new DNA sequences. Similar proteins can be found in the pathogenic bacterium Ralstonia solanacearum and Burkholderia rhizoxinica, as well as yet unidentified marine microorganisms. The term TALE-likes is used to refer to the putative protein family encompassing the TALEs and these related proteins.

Transcription activator-like effector nucleases (TALEN) are restriction enzymes that can be engineered to cut specific sequences of DNA. They are made by fusing a TAL effector DNA-binding domain to a DNA cleavage domain. Transcription activator-like effectors (TALEs) can be engineered to bind to practically any desired DNA sequence, so when combined with a nuclease, DNA can be cut at specific locations. The restriction enzymes can be introduced into cells, for use in gene editing or for genome editing in situ, a technique known as genome editing with engineered nucleases. Alongside zinc finger nucleases and CRISPR/Cas9, TALEN is a prominent tool in the field of genome editing.
The RNA-binding Proteins Database (RBPDB) is a biological database of RNA-binding protein specificities that includes experimental observations of RNA-binding sites. The experimental results included are both in vitro and in vivo from primary literature. It includes four metazoan species, which are Homo sapiens, Mus musculus, Drosophila melanogaster, and Caenorhabditis elegans. RNA-binding domains included in this database are RNA recognition motif, K homology, CCCH zinc finger, and more domains. As of 2021, the latest RBPDB release includes 1,171 RNA-binding proteins.

Genome editing, or genome engineering, or gene editing, is a type of genetic engineering in which DNA is inserted, deleted, modified or replaced in the genome of a living organism. Unlike early genetic engineering techniques that randomly inserts genetic material into a host genome, genome editing targets the insertions to site specific locations.
Zinc finger transcription factors or ZF-TFs, are transcription factors composed of a zinc finger-binding domain and any of a variety of transcription-factor effector-domains that exert their modulatory effect in the vicinity of any sequence to which the protein domain binds.
In molecular biology, a GC box, also known as a GSG box, is a distinct pattern of nucleotides found in the promoter region of some eukaryotic genes. The GC box is upstream of the TATA box and approximately 110 bases upstream from the transcription initiation site. It has a consensus sequence GGGCGG which is position-dependent and orientation-independent. The GC elements are bound by transcription factors and have similar functions to enhancers. Some known GC box-binding proteins include Sp1, Krox/Egr, Wilms' tumor, MIGI, and CREA.

The WRKY domain is found in the WRKY transcription factor family, a class of transcription factors. The WRKY domain is found almost exclusively in plants although WRKY genes appear present in some diplomonads, social amoebae and other amoebozoa, and fungi incertae sedis. They appear absent in other non-plant species. WRKY transcription factors have been a significant area of plant research for the past 20 years. The WRKY DNA-binding domain recognizes the W-box (T)TGAC(C/T) cis-regulatory element.
References
- 1 2 Wolfe SA, Grant RA, Elrod-Erickson M, Pabo CO (2001). "Beyond the "recognition code": structures of two Cys2His2 zinc finger/TATA box complexes". Structure. 9 (8): 717–23. doi: 10.1016/S0969-2126(01)00632-3 . PMID 11587646.
- 1 2 Beerli RR, Barbas CF (2002). "Engineering polydactyl zinc-finger transcription factors". Nat. Biotechnol. 20 (2): 135–41. doi:10.1038/nbt0202-135. PMID 11821858. S2CID 12685879.
- 1 2 3 Segal DJ, Dreier B, Beerli RR, Barbas CF (1999). "Toward controlling gene expression at will: selection and design of zinc finger domains recognizing each of the 5'-GNN-3' DNA target sequences". Proc. Natl. Acad. Sci. U.S.A. 96 (6): 2758–63. Bibcode:1999PNAS...96.2758S. doi: 10.1073/pnas.96.6.2758 . PMC 15842 . PMID 10077584.
- ↑ Dreier B, Fuller RP, Segal DJ, et al. (2005). "Development of zinc finger domains for recognition of the 5'-CNN-3' family DNA sequences and their use in the construction of artificial transcription factors". J. Biol. Chem. 280 (42): 35588–97. doi: 10.1074/jbc.M506654200 . PMID 16107335.