In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splicing to create monocistronic mRNAs that are translated separately, i.e. several strands of mRNA that each encode a single gene product. The result of this is that the genes contained in the operon are either expressed together or not at all. Several genes must be co-transcribed to define an operon.

The lac operon is an operon required for the transport and metabolism of lactose in Escherichia coli and many other enteric bacteria. Although glucose is the preferred carbon source for most bacteria, the lac operon allows for the effective digestion of lactose when glucose is not available through the activity of beta-galactosidase. Gene regulation of the lac operon was the first genetic regulatory mechanism to be understood clearly, so it has become a foremost example of prokaryotic gene regulation. It is often discussed in introductory molecular and cellular biology classes for this reason. This lactose metabolism system was used by François Jacob and Jacques Monod to determine how a biological cell knows which enzyme to synthesize. Their work on the lac operon won them the Nobel Prize in Physiology in 1965.
A transcriptional activator is a protein that increases gene transcription of a gene or set of genes. Most activators are DNA-binding proteins that bind to enhancers or promoter-proximal elements.

Tryptophan repressor is a transcription factor involved in controlling amino acid metabolism. It has been best forstudied in Escherichia coli, where it is a dimeric protein that regulates transcription of the 5 genes in the tryptophan operon. When the amino acid tryptophan is plentiful in the cell, it binds to the protein, which causes a conformational change in the protein. The repressor complex then binds to its operator sequence in the genes it regulates, shutting off the genes.
Charles Yanofsky was an American geneticist on the faculty of Stanford University who contributed to the establishment of the one gene-one enzyme hypothesis and discovered attenuation, a riboswitch mechanism in which messenger RNA changes shape in response to a small molecule and thus alters its binding ability for the regulatory region of a gene or operon.

Ribosomal ribonucleic acid (rRNA) is the RNA component of the ribosome, which is essential for protein synthesis in all living organisms. rRNA is the predominant RNA in most cells, composing around 80% of cellular RNA. Ribosomes are approximately 60% rRNA and 40% protein by weight. A ribosome contains two subunits, the large ribosomal subunit (LSU) and small ribosomal subunit (SSU).
Gene structure is the organisation of specialised sequence elements within a gene. Genes contain the information necessary for living cells to survive and reproduce. In most organisms, genes are made of DNA, where the particular DNA sequence determines the function of the gene. A gene is transcribed (copied) from DNA into RNA, which can either be non-coding (ncRNA) with a direct function, or an intermediate messenger (mRNA) that is then translated into protein. Each of these steps is controlled by specific sequence elements, or regions, within the gene. Every gene, therefore, requires multiple sequence elements to be functional. This includes the sequence that actually encodes the functional protein or ncRNA, as well as multiple regulatory sequence regions. These regions may be as short as a few base pairs, up to many thousands of base pairs long.
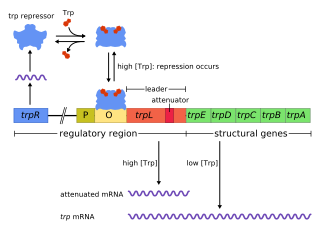
The trp operon is an operon—a group of genes that is used, or transcribed, together—that codes for the components for production of tryptophan. The trp operon is present in many bacteria, but was first characterized in Escherichia coli. The operon is regulated so that when tryptophan is present in the environment, the genes for tryptophan synthesis are not expressed. It was an important experimental system for learning about gene regulation, and is commonly used to teach gene regulation.
The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.

Spot 42 RNA is a regulatory non-coding bacterial small RNA encoded by the spf gene. Spf is found in gammaproteobacteria and the majority of experimental work on Spot42 has been performed in Escherichia coli and recently in Aliivibrio salmonicida. In the cell Spot42 plays essential roles as a regulator in carbohydrate metabolism and uptake, and its expression is activated by glucose, and inhibited by the cAMP-CRP complex.

(p)ppGpp, guanosine pentaphosphate or tetraphosphate is an alarmone which is involved in the stringent response in bacteria, causing the inhibition of RNA synthesis when there is a shortage of amino acids present. This causes translation to decrease and the amino acids present are therefore conserved. Furthermore, ppGpp causes the up-regulation of many other genes involved in stress response such as the genes for amino acid uptake and biosynthesis.
The gal operon is a prokaryotic operon, which encodes enzymes necessary for galactose metabolism. The operon contains two operators, OE and OI. The former is just before the promoter, and the latter is just after the galE gene.

The JUMPstart RNA motif describes a conserved RNA-based secondary structure associated with JUMPstart elements. The 39-base-pair JUMPstart sequence describes a conserved element upstream of genes that participate in polysaccharide synthesis. The JUMPstart element has been shown to function as an RNA, and is present in the 5' untranslated regions of the genes it regulates.
RegulonDB is a database of the regulatory network of gene expression in Escherichia coli K-12. RegulonDB also models the organization of the genes in transcription units, operons and regulons. A total of 120 sRNAs with 231 total interactions which all together regulate 192 genes are also included. RegulonDB was founded in 1998 and also contributes data to the EcoCyc database.
In DNA repair, the Ada Regulon is a set of genes whose expression is essential to adaptive response, which is triggered in prokaryotic cells by exposure to sub-lethal doses of alkylating agents. This allows the cells to tolerate the effects of such agents, which are otherwise toxic and mutagenic.
The gua operon is responsible for regulating the synthesis of guanosine mono phosphate(GMP), a purine nucleotide, from inosine monophosphate. It consists of two structural genes guaB(encodes for IMP dehydrogenase or IMPDH) and guaA(encodes for GMP synthetase) apart from the promoter and operator region.

The gab operon is responsible for the conversion of γ-aminobutyrate (GABA) to succinate. The gab operon comprises three structural genes – gabD, gabT and gabP – that encode for a succinate semialdehyde dehydrogenase, GABA transaminase and a GABA permease respectively. There is a regulatory gene csiR, downstream of the operon, that codes for a putative transcriptional repressor and is activated when nitrogen is limiting.
The locus of enterocyte effacement-encoded regulator (Ler) is a regulatory protein that controls bacterial pathogenicity of enteropathogenic Escherichia coli (EPEC) and enterohemorrhagic Escherichia coli (EHEC). More specifically, Ler regulates the locus of enterocyte effacement (LEE) pathogenicity island genes, which are responsible for creating intestinal attachment and effacing lesions and subsequent diarrhea: LEE1, LEE2, and LEE3. LEE1, 2, and 3 carry the information necessary for a type III secretion system. The transcript encoding the Ler protein is the open reading frame 1 on the LEE1 operon.