Related Research Articles
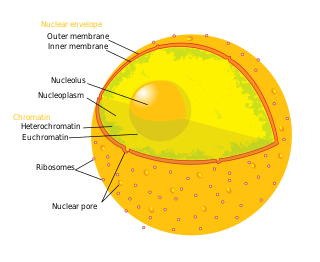
A nuclear pore is a part of a large complex of proteins, known as a nuclear pore complex that spans the nuclear envelope, which is the double membrane surrounding the eukaryotic cell nucleus. There are approximately 1,000 nuclear pore complexes (NPCs) in the nuclear envelope of a vertebrate cell, but this number varies depending on cell type and the stage in the life cycle. The human nuclear pore complex (hNPC) is a 110 megadalton (MDa) structure. The proteins that make up the nuclear pore complex are known as nucleoporins; each NPC contains at least 456 individual protein molecules and is composed of 34 distinct nucleoporin proteins. About half of the nucleoporins typically contain solenoid protein domains—either an alpha solenoid or a beta-propeller fold, or in some cases both as separate structural domains. The other half show structural characteristics typical of "natively unfolded" or intrinsically disordered proteins, i.e. they are highly flexible proteins that lack ordered tertiary structure. These disordered proteins are the FG nucleoporins, so called because their amino-acid sequence contains many phenylalanine–glycine repeats.

The nucleolus is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of signal recognition particles and plays a role in the cell's response to stress. Nucleoli are made of proteins, DNA and RNA, and form around specific chromosomal regions called nucleolar organizing regions. Malfunction of nucleoli can be the cause of several human conditions called "nucleolopathies" and the nucleolus is being investigated as a target for cancer chemotherapy.

Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins are said to produce messenger RNA (mRNA). Other segments of DNA are copied into RNA molecules called non-coding RNAs (ncRNAs). mRNA comprises only 1–3% of total RNA samples. Less than 2% of the human genome can be transcribed into mRNA, while at least 80% of mammalian genomic DNA can be actively transcribed, with the majority of this 80% considered to be ncRNA.

Phage display is a laboratory technique for the study of protein–protein, protein–peptide, and protein–DNA interactions that uses bacteriophages to connect proteins with the genetic information that encodes them. In this technique, a gene encoding a protein of interest is inserted into a phage coat protein gene, causing the phage to "display" the protein on its outside while containing the gene for the protein on its inside, resulting in a connection between genotype and phenotype. These displaying phages can then be screened against other proteins, peptides or DNA sequences, in order to detect interaction between the displayed protein and those other molecules. In this way, large libraries of proteins can be screened and amplified in a process called in vitro selection, which is analogous to natural selection.
A nuclear localization signalorsequence (NLS) is an amino acid sequence that 'tags' a protein for import into the cell nucleus by nuclear transport. Typically, this signal consists of one or more short sequences of positively charged lysines or arginines exposed on the protein surface. Different nuclear localized proteins may share the same NLS. An NLS has the opposite function of a nuclear export signal (NES), which targets proteins out of the nucleus.

Two-hybrid screening is a molecular biology technique used to discover protein–protein interactions (PPIs) and protein–DNA interactions by testing for physical interactions between two proteins or a single protein and a DNA molecule, respectively.
RNA-binding proteins are proteins that bind to the double or single stranded RNA in cells and participate in forming ribonucleoprotein complexes. RBPs contain various structural motifs, such as RNA recognition motif (RRM), dsRNA binding domain, zinc finger and others. They are cytoplasmic and nuclear proteins. However, since most mature RNA is exported from the nucleus relatively quickly, most RBPs in the nucleus exist as complexes of protein and pre-mRNA called heterogeneous ribonucleoprotein particles (hnRNPs). RBPs have crucial roles in various cellular processes such as: cellular function, transport and localization. They especially play a major role in post-transcriptional control of RNAs, such as: splicing, polyadenylation, mRNA stabilization, mRNA localization and translation. Eukaryotic cells express diverse RBPs with unique RNA-binding activity and protein–protein interaction. According to the Eukaryotic RBP Database (EuRBPDB), there are 2961 genes encoding RBPs in humans. During evolution, the diversity of RBPs greatly increased with the increase in the number of introns. Diversity enabled eukaryotic cells to utilize RNA exons in various arrangements, giving rise to a unique RNP (ribonucleoprotein) for each RNA. Although RBPs have a crucial role in post-transcriptional regulation in gene expression, relatively few RBPs have been studied systematically.It has now become clear that RNA–RBP interactions play important roles in many biological processes among organisms.
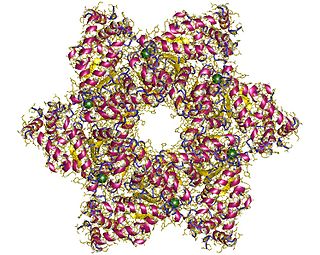
SV40 large T antigen is a hexamer protein that is a dominant-acting oncoprotein derived from the polyomavirus SV40. TAg is capable of inducing malignant transformation of a variety of cell types. The transforming activity of TAg is due in large part to its perturbation of the retinoblastoma (pRb) and p53 tumor suppressor proteins. In addition, TAg binds to several other cellular factors, including the transcriptional co-activators p300 and CBP, which may contribute to its transformation function. Similar proteins from related viruses are known as large tumor antigen in general.
Importin is a type of karyopherin that transports protein molecules from the cell's cytoplasm to the nucleus. It does so by binding to specific recognition sequences, called nuclear localization sequences (NLS).
Heterogeneous nuclear ribonucleoproteins (hnRNPs) are complexes of RNA and protein present in the cell nucleus during gene transcription and subsequent post-transcriptional modification of the newly synthesized RNA (pre-mRNA). The presence of the proteins bound to a pre-mRNA molecule serves as a signal that the pre-mRNA is not yet fully processed and therefore not ready for export to the cytoplasm. Since most mature RNA is exported from the nucleus relatively quickly, most RNA-binding protein in the nucleus exist as heterogeneous ribonucleoprotein particles. After splicing has occurred, the proteins remain bound to spliced introns and target them for degradation.
Nuclear transport refers to the mechanisms by which molecules move across the nuclear membrane of a cell. The entry and exit of large molecules from the cell nucleus is tightly controlled by the nuclear pore complexes (NPCs). Although small molecules can enter the nucleus without regulation, macromolecules such as RNA and proteins require association with transport factors known as nuclear transport receptors, like karyopherins called importins to enter the nucleus and exportins to exit.
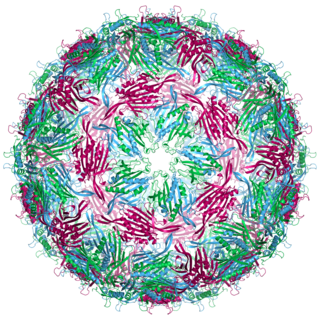
Bacteriophage MS2, commonly called MS2, is an icosahedral, positive-sense single-stranded RNA virus that infects the bacterium Escherichia coli and other members of the Enterobacteriaceae. MS2 is a member of a family of closely related bacterial viruses that includes bacteriophage f2, bacteriophage Qβ, R17, and GA.
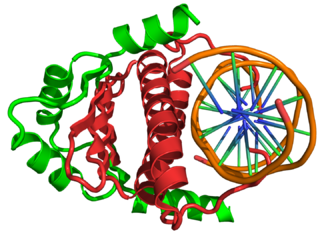
In the field of molecular biology, myocyte enhancer factor-2 (Mef2) proteins are a family of transcription factors which through control of gene expression are important regulators of cellular differentiation and consequently play a critical role in embryonic development. In adult organisms, Mef2 proteins mediate the stress response in some tissues. Mef2 proteins contain both MADS-box and Mef2 DNA-binding domains.

U1 spliceosomal RNA is the small nuclear RNA (snRNA) component of U1 snRNP, an RNA-protein complex that combines with other snRNPs, unmodified pre-mRNA, and various other proteins to assemble a spliceosome, a large RNA-protein molecular complex upon which splicing of pre-mRNA occurs. Splicing, or the removal of introns, is a major aspect of post-transcriptional modification, and takes place only in the nucleus of eukaryotes.

In molecular biology, heat shock factors (HSF), are the transcription factors that regulate the expression of the heat shock proteins. A typical example is the heat shock factor of Drosophila melanogaster.

Rev is a transactivating protein that is essential to the regulation of HIV-1 protein expression. A nuclear localization signal is encoded in the rev gene, which allows the Rev protein to be localized to the nucleus, where it is involved in the export of unspliced and incompletely spliced mRNAs. In the absence of Rev, mRNAs of the HIV-1 late (structural) genes are retained in the nucleus, preventing their translation.
The Streptavidin-Binding Peptide (SBP)-Tag is a 38-amino acid sequence that may be engineered into recombinant proteins. Recombinant proteins containing the SBP-Tag bind to streptavidin and this property may be utilized in specific purification, detection or immobilization strategies.
Importin alpha, or karyopherin alpha refers to a class of adaptor proteins that are involved in the import of proteins into the cell nucleus. They are a sub-family of karyopherin proteins.
CRISPR activation (CRISPRa) is a type of CRISPR tool that uses modified versions of CRISPR effectors without endonuclease activity, with added transcriptional activators on dCas9 or the guide RNAs (gRNAs).
Colin Dingwall is a British biochemist and cell biologist. He is a Fellow of the Royal Society of Biology and a Life Member of Clare Hall, Cambridge UK. Working with Ron Laskey and Sir John Gurdon he established the identified the bipartite nuclear localization sequence (NLS) which is the major signal for protein entry into the nucleus.
References
- ↑ Johansson HE, Liljas L, Uhlenbeck OC (1997). "RNA recognition by the MS2 phage coat protein". Seminars in Virology. 8 (3): 176–185. doi:10.1006/smvy.1997.0120.
- ↑ Bertrand E, Chartrand P, Schaefer M, Shenoy SM, Singer RH, Long RM (October 1998). "Localization of ASH1 mRNA particles in living yeast". Molecular Cell. 2 (4): 437–45. doi: 10.1016/s1097-2765(00)80143-4 . PMID 9809065.
- ↑ Golding I, Paulsson J, Zawilski SM, Cox EC (December 2005). "Real-time kinetics of gene activity in individual bacteria". Cell. 123 (6): 1025–36. doi: 10.1016/j.cell.2005.09.031 . PMID 16360033. S2CID 10319035.
- ↑ Chubb JR, Trcek T, Shenoy SM, Singer RH (May 2006). "Transcriptional pulsing of a developmental gene". Current Biology. 16 (10): 1018–25. doi:10.1016/j.cub.2006.03.092. PMC 4764056 . PMID 16713960.
- 1 2 3 4 5 6 7 "Live and In Color | The Scientist Magazine®". The Scientist. Retrieved 2017-03-12.
- 1 2 3 "Technology". Lucerna, Inc. Retrieved 2017-03-12.
- 1 2 3 Marchese D, de Groot NS, Lorenzo Gotor N, Livi CM, Tartaglia GG (November 2016). "Advances in the characterization of RNA-binding proteins". Wiley Interdisciplinary Reviews: RNA. 7 (6): 793–810. doi:10.1002/wrna.1378. PMC 5113702 . PMID 27503141.
- 1 2 3 Said N, Rieder R, Hurwitz R, Deckert J, Urlaub H, Vogel J (November 2009). "In vivo expression and purification of aptamer-tagged small RNA regulators". Nucleic Acids Research. 37 (20): e133. doi:10.1093/nar/gkp719. PMC 2777422 . PMID 19726584.