
Biotin (or vitamin B7) is one of the B vitamins. It is involved in a wide range of metabolic processes, both in humans and in other organisms, primarily related to the utilization of fats, carbohydrates, and amino acids. The name biotin, borrowed from the German Biotin, derives from the Ancient Greek word βίοτος (bíotos; 'life') and the suffix "-in" (a suffix used in chemistry usually to indicate 'forming').
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amine group is bonded to the carboxylic group of another amino acid, making it a chain. That leaves a free carboxylic group at one end of the peptide, called the C-terminus, and a free amine group on the other end called the N-terminus. By convention, peptide sequences are written N-terminus to C-terminus, left to right (in LTR writing systems). This correlates the translation direction to the text direction, because when a protein is translated from messenger RNA, it is created from the N-terminus to the C-terminus, as amino acids are added to the carboxyl end of the protein.

A ubiquitin ligase is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another thing by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases regulates diverse areas such as cell trafficking, DNA repair, and signaling and is of profound importance in cell biology. E3 ligases are also key players in cell cycle control, mediating the degradation of cyclins, as well as cyclin dependent kinase inhibitor proteins. The human genome encodes over 600 putative E3 ligases, allowing for tremendous diversity in substrates.
In biochemistry, biotinylation is the process of covalently attaching biotin to a protein, nucleic acid or other molecule. Biotinylation is rapid, specific and is unlikely to disturb the natural function of the molecule due to the small size of biotin. Biotin binds to streptavidin and avidin with an extremely high affinity, fast on-rate, and high specificity, and these interactions are exploited in many areas of biotechnology to isolate biotinylated molecules of interest. Biotin-binding to streptavidin and avidin is resistant to extremes of heat, pH and proteolysis, making capture of biotinylated molecules possible in a wide variety of environments. Also, multiple biotin molecules can be conjugated to a protein of interest, which allows binding of multiple streptavidin, avidin or neutravidin protein molecules and increases the sensitivity of detection of the protein of interest. There is a large number of biotinylation reagents available that exploit the wide range of possible labelling methods. Due to the strong affinity between biotin and streptavidin, the purification of biotinylated proteins has been a widely used approach to identify protein-protein interactions and post-translational events such as ubiquitylation in molecular biology.
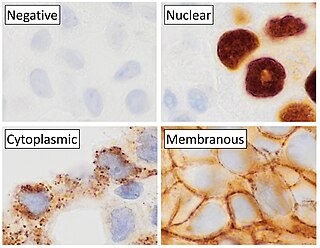
Immunohistochemistry (IHC) is the most common application of immunostaining. It involves the process of selectively identifying antigens (proteins) in cells of a tissue section by exploiting the principle of antibodies binding specifically to antigens in biological tissues. IHC takes its name from the roots "immuno", in reference to antibodies used in the procedure, and "histo", meaning tissue. Albert Coons conceptualized and first implemented the procedure in 1941.
A protein microarray is a high-throughput method used to track the interactions and activities of proteins, and to determine their function, and determining function on a large scale. Its main advantage lies in the fact that large numbers of proteins can be tracked in parallel. The chip consists of a support surface such as a glass slide, nitrocellulose membrane, bead, or microtitre plate, to which an array of capture proteins is bound. Probe molecules, typically labeled with a fluorescent dye, are added to the array. Any reaction between the probe and the immobilised protein emits a fluorescent signal that is read by a laser scanner. Protein microarrays are rapid, automated, economical, and highly sensitive, consuming small quantities of samples and reagents. The concept and methodology of protein microarrays was first introduced and illustrated in antibody microarrays in 1983 in a scientific publication and a series of patents. The high-throughput technology behind the protein microarray was relatively easy to develop since it is based on the technology developed for DNA microarrays, which have become the most widely used microarrays.
Chemical biology is a scientific discipline between the fields of chemistry and biology. The discipline involves the application of chemical techniques, analysis, and often small molecules produced through synthetic chemistry, to the study and manipulation of biological systems. In contrast to biochemistry, which involves the study of the chemistry of biomolecules and regulation of biochemical pathways within and between cells, chemical biology deals with chemistry applied to biology.

Streptavidin is a 52 kDa protein (tetramer) purified from the bacterium Streptomyces avidinii. Streptavidin homo-tetramers have an extraordinarily high affinity for biotin. With a dissociation constant (Kd) on the order of ≈10−14 mol/L, the binding of biotin to streptavidin is one of the strongest non-covalent interactions known in nature. Streptavidin is used extensively in molecular biology and bionanotechnology due to the streptavidin-biotin complex's resistance to organic solvents, denaturants, detergents, proteolytic enzymes, and extremes of temperature and pH.
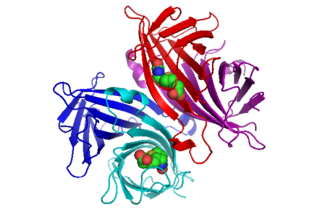
Avidin is a tetrameric biotin-binding protein produced in the oviducts of birds, reptiles and amphibians and deposited in the whites of their eggs. Dimeric members of the avidin family are also found in some bacteria. In chicken egg white, avidin makes up approximately 0.05% of total protein (approximately 1800 μg per egg). The tetrameric protein contains four identical subunits (homotetramer), each of which can bind to biotin (Vitamin B7, vitamin H) with a high degree of affinity and specificity. The dissociation constant of the avidin-biotin complex is measured to be KD ≈ 10−15 M, making it one of the strongest known non-covalent bonds.
Biotinidase, also known as biotinase, is an enzyme that in humans is encoded by the BTD gene.

Meir Wilchek is an Israeli biochemist. He is a professor at the Weizmann Institute of Science.

Isobaric tags for relative and absolute quantitation (iTRAQ) is an isobaric labeling method used in quantitative proteomics by tandem mass spectrometry to determine the amount of proteins from different sources in a single experiment. It uses stable isotope labeled molecules that can be covalent bonded to the N-terminus and side chain amines of proteins.
Alice Yen-Ping Ting is Taiwanese-born American chemist. She is a professor of Genetics, of Biology, and by courtesy, of Chemistry at Stanford University. She is also a Chan Zuckerberg Biohub investigator.
The Streptavidin-Binding Peptide (SBP)-Tag is a 38-amino acid sequence that may be engineered into recombinant proteins. Recombinant proteins containing the SBP-Tag bind to streptavidin and this property may be utilized in specific purification, detection or immobilization strategies.
Terminal amine isotopic labeling of substrates (TAILS) is a method in quantitative proteomics that identifies the protein content of samples based on N-terminal fragments of each protein and detects differences in protein abundance among samples.
Mono-BOC-cystamine is a tert-butyloxycarbonyl (BOC) derivative of cystamine used as crosslinker in biotechnology and molecular biology applications. This compound was originally reported by Hansen et al.

O-GlcNAc is a reversible enzymatic post-translational modification that is found on serine and threonine residues of nucleocytoplasmic proteins. The modification is characterized by a β-glycosidic bond between the hydroxyl group of serine or threonine side chains and N-acetylglucosamine (GlcNAc). O-GlcNAc differs from other forms of protein glycosylation: (i) O-GlcNAc is not elongated or modified to form more complex glycan structures, (ii) O-GlcNAc is almost exclusively found on nuclear and cytoplasmic proteins rather than membrane proteins and secretory proteins, and (iii) O-GlcNAc is a highly dynamic modification that turns over more rapidly than the proteins which it modifies. O-GlcNAc is conserved across metazoans.
Chemoproteomics entails a broad array of techniques used to identify and interrogate protein-small molecule interactions. Chemoproteomics complements phenotypic drug discovery, a paradigm that aims to discover lead compounds on the basis of alleviating a disease phenotype, as opposed to target-based drug discovery, in which lead compounds are designed to interact with predetermined disease-driving biological targets. As phenotypic drug discovery assays do not provide confirmation of a compound's mechanism of action, chemoproteomics provides valuable follow-up strategies to narrow down potential targets and eventually validate a molecule's mechanism of action. Chemoproteomics also attempts to address the inherent challenge of drug promiscuity in small molecule drug discovery by analyzing protein-small molecule interactions on a proteome-wide scale. A major goal of chemoproteomics is to characterize the interactome of drug candidates to gain insight into mechanisms of off-target toxicity and polypharmacology.
BLESS, also known as breaks labeling, enrichment on streptavidin and next-generation sequencing, is a method used to detect genome-wide double-strand DNA damage. In contrast to chromatin immunoprecipitation (ChIP)-based methods of identifying DNA double-strand breaks (DSBs) by labeling DNA repair proteins, BLESS utilizes biotinylated DNA linkers to directly label genomic DNA in situ which allows for high-specificity enrichment of samples on streptavidin beads and the subsequent sequencing-based DSB mapping to nucleotide resolution.
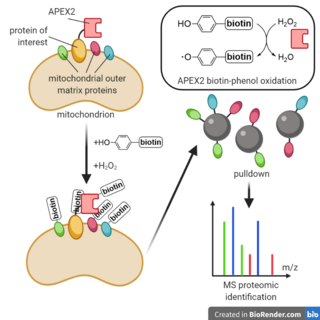
Enzyme-catalyzed proximity labeling (PL), also known as proximity-based labeling, is a laboratory technique that labels biomolecules, usually proteins or RNA, proximal to a protein of interest. By creating a gene fusion in a living cell between the protein of interest and an engineered labeling enzyme, biomolecules spatially proximal to the protein of interest can then be selectively marked with biotin for pulldown and analysis. Proximity labeling has been used for identifying the components of novel cellular structures and for determining protein-protein interaction partners, among other applications.









