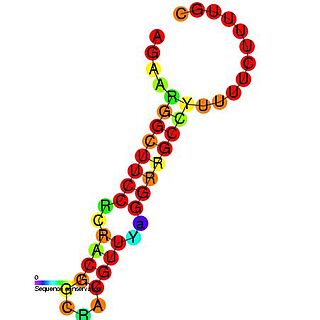
In molecular biology ctRNA is a plasmid encoded noncoding RNA that binds to the mRNA of repB and causes translational inhibition. ctRNA is encoded by plasmids and functions in rolling circle replication to maintain a low copy number. In Corynebacterium glutamicum, it achieves this by antisense pairing with the mRNA of RepB, a replication initiation protein. In Enterococcus faecium the plasmid pJB01 contains three open reading frames, copA, repB, and repC. The pJB01 ctRNA is coded on the opposite strand from the copA/repB intergenic region and partially overlaps an atypical ribosome binding site for repB.
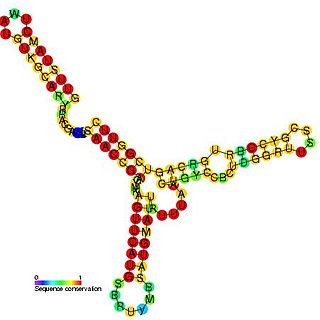
Anti-Q RNA is a small ncRNA from the conjugal plasmid pCF10 of Enterococcus faecalis. It is coded in cis to its regulatory target, prgQ, but can also act in trans. Anti-Q is known to interact with nascent prgQ transcripts to allow formation of an intrinsic terminator, or attenuator, thus preventing transcription of downstream genes. This mode of regulation is essentially the same as that of the countertranscript-driven attenuators that control copy number in pT181, pAMbeta1 and pIP501 and related Staphylococcal plasmids.

R1162-like plasmid antisense RNA is a 75-base RNA molecule which negatively regulates the RepI region of the plasmid. The protein product of this gene region, along with another protein, controls the copy number of the 8.75kB R1162 plasmid.
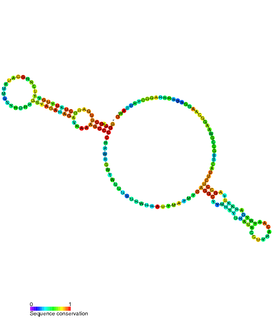
The hok/sok system is a postsegregational killing mechanism employed by the R1 plasmid in Escherichia coli. It was the first type I toxin-antitoxin pair to be identified through characterisation of a plasmid-stabilising locus. It is a type I system because the toxin is neutralised by a complementary RNA, rather than a partnered protein.

The Bacillus-plasmid RNA motif is a predicted conserved RNA structure usually located in plasmids. It is known in species under the genera Bacillus and Lactobacillus. In Bacillus subtilis, it is found upstream of the hypothetical gene ydcS, whose function is unknown.

The Chlorobi-RRM RNA motif is a conserved RNA structure identified by bioinformatics. It is found within bacteria in the phylum Chlorobi, and is exclusively detected in the presumed 5' untranslated regions of genes that encode putative RNA-binding proteins. Since many RNA-binding proteins regulate their own expression in a feedback mechanism by binding or acting up their 5' UTR, it was proposed that the Chlorobi-RRM is a component in an analogous feedback mechanism. Structurally, the motif consists of two stem-loops, the second of which might function as a rho-independent transcription terminator.

The L17 downstream element RNA motif is a conserved RNA structure identified in bacteria by bioinformatics. All known L17 downstream elements were detected immediately downstream of genes encoding the L17 subunit of the ribosome, and therefore might be in the 3' untranslated regions of these genes. The element is found in a variety of lactic acid bacteria and in the genus Listeria.

The lactis-plasmid RNA motif is a conserved RNA structure identified by bioinformatics. The RNAs are restricted to lactic acid bacteria, and are especially common in Lactococcus lactis. They typically lie near to repB genes, and are almost found in plasmids. This data suggested that lactis-plasmid RNAs participate in the control of plasmid abundance. However, many of the plasmids that carry lactis-plasmid RNAs also carry ctRNA-pND324 RNAs, which are involved in plasmid copy count regulation. Therefore lactis-plasmid RNAs might have a different function.

PhotoRC RNA motifs refer to conserved RNA structures that are associated with genes acting in the photosynthetic reaction centre of photosynthetic bacteria. Two such RNA classes were identified and called the PhotoRC-I and PhotoRC-II motifs. PhotoRC-I RNAs were detected in the genomes of some cyanobacteria. Although no PhotoRC-II RNA has been detected in cyanobacteria, one is found in the genome of a purified phage that infects cyanobacteria. Both PhotoRC-I and PhotoRC-II RNAs are present in sequences derived from DNA that was extracted from uncultivated marine bacteria.

The rmf RNA motif is a conserved RNA structure that was originally detected using bioinformatics. rmf RNAs are consistently foundwithin species classified into the genus Pseudomonas, and is located potentially in the 5' untranslated regions of rmf genes. These genes encodes the ribosome modulation factor protein, which affects the translation of genes by modifying ribosome structure in response to stress such as starvation. This ribosome modulation is a part of the stringent response in bacteria. The likely biological role of rmf RNAs is ambiguous. Since the RNA could be in the 5' UTRs of protein-coding genes, it was hypothesized that it functions as a cis-regulatory element. This hypothesis is bolstered by the observation that ribosome modulation factor binds ribosomal RNA, and many cis-regulatory RNAs called ribosomal protein leaders participate in a feedback regulation mechanism by binding to proteins that normally bind to ribosomal RNA. However, since rmf RNAs are not very close to the rmf genes, they might function as non-coding RNAs.

The rne-II RNA motif is a conserved RNA structure identified using bioinformatics. It is detected only in species classified within the family Pseudomonadaceae, a group of gammaproteobacteria. rne-II RNAs are consistently located in the presumed 5' untranslated regions of genes that encode Ribonuclease E. The RNase E 5' UTR element is a previously identified RNA structure that is also found in the 5' UTRs of RNase E genes. However, the latter motif is found only in enterobacteria, and the two motifs have apparently unrelated structure. In view of their differences, it was hypothesized that rne-II RNAs fulfill the same functional role as RNase E 5' UTR elements, which is to regulate the levels of RNase E proteins by acting as a substrate for RNase E. Thus, when concentrations of RNase E are high, they will degrade their own messenger RNA.

The traJ-II RNA motif is a conserved RNA structure discovered in bacteria by using bioinformatics. traJ-II RNAs appear to be in the 5' untranslated regions of protein-coding genes called traJ, which functions in the process of bacterial conjugation. A previously identified motif known as TraJ 5' UTR is also found upstream of traJ genes functions as the target of FinP antisense RNAs, so it is possible that traJ-II RNAs play a similar role as targets of an antisense RNA. However, some sequence features within the traJ-II RNA motif suggest that the biological RNA might be transcribed from the reverse-complement strand. Thus is it unclear whether traJ-II function as cis-regulatory elements. traJ-II RNAs are found in a variety of proteobacteria.

The yjdF RNA motif is a conserved RNA structure identified using bioinformatics. Most yjdF RNAs are located in bacteria classified within the phylum Firmicutes. A yjdF RNA is found in the presumed 5' untranslated region of the yjdF gene in Bacillus subtilis, and almost all yjdF RNAs are found in the 5' UTRs of homologs of this gene. The function of the yjdF gene is unknown, but the protein that it is predicted to encode is classified by the Pfam Database as DUF2992.
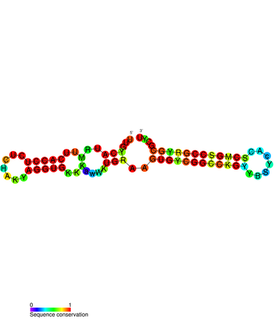
PtaRNA1 is a family of non-coding RNAs. Homologs of PtaRNA1 can be found in the proteobacteria families, Betaproteobacteria and Gammaproteobacteria. In all cases the PtaRNA1 is located anti-sense to a short protein-coding gene. In Xanthomonas campestris pv. vesicatoria, this gene is annotated as XCV2162 and is included in the plasmid toxin family of proteins.

A toxin-antitoxin system is a set of two or more closely linked genes that together encode both a "toxin" protein and a corresponding "antitoxin". When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK). Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies.
Plasmid-mediated resistance is the transfer of antibiotic resistance genes which are carried on plasmids. The plasmids can be transferred between bacteria within the same species or between different species via conjugation. Plasmids often carry multiple antibiotic resistance genes, contributing to the spread of multidrug-resistance (MDR). Antibiotic resistance mediated by MDR plasmids severely limits the treatment options for the infections caused by Gram-negative bacteria, especially Enterobacteriaceae family. The global spread of MDR plasmids has been enhanced by selective pressure from antibiotic usage in human and veterinary medicine.

The par stability determinant is a 400 bp locus of the pAD1 plasmid which encodes a type I toxin-antitoxin system in Enterococcus faecalis. It was the first such plasmid addiction module to be found in gram-positive bacteria.

ZEB2-AS1 is a long non-coding RNA, which is overlapping and antisense to the ZEB2 gene. It overlaps the 5' splice site of an intron within the 5'UTR of the ZEB2 gene. This intron contains an internal ribosome entry site (IRES), which is necessary for ZEB2 expression. ZEB2-AS1 prevents the splicing of this intron, and therefore activates ZEB2 expression.



















