Adenosine triphosphate (ATP) is a nucleotide that provides energy to drive and support many processes in living cells, such as muscle contraction, nerve impulse propagation, condensate dissolution, and chemical synthesis. Found in all known forms of life, it is often referred to as the "molecular unit of currency" of intracellular energy transfer.

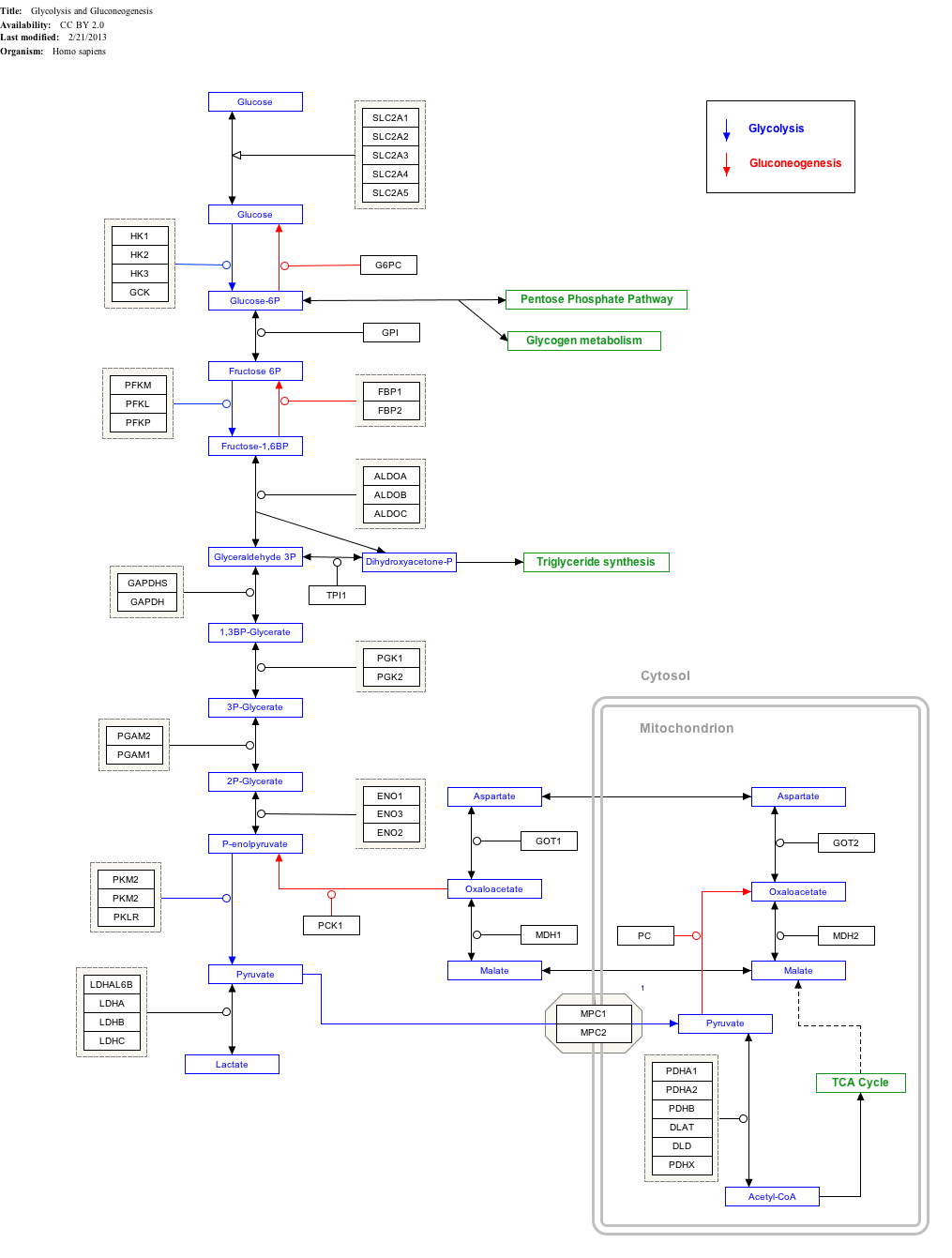
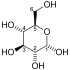
Glycolysis is the metabolic pathway that converts glucose into pyruvate and, in most organisms, occurs in the liquid part of cells. The free energy released in this process is used to form the high-energy molecules adenosine triphosphate (ATP) and reduced nicotinamide adenine dinucleotide (NADH). Glycolysis is a sequence of ten reactions catalyzed by enzymes.

In biochemistry, phosphorylation is the attachment of a phosphate group to a molecule or an ion. This process and its inverse, dephosphorylation, are common in biology. Protein phosphorylation often activates many enzymes.

Tumor hypoxia is the situation where tumor cells have been deprived of oxygen. As a tumor grows, it rapidly outgrows its blood supply, leaving portions of the tumor with regions where the oxygen concentration is significantly lower than in healthy tissues. Hypoxic microenvironments in solid tumors are a result of available oxygen being consumed within 70 to 150 μm of tumor vasculature by rapidly proliferating tumor cells thus limiting the amount of oxygen available to diffuse further into the tumor tissue. In order to support continuous growth and proliferation in challenging hypoxic environments, cancer cells are found to alter their metabolism. Furthermore, hypoxia is known to change cell behavior and is associated with extracellular matrix remodeling and increased migratory and metastatic behavior.

Aldolase A, also known as fructose-bisphosphate aldolase, is an enzyme that in humans is encoded by the ALDOA gene on chromosome 16.

Human iron metabolism is the set of chemical reactions that maintain human homeostasis of iron at the systemic and cellular level. Iron is both necessary to the body and potentially toxic. Controlling iron levels in the body is a critically important part of many aspects of human health and disease. Hematologists have been especially interested in systemic iron metabolism, because iron is essential for red blood cells, where most of the human body's iron is contained. Understanding iron metabolism is also important for understanding diseases of iron overload, such as hereditary hemochromatosis, and iron deficiency, such as iron-deficiency anemia.
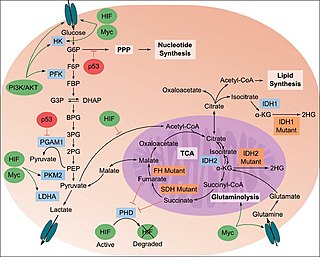
The study of the tumor metabolism, also known as tumor metabolome describes the different characteristic metabolic changes in tumor cells. The characteristic attributes of the tumor metabolome are high glycolytic enzyme activities, the expression of the pyruvate kinase isoenzyme type M2, increased channeling of glucose carbons into synthetic processes, such as nucleic acid, amino acid and phospholipid synthesis, a high rate of pyrimidine and purine de novo synthesis, a low ratio of Adenosine triphosphate and Guanosine triphosphate to Cytidine triphosphate and Uridine triphosphate, low Adenosine monophosphate levels, high glutaminolytic capacities, release of immunosuppressive substances and dependency on methionine.
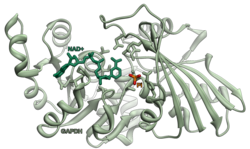
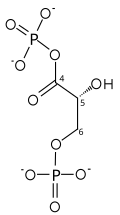
Glyceraldehyde-3-phosphate dehydrogenase (NADP+) (GAPN) is an enzyme that irreversibly catalyzes the oxidation of glyceraldehyde-3-phosphate (GAP) to 3-phosphoglycerate using the reduction of NADP+ to NADPH. GAPN is used in a variant of glycolysis that conserves energy as NADPH rather than as ATP. The NADPH and 3-PG can then be used for synthesis. The most familiar variant of glycolysis uses glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and phosphoglycerate kinase to produce ATP. GAPDH is phosphorylating. GAPN is non-phosphorylating.

Transferrin receptor (TfR) is a carrier protein for transferrin. It is needed for the import of iron into cells and is regulated in response to intracellular iron concentration. It imports iron by internalizing the transferrin-iron complex through receptor-mediated endocytosis. The existence of a receptor for transferrin iron uptake has been recognized since the late 1950s. Earlier two transferrin receptors in humans, transferrin receptor 1 and transferrin receptor 2 had been characterized and until recently cellular iron uptake was believed to occur chiefly via these two well documented transferrin receptors. Both these receptors are transmembrane glycoproteins. TfR1 is a high affinity ubiquitously expressed receptor while expression of TfR2 is restricted to certain cell types and is unaffected by intracellular iron concentrations. TfR2 binds to transferrin with a 25-30 fold lower affinity than TfR1. Although TfR1 mediated iron uptake is the major pathway for iron acquisition by most cells and especially developing erythrocytes, several studies have indicated that the uptake mechanism varies depending upon the cell type. It is also reported that Tf uptake exists independent of these TfRs although the mechanisms are not well characterized. The multifunctional glycolytic enzyme glyceraldehyde 3-phosphate dehydrogenase has been shown to utilize post translational modifications to exhibit higher order moonlighting behavior wherein it switches its function as a holo or apo transferrin receptor leading to either iron delivery or iron export respectively.

The glycerol-3-phosphate shuttle is a mechanism used in skeletal muscle and the brain that regenerates NAD+ from NADH, a by-product of glycolysis. The NADH generated during glycolysis is found in the cytoplasm and must be transported into the mitochondria to enter the oxidative phosphorylation pathway. However, the inner mitochondrial membrane is impermeable to NADH and NAD+ and does not contain a transport system for these electron carriers. Either the glycerol-3-phosphate shuttle pathway or the malate-aspartate shuttle pathway, depending on the tissue of the organism, must be taken to transport electrons from cytoplasmic NADH into the mitochondria. The shuttle consists of the sequential activity of two proteins; Cytoplasmic glycerol-3-phosphate dehydrogenase (cGPD) transfers an electron pair from NADH to dihydroxyacetone phosphate (DHAP), forming glycerol-3-phosphate (G3P) and regenerating NAD+ needed to generate energy via glycolysis. The other protein, mitochondrial glycerol-3-phosphate dehydrogenase (mGPD) catalyzes the oxidation of G3P by FAD, regenerating DHAP in the cytosol and forming FADH2 in the mitochondrial matrix. In mammals, its activity in transporting reducing equivalents across the mitochondrial membrane is considered secondary to the malate-aspartate shuttle.

Fructose-bisphosphate aldolase, often just aldolase, is an enzyme catalyzing a reversible reaction that splits the aldol, fructose 1,6-bisphosphate, into the triose phosphates dihydroxyacetone phosphate (DHAP) and glyceraldehyde 3-phosphate (G3P). Aldolase can also produce DHAP from other (3S,4R)-ketose 1-phosphates such as fructose 1-phosphate and sedoheptulose 1,7-bisphosphate. Gluconeogenesis and the Calvin cycle, which are anabolic pathways, use the reverse reaction. Glycolysis, a catabolic pathway, uses the forward reaction. Aldolase is divided into two classes by mechanism.

Glycerol-3-phosphate dehydrogenase (GPDH) is an enzyme that catalyzes the reversible redox conversion of dihydroxyacetone phosphate to sn-glycerol 3-phosphate.

This family represents the internal ribosome entry site (IRES) of the hepatitis A virus. HAV IRES is a 450 nucleotide long sequence located in the 735 nt long 5’ UTR of Hepatitis A viral RNA genome. IRES elements allow cap and end-independent translation of mRNA in the host cell. The IRES achieves this by mediating the internal initiation of translation by recruiting a ribosomal 40S pre-initiation complex directly to the initiation codon and eliminates the requirement for eukaryotic initiation factor, eIF4F.

Lactate dehydrogenase A (LDHA) is an enzyme which in humans is encoded by the LDHA gene. It is a monomer of Lactate dehydrogenase, which exists as a tetramer. The other main subunit is lactate dehydrogenase B (LDHB).

Protein kinase C iota type is an enzyme that in humans is encoded by the PRKCI gene.

Glyceraldehyde-3-phosphate dehydrogenase, spermatogenic or glyceraldehyde-3-phosphate dehydrogenase, testis-specific is an enzyme that in humans is encoded by the GAPDHS gene.

Aldolase C, fructose-bisphosphate, is an enzyme that, in humans, is encoded by the ALDOC gene on chromosome 17. This gene encodes a member of the class I fructose-bisphosphate aldolase gene family. Expressed specifically in the hippocampus and Purkinje cells of the brain, the encoded protein is a glycolytic enzyme that catalyzes the reversible aldol cleavage of fructose 1,6-bisphosphate and fructose-1-phosphate to dihydroxyacetone phosphate and either glyceraldehyde 3-phosphate or glyceraldehyde, respectively.[provided by RefSeq, Jul 2008]

Protein moonlighting is a phenomenon by which a protein can perform more than one function. It is an excellent example of gene sharing.

The TP53-inducible glycolysis and apoptosis regulator (TIGAR) also known as fructose-2,6-bisphosphatase TIGAR is an enzyme that in humans is encoded by the C12orf5 gene.

1-Arseno-3-phosphoglycerate is a compound produced by the enzyme glyceraldehyde 3-phosphate dehydrogenase, present in high concentrations in many organisms, from glyceraldehyde 3-phosphate and arsenate in the glycolysis pathway. The compound is unstable and hydrolyzes spontaneously to 3-phosphoglycerate, bypassing the energy producing step of glycolysis.