
Plasmodium malariae is a parasitic protozoan that causes malaria in humans. It is one of several species of Plasmodium parasites that infect other organisms as pathogens, also including Plasmodium falciparum and Plasmodium vivax, responsible for most malarial infection. Found worldwide, it causes a so-called "benign malaria", not nearly as dangerous as that produced by P. falciparum or P. vivax. The signs include fevers that recur at approximately three-day intervals – a quartan fever or quartan malaria – longer than the two-day (tertian) intervals of the other malarial parasite.
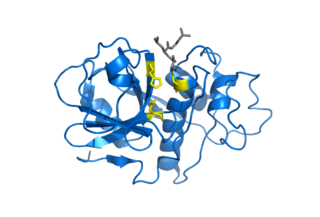
Cysteine proteases, also known as thiol proteases, are hydrolase enzymes that degrade proteins. These proteases share a common catalytic mechanism that involves a nucleophilic cysteine thiol in a catalytic triad or dyad.
Aspergillopepsin I is an enzyme. This enzyme catalyses the following chemical reaction
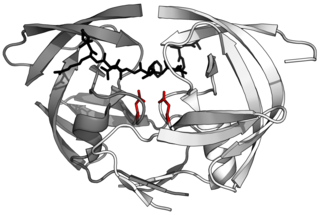
Aspartic proteases are a catalytic type of protease enzymes that use an activated water molecule bound to one or more aspartate residues for catalysis of their peptide substrates. In general, they have two highly conserved aspartates in the active site and are optimally active at acidic pH. Nearly all known aspartyl proteases are inhibited by pepstatin.
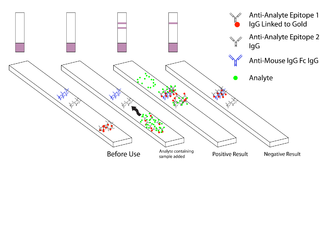
Malaria antigen detection tests are a group of commercially available rapid diagnostic tests of the rapid antigen test type that allow quick diagnosis of malaria by people who are not otherwise skilled in traditional laboratory techniques for diagnosing malaria or in situations where such equipment is not available. There are currently over 20 such tests commercially available. The first malaria antigen suitable as target for such a test was a soluble glycolytic enzyme Glutamate dehydrogenase. None of the rapid tests are currently as sensitive as a thick blood film, nor as cheap. A major drawback in the use of all current dipstick methods is that the result is essentially qualitative. In many endemic areas of tropical Africa, however, the quantitative assessment of parasitaemia is important, as a large percentage of the population will test positive in any qualitative assay.

6-Phosphogluconolactonase (EC 3.1.1.31, 6PGL, PGLS, systematic name 6-phospho-D-glucono-1,5-lactone lactonohydrolase) is a cytosolic enzyme found in all organisms that catalyzes the hydrolysis of 6-phosphogluconolactone to 6-phosphogluconic acid in the oxidative phase of the pentose phosphate pathway:

Ribose-5-phosphate isomerase (Rpi) encoded by the RPIA gene is an enzyme that catalyzes the conversion between ribose-5-phosphate (R5P) and ribulose-5-phosphate (Ru5P). It is a member of a larger class of isomerases which catalyze the interconversion of chemical isomers. It plays a vital role in biochemical metabolism in both the pentose phosphate pathway and the Calvin cycle. The systematic name of this enzyme class is D-ribose-5-phosphate aldose-ketose-isomerase.

Cathepsin D is a protein that in humans is encoded by the CTSD gene. This gene encodes a lysosomal aspartyl protease composed of a protein dimer of disulfide-linked heavy and light chains, both produced from a single protein precursor. Cathepsin D is an aspartic endo-protease that is ubiquitously distributed in lysosomes. The main function of cathepsin D is to degrade proteins and activate precursors of bioactive proteins in pre-lysosomal compartments. This proteinase, which is a member of the peptidase A1 family, has a specificity similar to but narrower than that of pepsin A. Transcription of the CTSD gene is initiated from several sites, including one that is a start site for an estrogen-regulated transcript. Mutations in this gene are involved in the pathogenesis of several diseases, including breast cancer and possibly Alzheimer disease. Homozygous deletion of the CTSD gene leads to early lethality in the postnatal phase. Deficiency of CTSD gene has been reported an underlying cause of neuronal ceroid lipofuscinosis (NCL).

Cathepsin E is an enzyme that in humans is encoded by the CTSE gene. The enzyme is also known as slow-moving proteinase, erythrocyte membrane aspartic proteinase, SMP, EMAP, non-pepsin proteinase, cathepsin D-like acid proteinase, cathepsin E-like acid proteinase, cathepsin D-type proteinase) is an enzyme.

Piperaquine is an antiparasitic drug used in combination with dihydroartemisinin to treat malaria. Piperaquine was developed under the Chinese National Malaria Elimination Programme in the 1960s and was adopted throughout China as a replacement for the structurally similar antimalarial drug chloroquine. Due to widespread parasite resistance to piperaquine, the drug fell out of use as a monotherapy, and is instead used as a partner drug for artemisinin combination therapy. Piperaquine kills parasites by disrupting the detoxification of host heme.
Human genetic resistance to malaria refers to inherited changes in the DNA of humans which increase resistance to malaria and result in increased survival of individuals with those genetic changes. The existence of these genotypes is likely due to evolutionary pressure exerted by parasites of the genus Plasmodium which cause malaria. Since malaria infects red blood cells, these genetic changes are most common alterations to molecules essential for red blood cell function, such as hemoglobin or other cellular proteins or enzymes of red blood cells. These alterations generally protect red blood cells from invasion by Plasmodium parasites or replication of parasites within the red blood cell.
The N-end rule is a rule that governs the rate of protein degradation through recognition of the N-terminal residue of proteins. The rule states that the N-terminal amino acid of a protein determines its half-life. The rule applies to both eukaryotic and prokaryotic organisms, but with different strength, rules, and outcome. In eukaryotic cells, these N-terminal residues are recognized and targeted by ubiquitin ligases, mediating ubiquitination thereby marking the protein for degradation. The rule was initially discovered by Alexander Varshavsky and co-workers in 1986. However, only rough estimations of protein half-life can be deduced from this 'rule', as N-terminal amino acid modification can lead to variability and anomalies, whilst amino acid impact can also change from organism to organism. Other degradation signals, known as degrons, can also be found in sequence.
Nepenthesin is an aspartic protease of plant origin that has so far been identified in the pitcher secretions of Nepenthes and in the leaves of Drosera peltata. It is similar to pepsin, but differs in that it also cleaves on either side of Asp residues and at Lys┼Arg. While more pH and temperature stable than porcine pepsin A, it is considerably less stable in urea or guanidine hydrochloride. It is the only known protein with such a stability profile.

Aspergilloglutamic peptidase, also called aspergillopepsin II is a proteolytic enzyme. The enzyme was previously thought be an aspartic protease, but it was later shown to be a glutamic protease with a catalytic Glu residue at the active site, and was therefore renamed aspergilloglutamic peptidase.
Penicillopepsin is an enzyme. This enzyme catalyses the following chemical reaction
Endothiapepsin is an enzyme. This enzyme catalyses the following chemical reaction
Mucorpepsin is an enzyme. This enzyme catalyses the following chemical reaction

Scytalidocarboxyl peptidase B, also known as Scytalidoglutamic peptidase and Scytalidopepsin B is a proteolytic enzyme. It was previously thought to be an aspartic protease, but determination of its molecular structure showed it to belong a novel group of proteases, glutamic protease.
Plasmepsin I is an enzyme. This enzyme catalyses the following chemical reaction

Plasmepsin II (EC 3.4.23.39, aspartic hemoglobinase II, PFAPD) is an enzyme. This enzyme catalyses the following chemical reaction












