
Ribonucleic acid (RNA) is a polymeric molecule essential in various biological roles in coding, decoding, regulation and expression of genes. RNA and deoxyribonucleic acid (DNA) are nucleic acids. Along with lipids, proteins, and carbohydrates, nucleic acids constitute one of the four major macromolecules essential for all known forms of life. Like DNA, RNA is assembled as a chain of nucleotides, but unlike DNA, RNA is found in nature as a single strand folded onto itself, rather than a paired double strand. Cellular organisms use messenger RNA (mRNA) to convey genetic information that directs synthesis of specific proteins. Many viruses encode their genetic information using an RNA genome.

A non-coding RNA (ncRNA) is an RNA molecule that is not translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally important types of non-coding RNAs include transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs), as well as small RNAs such as microRNAs, siRNAs, piRNAs, snoRNAs, snRNAs, exRNAs, scaRNAs and the long ncRNAs such as Xist and HOTAIR.
Rfam is a database containing information about non-coding RNA (ncRNA) families and other structured RNA elements. It is an annotated, open access database originally developed at the Wellcome Trust Sanger Institute in collaboration with Janelia Farm, and currently hosted at the European Bioinformatics Institute. Rfam is designed to be similar to the Pfam database for annotating protein families.

Small nucleolar RNA R105/R108 refers to a group of related non-coding RNA (ncRNA) molecules which function in the biogenesis of other small nuclear RNAs (snRNAs). These small nucleolar RNAs (snoRNAs) are modifying RNAs and usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis.

Small nucleolar RNA R44/J54/Z268 refers to a group of related non-coding RNA (ncRNA) molecules which function in the biogenesis of other small nuclear RNAs (snRNAs). These small nucleolar RNAs (snoRNAs) are modifying RNAs and usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis.

Small nucleolar RNA RZ107/R87 refers to a group of related non-coding RNA (ncRNA) molecules which function in the biogenesis of other small nuclear RNAs (snRNAs). These small nucleolar RNAs (snoRNAs) are modifying RNAs and usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis.
Small nucleolar RNA J26 is a non-coding RNA (ncRNA) molecule identified in rice which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.

Small nucleolar RNA R12 is a non-coding RNA (ncRNA) molecule which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.
Samm nucleolar RNA R160 is a non-coding RNA (ncRNA) molecule identified in plants which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.
Small nucleolar RNA R41 is a non-coding RNA (ncRNA) molecule identified in plants which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.

Small nucleolar RNA R66 is a non-coding RNA (ncRNA) molecule which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.
Small nucleolar RNA R72 is a non-coding RNA (ncRNA) molecule identified in plants which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.
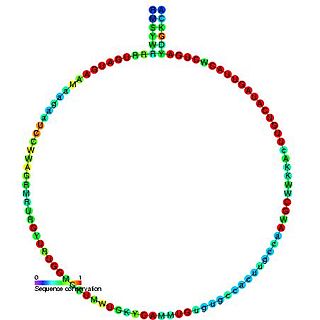
snoRNA U46 is a non-coding RNA (ncRNA) molecule which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.

Small nucleolar RNA Z122 is a non-coding RNA (ncRNA) molecule which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA.

Long non-coding RNAs are a type of RNA, defined as being transcripts with lengths exceeding 200 nucleotides that are not translated into protein. This somewhat arbitrary limit distinguishes long ncRNAs from small non-coding RNAs such as microRNAs (miRNAs), small interfering RNAs (siRNAs), Piwi-interacting RNAs (piRNAs), small nucleolar RNAs (snoRNAs), and other short RNAs. Long intervening/intergenic noncoding RNAs (lincRNAs) are sequences of lncRNA which do not overlap protein-coding genes.

In bioinformatics, miRBase is a biological database that acts as an archive of microRNA sequences and annotations. As of September 2010 it contained information about 15,172 microRNAs. This number has risen to 38,589 by March 2018. The miRBase registry provides a centralised system for assigning new names to microRNA genes.
StarBase is a database for decoding miRNA-mRNA, miRNA-lncRNA, miRNA-sncRNA, miRNA-circRNA, miRNA-pseudogene, protein-lncRNA, protein-ncRNA, protein-mRNA interactions, and ceRNA networks from CLIP-Seq and degradome sequencing data. StarBase provides miRFunction and ceRNAFunction web tools to predict the function of ncRNAs and protein-coding genes from the miRNA and ceRNA regulatory networks. StarBase also developed Pan-Cancer Analysis Platform to decipher Pan-Cancer Analysis Networks of lncRNAs, miRNAs, ceRNAs, and RNA-binding proteins (RBPs) by mining clinical and expression profiles of 14 cancer types from The Cancer Genome Atlas (TCGA) Data Portal.

HOXA11-AS lncRNA is a long non-coding RNA from the antisense strand in the homeobox A. The HOX gene contains four clusters. The sense strand of the HOXA gene codes for proteins. Alternative names for HOXA11-AS lncRNA are: HOXA-AS5, HOXA11S, HOXA11-AS1, HOXA11AS, or NCRNA00076. This gene is 3,885 nucleotides long and resides at chromosome 7 (7p15.2) and is transcribed from an independent gene promoter. Being a lncRNA, it is longer than 200 nucleotides in length, in contrast to regular non-coding RNAs.
CRISPR-Display (CRISP-Disp) is a modification of the CRISPR/Cas9 system for genome editing. The CRISPR/Cas9 system uses a short guide RNA (sgRNA) sequence to direct a Streptococcus pyogenes Cas9 nuclease, acting as a programmable DNA binding protein, to cleave DNA at a site of interest.
















