Related Research Articles

Rhizobium is a genus of Gram-negative soil bacteria that fix nitrogen. Rhizobium species form an endosymbiotic nitrogen-fixing association with roots of (primarily) legumes and other flowering plants.

Ensifer meliloti are an aerobic, Gram-negative, and diazotrophic species of bacteria. S. meliloti are motile and possess a cluster of peritrichous flagella. S. meliloti fix atmospheric nitrogen into ammonia for their legume hosts, such as alfalfa. S. meliloti forms a symbiotic relationship with legumes from the genera Medicago, Melilotus and Trigonella, including the model legume Medicago truncatula. This symbiosis promotes the development of a plant organ, termed a root nodule. Because soil often contains a limited amount of nitrogen for plant use, the symbiotic relationship between S. meliloti and their legume hosts has agricultural applications. These techniques reduce the need for inorganic nitrogenous fertilizers.

The Hyphomicrobiales are an order of Gram-negative Alphaproteobacteria.

Alphaproteobacteria is a class of bacteria in the phylum Pseudomonadota. The Magnetococcales and Mariprofundales are considered basal or sister to the Alphaproteobacteria. The Alphaproteobacteria are highly diverse and possess few commonalities, but nevertheless share a common ancestor. Like all Proteobacteria, its members are gram-negative and some of its intracellular parasitic members lack peptidoglycan and are consequently gram variable.

Bradyrhizobium is a genus of Gram-negative soil bacteria, many of which fix nitrogen. Nitrogen fixation is an important part of the nitrogen cycle. Plants cannot use atmospheric nitrogen (N2); they must use nitrogen compounds such as nitrates.
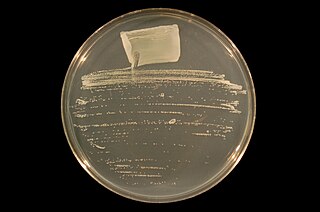
Ensifer is a genus of nitrogen-fixing bacteria (rhizobia), three of which have been sequenced.

MicrobesOnline is a publicly and freely accessible website that hosts multiple comparative genomic tools for comparing microbial species at the genomic, transcriptomic and functional levels. MicrobesOnline was developed by the Virtual Institute for Microbial Stress and Survival, which is based at the Lawrence Berkeley National Laboratory in Berkeley, California. The site was launched in 2005, with regular updates until 2011.
Actinorhizal plants are a group of angiosperms characterized by their ability to form a symbiosis with the nitrogen fixing actinomycetota Frankia. This association leads to the formation of nitrogen-fixing root nodules.
Bradyrhizobium japonicum is a species of legume-root nodulating, microsymbiotic nitrogen-fixing bacteria. The species is one of many Gram-negative, rod-shaped bacteria commonly referred to as rhizobia. Within that broad classification, which has three groups, taxonomy studies using DNA sequencing indicate that B. japonicum belongs within homology group II.
αr7 is a family of bacterial small non-coding RNAs with representatives in a broad group of Alphaproteobacterial species from the order Hyphomicrobiales. The first member of this family was found in a Sinorhizobium meliloti 1021 locus located in the chromosome (C). Further homology and structure conservation analysis identified full-length homologs in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species. αr7 RNA species are 134-159 nucleotides (nt) long and share a well defined common secondary structure. αr7 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions (IGRs) of the Alphaproteobacterial genomes.
αr9 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria from the order Hyphomicrobiales. The first member of this family (Smr9C) was found in a Sinorhizobium meliloti 1021 locus located in the chromosome (C). Further homology and structure conservation analysis have identified full-length Smr9C homologs in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species. αr9C RNA species are 144-158 nt long and share a well defined common secondary structure consisting of seven conserved regions. Most of the αr9 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions (IGRs) of the α-proteobacterial genomes.
αr14 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria. The first member of this family (Smr14C2) was found in a Sinorhizobium meliloti 1021 locus located in the chromosome (C). It was later renamed NfeR1 and shown to be highly expressed in salt stress and during the symbiotic interaction on legume roots. Further homology and structure conservation analysis identified 2 other chromosomal copies and 3 plasmidic ones. Moreover, full-length Smr14C homologs have been identified in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species. αr14C RNA species are 115-125 nt long and share a well defined common secondary structure. Most of the αr14 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions (IGRs) of the α-proteobacterial genomes.
αr15 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria from the order Rhizobiales. The first members of this family were found tandemly arranged in the same intergenic region (IGR) of the Sinorhizobium meliloti 1021 chromosome (C). Further homology and structure conservation analysis have identified full-length Smr15C1 and Smr15C2 homologs in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species. The Smr15C1 and Smr15C2 homologs are also encoded in tandem within the same IGR region of Rhizobium and Agrobacterium species, whereas in Brucella species the αr15C loci are spread in the IGRs of Chromosome I. Moreover, this analysis also identified a third αr15 loci in extrachromosomal replicons of the mentioned nitrogen-fixing α-proteobacteria and in the Chromosome II of Brucella species. αr15 RNA species are 99-121 nt long and share a well defined common secondary structure consisting of three stem loops. The transcripts of the αr15 family can be catalogued as trans-acting sRNAs encoded by independent transcription units with recognizable promoter and transcription termination signatures within intergenic regions (IGRs) of the α-proteobacterial genomes.
αr35 is a family of bacterial small non-coding RNAs with representatives in a reduced group of Alphaproteobacteria from the order Hyphomicrobiales. The first member of this family (Smr35B) was found in a Sinorhizobium meliloti 1021 locus located in the symbiotic plasmid B (pSymB). Further homology and structure conservation analysis have identified full-length SmrB35 homologs in other legume symbionts, as well as in the human and plant pathogens Brucella anthropi and Agrobacterium tumefaciens, respectively. αr35 RNA species are 139-142 nt long and share a common secondary structure consisting of two stem loops and a well conserved rho independent terminator. Most of the αr35 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions of the Alphaproteobacterial genomes.
αr45 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria from the order Hyphomicrobiales. The first member of this family (Smr45C) was found in a Sinorhizobium meliloti 1021 locus located in the chromosome (C). Further homology and structure conservation analysis identified homologs in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species, in Bartonella species, in several members of the Xanthobactereacea family, and in some representatives of the Beijerinckiaceae family. αr45C RNA species are 147-153 nt long and share a well defined common secondary structure. All of the αr45 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions (IGRs) of the α-proteobacterial genomes.
Mesorhizobium mediterraneum is a bacterium from the genus Mesorhizobium, which was isolated from root nodule of the Chickpea in Spain. The species Rhizobium mediterraneum was subsequently transferred to Mesorhizobium mediterraneum. This species, along with many other closely related taxa, have been found to promote production of chickpea and other crops worldwide by forming symbiotic relationships.
Streptomyces cattleya is a Gram-positive bacterium which makes cephamycin, penicillin and thienamycin. The bacterium expresses a fluorinase enzyme, and the organism has been used to understand the biosynthesis of fluoroacetate and the antibacterial 4-fluoro-L-threonine. The γ-Glu-βes pathway to biosynthesis of non-traditional amino acids β-ethynylserine (βes) and L-propargylglycine (Pra) was first characterized in this species.
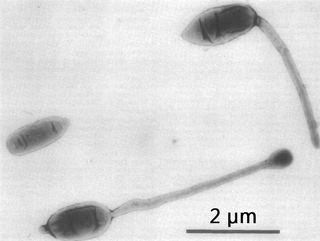
Hyphomicrobium is a genus of Gram-negative, non-spore-forming, rod-shaped bacteria from the family of Hyphomicrobiaceae. It has a large polar or sub-polar filiform prostheca very similar to that of Caulobacter. In addition to having a nutritional function, the prostheca also plays a role in the initiation of DNA replication.
Ensifer medicae is a species of gram-negative, nitrogen-fixing, rod-shaped bacteria. They can be free-living or symbionts of leguminous plants in root nodules. E.medicae was first isolated from root nodules on plants in the genus Medicago. Some strains of E.medicae, like WSM419, are aerobic. They are chemoorganotrophic mesophiles that prefer temperatures around 28 °C. In addition to their primary genome, these organisms also have three known plasmids, sized 1,570,951 bp, 1,245,408 bp and 219,313 bp.
Michael J. Sadowsky is an American microbiologist at the University of Minnesota. He is the director of the BioTechnology Institute and a Professor in the Department of Soil, Water, and Climate. Sadowsky's scientific career spans over 40 years, most of it focused on research studying the nature of bacteria and bacterial genes in ecological settings, with a particular emphasis on soil bacteria that are involved in nitrogen fixation.
References
- ↑ List of Prokaryotic names with Standing in Nomenclature: https://lpsn.dsmz.de/genus/cupriavidus
- ↑ Strainingo of Cupriavidus taiwanensis http://www.straininfo.net/strains/49173 Archived 2021-02-20 at the Wayback Machine
- ↑ UniProt.org https://www.uniprot.org/taxonomy/164546
- ↑ Chen; Laevens, S; Lee, TM; Coenye, T; De Vos, P; Mergeay, M; Vandamme, P; et al. (Sep 2001). "Ralstonia taiwanensis sp. nov., isolated from root nodules of Mimosa species and sputum of a cystic fibrosis patient". Int J Syst Evol Microbiol. 51 (5): 1729–35. doi: 10.1099/00207713-51-5-1729 . PMID 11594603.
- ↑ Applied and Environmental Microbiology http://aem.asm.org/content/77/6/2161
- ↑ Amadou C, Pascal G, Mangenot S, Glew M, Bontemps C, Capela D, Carrère S, Cruveiller S, Dossat C, Lajus A, Marchetti M, Poinsot V, Rouy Z, Servin B, Saad M, Schenowitz C, Barbe V, Batut J, Médigue C, Masson-Boivin C (2008). "Genome sequence of the beta-rhizobium Cupriavidus taiwanensis and comparative genomics of rhizobia". Genome Res. 18 (9): 1472–83. doi:10.1101/gr.076448.108. PMC 2527706 . PMID 18490699.
- ↑ NCBI https://www.ncbi.nlm.nih.gov/gene/6455401
- ↑ Amadou C, Pascal G, Mangenot S, Glew M, Bontemps C, Capela D, Carrère S, Cruveiller S, Dossat C, Lajus A, Marchetti M, Poinsot V, Rouy Z, Servin B, Saad M, Schenowitz C, Barbe V, Batut J, Médigue C, Masson-Boivin C (2008). "Genome sequence of the beta-rhizobium Cupriavidus taiwanensis and comparative genomics of rhizobia". Genome Res. 18 (9): 1472–83. doi:10.1101/gr.076448.108. PMC 2527706 . PMID 18490699.