Related Research Articles

Proteomics is the large-scale study of proteins. Proteins are vital parts of living organisms, with many functions such as the formation of structural fibers of muscle tissue, enzymatic digestion of food, or synthesis and replication of DNA. In addition, other kinds of proteins include antibodies that protect an organism from infection, and hormones that send important signals throughout the body.
Glycomics is the comprehensive study of glycomes, including genetic, physiologic, pathologic, and other aspects. Glycomics "is the systematic study of all glycan structures of a given cell type or organism" and is a subset of glycobiology. The term glycomics is derived from the chemical prefix for sweetness or a sugar, "glyco-", and was formed to follow the omics naming convention established by genomics and proteomics.
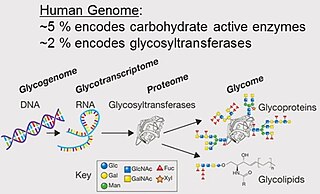
A glycome is the entire complement or complete set of all sugars, whether free or chemically bound in more complex molecules, of an organism. An alternative definition is the entirety of carbohydrates in a cell. The glycome may in fact be one of the most complex entities in nature. "Glycomics, analogous to genomics and proteomics, is the systematic study of all glycan structures of a given cell type or organism" and is a subset of glycobiology.
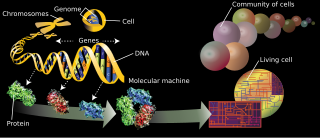
The branches of science known informally as omics are various disciplines in biology whose names end in the suffix -omics, such as genomics, proteomics, metabolomics, metagenomics, phenomics and transcriptomics. Omics aims at the collective characterization and quantification of pools of biological molecules that translate into the structure, function, and dynamics of an organism or organisms.

Metabolomics is the scientific study of chemical processes involving metabolites, the small molecule substrates, intermediates, and products of cell metabolism. Specifically, metabolomics is the "systematic study of the unique chemical fingerprints that specific cellular processes leave behind", the study of their small-molecule metabolite profiles. The metabolome represents the complete set of metabolites in a biological cell, tissue, organ, or organism, which are the end products of cellular processes. Messenger RNA (mRNA), gene expression data, and proteomic analyses reveal the set of gene products being produced in the cell, data that represents one aspect of cellular function. Conversely, metabolic profiling can give an instantaneous snapshot of the physiology of that cell, and thus, metabolomics provides a direct "functional readout of the physiological state" of an organism. There are indeed quantifiable correlations between the metabolome and the other cellular ensembles, which can be used to predict metabolite abundances in biological samples from, for example mRNA abundances. One of the ultimate challenges of systems biology is to integrate metabolomics with all other -omics information to provide a better understanding of cellular biology.

The metabolome refers to the complete set of small-molecule chemicals found within a biological sample. The biological sample can be a cell, a cellular organelle, an organ, a tissue, a tissue extract, a biofluid or an entire organism. The small molecule chemicals found in a given metabolome may include both endogenous metabolites that are naturally produced by an organism as well as exogenous chemicals that are not naturally produced by an organism.

Personalized medicine, also referred to as precision medicine, is a medical model that separates people into different groups—with medical decisions, practices, interventions and/or products being tailored to the individual patient based on their predicted response or risk of disease. The terms personalized medicine, precision medicine, stratified medicine and P4 medicine are used interchangeably to describe this concept though some authors and organisations use these expressions separately to indicate particular nuances.
In biomedical contexts, a biomarker, or biological marker, is a measurable indicator of some biological state or condition. Biomarkers are often measured and evaluated using blood, urine, or soft tissues to examine normal biological processes, pathogenic processes, or pharmacologic responses to a therapeutic intervention. Biomarkers are used in many scientific fields.
In medicine, a biomarker is a measurable indicator of the severity or presence of some disease state. It may be defined as a "cellular, biochemical or molecular alteration in cells, tissues or fluids that can be measured and evaluated to indicate normal biological processes, pathogenic processes, or pharmacological responses to a therapeutic intervention." More generally a biomarker is anything that can be used as an indicator of a particular disease state or some other physiological state of an organism. According to the WHO, the indicator may be chemical, physical, or biological in nature - and the measurement may be functional, physiological, biochemical, cellular, or molecular.
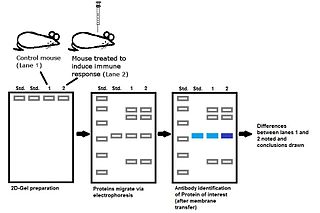
Immunoproteomics is the study of large sets of proteins (proteomics) involved in the immune response.
Surface-enhanced laser desorption/ionization (SELDI) is a soft ionization method in mass spectrometry (MS) used for the analysis of protein mixtures. It is a variation of matrix-assisted laser desorption/ionization (MALDI). In MALDI, the sample is mixed with a matrix material and applied to a metal plate before irradiation by a laser, whereas in SELDI, proteins of interest in a sample become bound to a surface before MS analysis. The sample surface is a key component in the purification, desorption, and ionization of the sample. SELDI is typically used with time-of-flight (TOF) mass spectrometers and is used to detect proteins in tissue samples, blood, urine, or other clinical samples, however, SELDI technology can potentially be used in any application by simply modifying the sample surface.
Glycoproteomics is a branch of proteomics that identifies, catalogs, and characterizes proteins containing carbohydrates as a result of post-translational modifications. Glycosylation is the most common post-translational modification of proteins, but continues to be the least studied on the proteome level. Mass spectrometry (MS) is an analytical technique used to improve the study of these proteins on the proteome level. Glycosylation contributes to several concerted biological mechanisms essential to maintaining physiological function. The study of the glycosylation of proteins is important to understanding certain diseases, like cancer, because a connection between a change in glycosylation and these diseases has been discovered. To study this post-translational modification of proteins, advanced mass spectrometry techniques based on glycoproteomics have been developed to help in terms of therapeutic applications and the discovery of biomarkers.

MALDI mass spectrometry imaging (MALDI-MSI) is the use of matrix-assisted laser desorption ionization as a mass spectrometry imaging technique in which the sample, often a thin tissue section, is moved in two dimensions while the mass spectrum is recorded. Advantages, like measuring the distribution of a large amount of analytes at one time without destroying the sample, make it a useful method in tissue-based study.
Mass spectrometry imaging (MSI) is a technique used in mass spectrometry to visualize the spatial distribution of molecules, as biomarkers, metabolites, peptides or proteins by their molecular masses. After collecting a mass spectrum at one spot, the sample is moved to reach another region, and so on, until the entire sample is scanned. By choosing a peak in the resulting spectra that corresponds to the compound of interest, the MS data is used to map its distribution across the sample. This results in pictures of the spatially resolved distribution of a compound pixel by pixel. Each data set contains a veritable gallery of pictures because any peak in each spectrum can be spatially mapped. Despite the fact that MSI has been generally considered a qualitative method, the signal generated by this technique is proportional to the relative abundance of the analyte. Therefore, quantification is possible, when its challenges are overcome. Although widely used traditional methodologies like radiochemistry and immunohistochemistry achieve the same goal as MSI, they are limited in their abilities to analyze multiple samples at once, and can prove to be lacking if researchers do not have prior knowledge of the samples being studied. Most common ionization technologies in the field of MSI are DESI imaging, MALDI imaging, secondary ion mass spectrometry imaging and Nanoscale SIMS (NanoSIMS).

Adipose differentiation-related protein, also known as perilipin 2, ADRP or adipophilin, is a protein which belongs to the perilipin (PAT) family of cytoplasmic lipid droplet (CLD)–binding proteins. In humans it is encoded by the ADFP gene. This protein surrounds the lipid droplet along with phospholipids and is involved in assisting the storage of neutral lipids within the lipid droplets.

Proteogenomics is a field of biological research that utilizes a combination of proteomics, genomics, and transcriptomics to aid in the discovery and identification of peptides. Proteogenomics is used to identify new peptides by comparing MS/MS spectra against a protein database that has been derived from genomic and transcriptomic information. Proteogenomics often refers to studies that use proteomic information, often derived from mass spectrometry, to improve gene annotations. The utilization of both proteomics and genomics data alongside advances in the availability and power of spectrographic and chromatographic technology led to the emergence of proteogenomics as its own field in 2004.
Secretomics is a type of proteomics which involves the analysis of the secretome—all the secreted proteins of a cell, tissue or organism. Secreted proteins are involved in a variety of physiological processes, including cell signaling and matrix remodeling, but are also integral to invasion and metastasis of malignant cells. Secretomics has thus been especially important in the discovery of biomarkers for cancer and understanding molecular basis of pathogenesis. The analysis of the insoluble fraction of the secretome has been termed matrisomics.
Pharmacometabolomics, also known as pharmacometabonomics, is a field which stems from metabolomics, the quantification and analysis of metabolites produced by the body. It refers to the direct measurement of metabolites in an individual's bodily fluids, in order to predict or evaluate the metabolism of pharmaceutical compounds, and to better understand the pharmacokinetic profile of a drug. Alternatively, pharmacometabolomics can be applied to measure metabolite levels following the administration of a pharmaceutical compound, in order to monitor the effects of the compound on certain metabolic pathways(pharmacodynamics). This provides detailed mapping of drug effects on metabolism and the pathways that are implicated in mechanism of variation of response to treatment. In addition, the metabolic profile of an individual at baseline (metabotype) provides information about how individuals respond to treatment and highlights heterogeneity within a disease state. All three approaches require the quantification of metabolites found in bodily fluids and tissue, such as blood or urine, and can be used in the assessment of pharmaceutical treatment options for numerous disease states.

A cancer biomarker refers to a substance or process that is indicative of the presence of cancer in the body. A biomarker may be a molecule secreted by a tumor or a specific response of the body to the presence of cancer. Genetic, epigenetic, proteomic, glycomic, and imaging biomarkers can be used for cancer diagnosis, prognosis, and epidemiology. Ideally, such biomarkers can be assayed in non-invasively collected biofluids like blood or serum.

In the field of cellular biology, single-cell analysis and subcellular analysis is the study of genomics, transcriptomics, proteomics, metabolomics and cell–cell interactions at the single cell level. The concept of single-cell analysis originated in the 1970s. Before the discovery of heterogeneity, single-cell analysis mainly referred to the analysis or manipulation of an individual cell in a bulk population of cells at a particular condition using optical or electronic microscope. To date, due to the heterogeneity seen in both eukaryotic and prokaryotic cell populations, analyzing a single cell makes it possible to discover mechanisms not seen when studying a bulk population of cells. Technologies such as fluorescence-activated cell sorting (FACS) allow the precise isolation of selected single cells from complex samples, while high throughput single cell partitioning technologies, enable the simultaneous molecular analysis of hundreds or thousands of single unsorted cells; this is particularly useful for the analysis of transcriptome variation in genotypically identical cells, allowing the definition of otherwise undetectable cell subtypes. The development of new technologies is increasing our ability to analyze the genome and transcriptome of single cells, as well as to quantify their proteome and metabolome. Mass spectrometry techniques have become important analytical tools for proteomic and metabolomic analysis of single cells. Recent advances have enabled quantifying thousands of protein across hundreds of single cells, and thus make possible new types of analysis. In situ sequencing and fluorescence in situ hybridization (FISH) do not require that cells be isolated and are increasingly being used for analysis of tissues.
References
- ↑ Hathout, Yetrib (2007). "Approaches to the study of the cell secretome". Expert Review of Proteomics. 4 (2): 239–48. doi:10.1586/14789450.4.2.239. PMID 17425459.
- ↑ Singer, E. A.; Penson, D. F.; Palapattu, G. S. (2007). "PSA Screening and Elderly Men". JAMA. 297 (9): 949, author reply 949–50. doi:10.1001/jama.297.9.949-a. PMID 17341705.
- ↑ Crawford, D. C.; Sanders, C. L.; Qin, X.; Smith, J. D.; Shephard, C.; Wong, M.; Witrak, L.; Rieder, M. J.; Nickerson, D. A. (2006). "Genetic Variation is Associated with C-Reactive Protein Levels in the Third National Health and Nutrition Examination Survey". Circulation. 114 (23): 2458–65. doi: 10.1161/CIRCULATIONAHA.106.615740 . PMID 17101857.
- ↑ Jacobs, Jon M.; Adkins, Joshua N.; Qian, Wei-Jun; Liu, Tao; Shen, Yufeng; Camp, David G.; Smith, Richard D. (2005). "Utilizing Human Blood Plasma for Proteomic Biomarker Discovery†". Journal of Proteome Research. 4 (4): 1073–85. doi:10.1021/pr0500657. PMID 16083256.
- ↑ Anderson, NL; Anderson, NG (2002). "The human plasma proteome: history, character, and diagnostic prospects". Molecular & Cellular Proteomics. 1 (11): 845–67. doi: 10.1074/mcp.R200007-MCP200 . PMID 12488461.
- ↑ He, YD (2006). "Genomic approach to biomarker identification and its recent applications". Cancer Biomarkers. 2 (3–4): 103–33. PMID 17192065.
- ↑ Loukopoulos P, Shibata T, Katoh H, et al. (March 2007). "Genome-wide array-based comparative genomic hybridization analysis of pancreatic adenocarcinoma: identification of genetic indicators that predict patient outcome". Cancer Sci. 98 (3): 392–400. doi: 10.1111/j.1349-7006.2007.00395.x . PMID 17233815.
- ↑ "NIH Researchers Identify Striking Genomic Signature Shared by 5 Types of Cancer - ScienceNewsline". www.sciencenewsline.com. Retrieved 2016-04-24.[ permanent dead link ]
- ↑ Bergman, Nina; Bergquist, Jonas (2014). "Recent developments in proteomic methods and disease biomarkers". The Analyst. 139 (16): 3836–3851. doi:10.1039/C4AN00627E. ISSN 0003-2654.
- ↑ Aizpurua-Olaizola, O.; Toraño, J. Sastre; Falcon-Perez, J.M.; Williams, C.; Reichardt, N.; Boons, G.-J. (2018). "Mass spectrometry for glycan biomarker discovery". TrAC Trends in Analytical Chemistry. 100: 7–14. doi:10.1016/j.trac.2017.12.015.
- ↑ "Ex Vivo Blood Stimulation in Biomarker Discovery". Archived from the original on 2009-11-29. Retrieved 2009-10-23.