Related Research Articles

The genetic code is the set of rules used by living cells to translate information encoded within genetic material into proteins. Translation is accomplished by the ribosome, which links proteinogenic amino acids in an order specified by messenger RNA (mRNA), using transfer RNA (tRNA) molecules to carry amino acids and to read the mRNA three nucleotides at a time. The genetic code is highly similar among all organisms and can be expressed in a simple table with 64 entries.

In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mitosis, or meiosis or other types of damage to DNA, which then may undergo error-prone repair, cause an error during other forms of repair, or cause an error during replication. Mutations may also result from insertion or deletion of segments of DNA due to mobile genetic elements.
The quasispecies model is a description of the process of the Darwinian evolution of certain self-replicating entities within the framework of physical chemistry. A quasispecies is a large group or "cloud" of related genotypes that exist in an environment of high mutation rate, where a large fraction of offspring are expected to contain one or more mutations relative to the parent. This is in contrast to a species, which from an evolutionary perspective is a more-or-less stable single genotype, most of the offspring of which will be genetically accurate copies.

Genetic recombination is the exchange of genetic material between different organisms which leads to production of offspring with combinations of traits that differ from those found in either parent. In eukaryotes, genetic recombination during meiosis can lead to a novel set of genetic information that can be further passed on from parents to offspring. Most recombination occurs naturally and can be classified into two types: (1) interchromosomal recombination, occurring through independent assortment of alleles whose loci are on different but homologous chromosomes ; & (2) intrachromosomal recombination, occurring through crossing over.
Molecular evolution is the process of change in the sequence composition of cellular molecules such as DNA, RNA, and proteins across generations. The field of molecular evolution uses principles of evolutionary biology and population genetics to explain patterns in these changes. Major topics in molecular evolution concern the rates and impacts of single nucleotide changes, neutral evolution vs. natural selection, origins of new genes, the genetic nature of complex traits, the genetic basis of speciation, the evolution of development, and ways that evolutionary forces influence genomic and phenotypic changes.

The neutral theory of molecular evolution holds that most evolutionary changes occur at the molecular level, and most of the variation within and between species are due to random genetic drift of mutant alleles that are selectively neutral. The theory applies only for evolution at the molecular level, and is compatible with phenotypic evolution being shaped by natural selection as postulated by Charles Darwin.
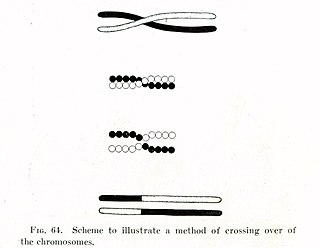
In evolutionary genetics, Muller's ratchet is a process which, in the absence of recombination, results in an accumulation of irreversible deleterious mutations. This happens because in the absence of recombination, and assuming reverse mutations are rare, offspring bear at least as much mutational load as their parents. Muller proposed this mechanism as one reason why sexual reproduction may be favored over asexual reproduction, as sexual organisms benefit from recombination and consequent elimination of deleterious mutations. The negative effect of accumulating irreversible deleterious mutations may not be prevalent in organisms which, while they reproduce asexually, also undergo other forms of recombination. This effect has also been observed in those regions of the genomes of sexual organisms that do not undergo recombination.

Eugene Viktorovich Koonin is a Russian-American biologist and Senior Investigator at the National Center for Biotechnology Information (NCBI). He is a recognised expert in the field of evolutionary and computational biology.
Antigenic drift is a kind of genetic variation in viruses, arising from the accumulation of mutations in the virus genes that code for virus-surface proteins that host antibodies recognize. This results in a new strain of virus particles that is not effectively inhibited by the antibodies that prevented infection by previous strains. This makes it easier for the changed virus to spread throughout a partially immune population. Antigenic drift occurs in both influenza A and influenza B viruses.

Sexual reproduction is an adaptive feature which is common to almost all multicellular organisms and various unicellular organisms. Currently, the adaptive advantage of sexual reproduction is widely regarded as a major unsolved problem in biology. As discussed below, one prominent theory is that sex evolved as an efficient mechanism for producing variation, and this had the advantage of enabling organisms to adapt to changing environments. Another prominent theory, also discussed below, is that a primary advantage of outcrossing sex is the masking of the expression of deleterious mutations. Additional theories concerning the adaptive advantage of sex are also discussed below. Sex does, however, come with a cost. In reproducing asexually, no time nor energy needs to be expended in choosing a mate and, if the environment has not changed, then there may be little reason for variation, as the organism may already be well-adapted. However, very few environments have not changed over the millions of years that reproduction has existed. Hence it is easy to imagine that being able to adapt to changing environment imparts a benefit. Sex also halves the amount of offspring a given population is able to produce. Sex, however, has evolved as the most prolific means of species branching into the tree of life. Diversification into the phylogenetic tree happens much more rapidly via sexual reproduction than it does by way of asexual reproduction.
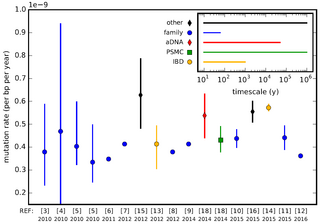
In genetics, the mutation rate is the frequency of new mutations in a single gene or organism over time. Mutation rates are not constant and are not limited to a single type of mutation; there are many different types of mutations. Mutation rates are given for specific classes of mutations. Point mutations are a class of mutations which are changes to a single base. Missense and Nonsense mutations are two subtypes of point mutations. The rate of these types of substitutions can be further subdivided into a mutation spectrum which describes the influence of the genetic context on the mutation rate.

In biology, the word gene can have several different meanings. The Mendelian gene is a basic unit of heredity and the molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protein-coding genes and non-coding genes.

Directed evolution (DE) is a method used in protein engineering that mimics the process of natural selection to steer proteins or nucleic acids toward a user-defined goal. It consists of subjecting a gene to iterative rounds of mutagenesis, selection and amplification. It can be performed in vivo, or in vitro. Directed evolution is used both for protein engineering as an alternative to rationally designing modified proteins, as well as for experimental evolution studies of fundamental evolutionary principles in a controlled, laboratory environment.
A viral quasispecies is a population structure of viruses with a large number of variant genomes. Quasispecies result from high mutation rates as mutants arise continually and change in relative frequency as viral replication and selection proceeds.
The evolution of biological complexity is one important outcome of the process of evolution. Evolution has produced some remarkably complex organisms – although the actual level of complexity is very hard to define or measure accurately in biology, with properties such as gene content, the number of cell types or morphology all proposed as possible metrics.
Gary Bruce Ruvkun is an American molecular biologist at Massachusetts General Hospital and professor of genetics at Harvard Medical School in Boston. Ruvkun discovered the mechanism by which lin-4, the first microRNA (miRNA) discovered by Victor Ambros, regulates the translation of target messenger RNAs via imperfect base-pairing to those targets, and discovered the second miRNA, let-7, and that it is conserved across animal phylogeny, including in humans. These miRNA discoveries revealed a new world of RNA regulation at an unprecedented small size scale, and the mechanism of that regulation. Ruvkun also discovered many features of insulin-like signaling in the regulation of aging and metabolism. He was elected a Member of the American Philosophical Society in 2019.
Experimental evolution studies are a means of testing evolutionary theory under carefully designed, reproducible experiments. Given enough time, space, and money, any organism could be used for experimental evolution studies. However, those with rapid generation times, high mutation rates, large population sizes, and small sizes increase the feasibility of experimental studies in a laboratory context. For these reasons, bacteriophages are especially favored by experimental evolutionary biologists. Bacteriophages, and microbial organisms, can be frozen in stasis, facilitating comparison of evolved strains to ancestors. Additionally, microbes are especially labile from a molecular biologic perspective. Many molecular tools have been developed to manipulate the genetic material of microbial organisms, and because of their small genome sizes, sequencing the full genomes of evolved strains is trivial. Therefore, comparisons can be made for the exact molecular changes in evolved strains during adaptation to novel conditions.

In evolutionary biology, robustness of a biological system is the persistence of a certain characteristic or trait in a system under perturbations or conditions of uncertainty. Robustness in development is known as canalization. According to the kind of perturbation involved, robustness can be classified as mutational, environmental, recombinational, or behavioral robustness etc. Robustness is achieved through the combination of many genetic and molecular mechanisms and can evolve by either direct or indirect selection. Several model systems have been developed to experimentally study robustness and its evolutionary consequences.
An overlapping gene is a gene whose expressible nucleotide sequence partially overlaps with the expressible nucleotide sequence of another gene. In this way, a nucleotide sequence may make a contribution to the function of one or more gene products. Overlapping genes are present in and a fundamental feature of both cellular and viral genomes. The current definition of an overlapping gene varies significantly between eukaryotes, prokaryotes, and viruses. In prokaryotes and viruses overlap must be between coding sequences but not mRNA transcripts, and is defined when these coding sequences share a nucleotide on either the same or opposite strands. In eukaryotes, gene overlap is almost always defined as mRNA transcript overlap. Specifically, a gene overlap in eukaryotes is defined when at least one nucleotide is shared between the boundaries of the primary mRNA transcripts of two or more genes, such that a DNA base mutation at any point of the overlapping region would affect the transcripts of all genes involved. This definition includes 5′ and 3′ untranslated regions (UTRs) along with introns.

Reverse genetics is a method in molecular genetics that is used to help understand the function(s) of a gene by analysing the phenotypic effects caused by genetically engineering specific nucleic acid sequences within the gene. The process proceeds in the opposite direction to forward genetic screens of classical genetics. While forward genetics seeks to find the genetic basis of a phenotype or trait, reverse genetics seeks to find what phenotypes are controlled by particular genetic sequences.
References
- ↑ Chao, L.; Levin, B. R. (1 October 1981). "Structured habitats and the evolution of anticompetitor toxins in bacteria". PNAS. 78 (10): 6324–6328. Bibcode:1981PNAS...78.6324C. doi: 10.1073/pnas.78.10.6324 . PMC 349031 . PMID 7031647.
- ↑ Chao, Lin (29 November 1990). "Fitness of RNA virus decreased by Muller's ratchet". Nature. 348 (6300): 454–455. Bibcode:1990Natur.348..454C. doi:10.1038/348454a0. PMID 2247152. S2CID 4235839.
- ↑ Chao, Lin; Tran, Thu T.; Tran, Thutrang T. (1 November 1997). "The Advantage of Sex in the RNA Virus φ6". Genetics. 147 (3): 953–959. doi:10.1093/genetics/147.3.953. PMC 1208270 . PMID 9383044.
- ↑ Turner, Paul E.; Chao, Lin (1 April 1999). "Prisoner's dilemma in an RNA virus". Nature. 398 (6726): 441–443. Bibcode:1999Natur.398..441T. doi:10.1038/18913. PMID 10201376. S2CID 4422956.
- ↑ Chao, L; Hanley, K. A.; Burch, C. L.; Dahlberg, C; Turner, P. E. (2000). "Kin selection and parasite evolution: Higher and lower virulence with hard and soft selection". The Quarterly Review of Biology. 75 (3): 261–75. doi:10.1086/393499. JSTOR 2665189. PMID 11008699. S2CID 24404768.
- ↑ Burch, Christina L.; Chao, Lin (1 March 1999). "Evolution by Small Steps and Rugged Landscapes in the RNA Virus ϕ6". Genetics. 151 (3): 921–927. doi:10.1093/genetics/151.3.921. PMC 1460516 . PMID 10049911.
- ↑ Chao, LIN (2000). "The Meaning of Life". BioScience. 50 (3): 245–250. doi: 10.1641/0006-3568(2000)050[0245:tmol]2.3.co;2 . JSTOR [0245:tmol2.3.co;2 10.1641/0006-3568(2000)050[0245:tmol]2.3.co;2].
- ↑ "40th Faculty Excellence Awards Honors Outstanding Teaching, Research and Service".