
Photosynthesis is a process used by plants and other organisms to convert light energy into chemical energy that, through cellular respiration, can later be released to fuel the organism's activities. Some of this chemical energy is stored in carbohydrate molecules, such as sugars and starches, which are synthesized from carbon dioxide and water – hence the name photosynthesis, from the Greek phōs, "light", and synthesis, "putting together". Most plants, algae, and cyanobacteria perform photosynthesis; such organisms are called photoautotrophs. Photosynthesis is largely responsible for producing and maintaining the oxygen content of the Earth's atmosphere, and supplies most of the energy necessary for life on Earth.
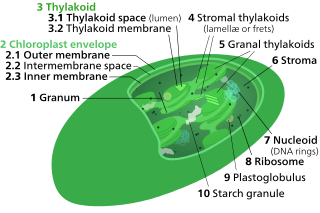
Thylakoids are membrane-bound compartments inside chloroplasts and cyanobacteria. They are the site of the light-dependent reactions of photosynthesis. Thylakoids consist of a thylakoid membrane surrounding a thylakoid lumen. Chloroplast thylakoids frequently form stacks of disks referred to as grana. Grana are connected by intergranal/stromal thylakoids, which join granum stacks together as a single functional compartment.

Photosystems are functional and structural units of protein complexes involved in photosynthesis. Together they carry out the primary photochemistry of photosynthesis: the absorption of light and the transfer of energy and electrons. Photosystems are found in the thylakoid membranes of plants, algae, and cyanobacteria. These membranes are located inside the chloroplasts of plants and algae, and in the cytoplasmic membrane of photosynthetic bacteria. There are two kinds of photosystems: PSI and PSII.

Photosystem II is the first protein complex in the light-dependent reactions of oxygenic photosynthesis. It is located in the thylakoid membrane of plants, algae, and cyanobacteria. Within the photosystem, enzymes capture photons of light to energize electrons that are then transferred through a variety of coenzymes and cofactors to reduce plastoquinone to plastoquinol. The energized electrons are replaced by oxidizing water to form hydrogen ions and molecular oxygen.

Photosystem I is one of two photosystems in the photosynthetic light reactions of algae, plants, and cyanobacteria. Photosystem I is an integral membrane protein complex that uses light energy to catalyze the transfer of electrons across the thylakoid membrane from plastocyanin to ferredoxin. Ultimately, the electrons that are transferred by Photosystem I are used to produce the moderate-energy hydrogen carrier NADPH. The photon energy absorbed by Photosystem I also produces a proton-motive force that is used to generate ATP. PSI is composed of more than 110 cofactors, significantly more than Photosystem II.
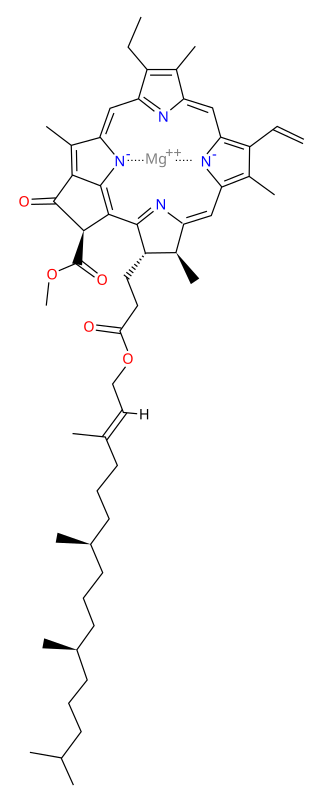
Chlorophyll a is a specific form of chlorophyll used in oxygenic photosynthesis. It absorbs most energy from wavelengths of violet-blue and orange-red light, and it is a poor absorber of green and near-green portions of the spectrum. Chlorophyll does not reflect light but chlorophyll-containing tissues appear green because green light, diffusively reflected by structures like cell walls, becomes enriched in the reflected light. This photosynthetic pigment is essential for photosynthesis in eukaryotes, cyanobacteria and prochlorophytes because of its role as primary electron donor in the electron transport chain. Chlorophyll a also transfers resonance energy in the antenna complex, ending in the reaction center where specific chlorophylls P680 and P700 are located.
Site-directed spin labeling (SDSL) is a technique for investigating the structure and local dynamics of proteins using electron spin resonance. The theory of SDSL is based on the specific reaction of spin labels with amino acids. A spin label's built-in protein structure can be detected by EPR spectroscopy. SDSL is also a useful tool in examinations of the protein folding process.
The oxygen-evolving complex (OEC), also known as the water-splitting complex, is the portion of photosystem II where photo-oxidation of water occurs during the light reactions of photosynthesis. The OEC is surrounded by four core protein subunits of photosystem II at the membrane-lumen interface.

The cytochrome b6f complex is an enzyme found in the thylakoid membrane in chloroplasts of plants, cyanobacteria, and green algae, that catalyzes the transfer of electrons from plastoquinol to plastocyanin. The reaction is analogous to the reaction catalyzed by cytochrome bc1 of the mitochondrial electron transport chain. During photosynthesis, the cytochrome b6f complex is one step along the chain that transfers electrons from Photosystem II to Photosystem I, and at the same time pumps protons into the thylakoid space, contributing to the generation of an electrochemical (energy) gradient that is later used to synthesize ATP from ADP.
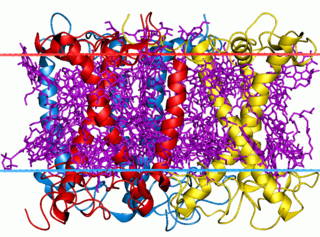
The light-harvesting complex is an array of protein and chlorophyll molecules embedded in the thylakoid membrane of plants and cyanobacteria, which transfer light energy to one chlorophyll a molecule at the reaction center of a photosystem.

A photosynthetic reaction center is a complex of several proteins, pigments and other co-factors that together execute the primary energy conversion reactions of photosynthesis. Molecular excitations, either originating directly from sunlight or transferred as excitation energy via light-harvesting antenna systems, give rise to electron transfer reactions along the path of a series of protein-bound co-factors. These co-factors are light-absorbing molecules (also named chromophores or pigments) such as chlorophyll and pheophytin, as well as quinones. The energy of the photon is used to excite an electron of a pigment. The free energy created is then used, via a chain of nearby electron acceptors, for a transfer of hydrogen atoms (as protons and electrons) from H2O or hydrogen sulfide towards carbon dioxide, eventually producing glucose. These electron transfer steps ultimately result in the conversion of the energy of photons to chemical energy.
A light-harvesting complex consists of a number of chromophores which are complex subunit proteins that may be part of a larger super complex of a photosystem, the functional unit in photosynthesis. It is used by plants and photosynthetic bacteria to collect more of the incoming light than would be captured by the photosynthetic reaction center alone. The light which is captured by the chromophores is capable of exciting molecules from their ground state to a higher energy state, known as the excited state. This excited state does not last very long and is known to be short-lived.

Photoinhibition is light-induced reduction in the photosynthetic capacity of a plant, alga, or cyanobacterium. Photosystem II (PSII) is more sensitive to light than the rest of the photosynthetic machinery, and most researchers define the term as light-induced damage to PSII. In living organisms, photoinhibited PSII centres are continuously repaired via degradation and synthesis of the D1 protein of the photosynthetic reaction center of PSII. Photoinhibition is also used in a wider sense, as dynamic photoinhibition, to describe all reactions that decrease the efficiency of photosynthesis when plants are exposed to light.
In enzymology, a ferredoxin-NADP+ reductase (EC 1.18.1.2) abbreviated FNR, is an enzyme that catalyzes the chemical reaction
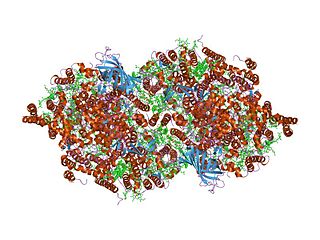
Cytochrome b559 is an important component of Photosystem II.

Photosystem II light-harvesting proteins are the intrinsic transmembrane proteins CP43 (PsbC) and CP47 (PsbB) occurring in the reaction centre of photosystem II (PSII). These polypeptides bind to chlorophyll a and β-Carotene and pass the excitation energy on to the reaction centre.

Light-dependent reactions is jargon for certain photochemical reactions that are involved in photosynthesis, the main process by which plants acquire energy. There are two light dependent reactions, the first occurs at photosystem II (PSII) and the second occurs at photosystem I (PSI),

Bacterial anoxygenic photosynthesis differs from the better known oxygenic photosynthesis in plants by the reductant used and the byproduct generated.

In molecular biology, the PsbZ (Ycf9) is a protein domain, which is low in molecular weight. It is a transmembrane protein and therefore is located in the thylakoid membrane of chloroplasts in cyanobacteria and plants. More specifically, it is located in Photosystem II (PSII) and in the light-harvesting complex II (LHCII). Ycf9 acts as a structural linker, that stabilises the PSII-LHCII supercomplexes. Moreover, the supercomplex fails to form in PsbZ-deficient mutants, providing further evidence to suggest Ycf9's role as a structural linker. This may be caused by a marked decrease in two LHCII antenna proteins, CP26 and CP29, found in PsbZ-deficient mutants, which result in structural changes, as well as functional modifications in PSII.

Alfred William Rutherford FRS is Professor and Chair in Biochemistry of Solar energy in the Department of Life sciences at Imperial College London.













