
Bioinformatics is an interdisciplinary field of science that develops methods and software tools for understanding biological data, especially when the data sets are large and complex. Bioinformatics uses biology, chemistry, physics, computer science, computer programming, information engineering, mathematics and statistics to analyze and interpret biological data. The subsequent process of analyzing and interpreting data is referred to as computational biology.

In computer science, evolutionary computation is a family of algorithms for global optimization inspired by biological evolution, and the subfield of artificial intelligence and soft computing studying these algorithms. In technical terms, they are a family of population-based trial and error problem solvers with a metaheuristic or stochastic optimization character.
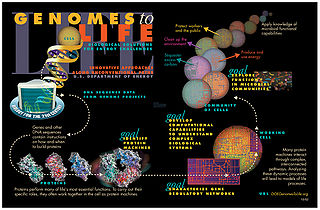
Systems biology is the computational and mathematical analysis and modeling of complex biological systems. It is a biology-based interdisciplinary field of study that focuses on complex interactions within biological systems, using a holistic approach to biological research.

Eugene Viktorovich Koonin is a Russian-American biologist and Senior Investigator at the National Center for Biotechnology Information (NCBI). He is a recognised expert in the field of evolutionary and computational biology.
In artificial intelligence, artificial immune systems (AIS) are a class of computationally intelligent, rule-based machine learning systems inspired by the principles and processes of the vertebrate immune system. The algorithms are typically modeled after the immune system's characteristics of learning and memory for use in problem-solving.
Vasant G. Honavar is an Indian-American computer scientist, and artificial intelligence, machine learning, big data, data science, causal inference, knowledge representation, bioinformatics and health informatics researcher and professor.
David B. Fogel is a pioneer in evolutionary computation.
Dr. Lawrence Jerome Fogel was a pioneer in evolutionary computation and human factors analysis. He is known as the inventor of active noise cancellation and the father of evolutionary programming. His scientific career spanned nearly six decades and included electrical engineering, aerospace engineering, communication theory, human factors research, information processing, cybernetics, biotechnology, artificial intelligence, and computer science.
Paulien Hogeweg is a Dutch theoretical biologist and complex systems researcher studying biological systems as dynamic information processing systems at many interconnected levels. In 1970, together with Ben Hesper, she defined the term bioinformatics as "the study of informatic processes in biotic systems".

Lawrence E. Hunter is a Professor and Director of the Center for Computational Pharmacology and of the Computational Bioscience Program at the University of Colorado School of Medicine and Professor of Computer Science at the University of Colorado Boulder. He is an internationally known scholar, focused on computational biology, knowledge-driven extraction of information from the primary biomedical literature, the semantic integration of knowledge resources in molecular biology, and the use of knowledge in the analysis of high-throughput data, as well as for his foundational work in computational biology, which led to the genesis of the major professional organization in the field and two international conferences.
Computational Resources for Drug Discovery (CRDD) is an important module of the in silico module of Open Source for Drug Discovery (OSDD). The CRDD web portal provides computer resources related to drug discovery on a single platform. It caters to researchers researching computer-aided drug design, provides computational resources, hosts a discussion forum, and has resources related to drug discovery, predicting inhibitors, and predicting the ADME-Tox properties of molecules. One of the major objectives of CRDD is to promote open source software in the field of cheminformatics and pharmacoinformatics.
Christopher Boyce Burge is Professor of Biology and Biological Engineering at Massachusetts Institute of Technology.
Osnat Penn, born in 1981, is an Israeli computational biologist whose work focuses on molecular evolution, cell research, and immunoinformatics. She is the third Israeli scientist in three years to win the UNESCO-L’Oréal fellowship, which she received in 2013 for her work on the genetic origins of autism. Penn is currently a postdoctorate fellow at the University of Washington in Seattle, where she has been working since 2012.

Debora S. Marks is a researcher in computational biology and a Professor of Systems Biology at Harvard Medical School. Her research uses computational approaches to address a variety of biological problems.
Transcriptomics technologies are the techniques used to study an organism's transcriptome, the sum of all of its RNA transcripts. The information content of an organism is recorded in the DNA of its genome and expressed through transcription. Here, mRNA serves as a transient intermediary molecule in the information network, whilst non-coding RNAs perform additional diverse functions. A transcriptome captures a snapshot in time of the total transcripts present in a cell. Transcriptomics technologies provide a broad account of which cellular processes are active and which are dormant. A major challenge in molecular biology is to understand how a single genome gives rise to a variety of cells. Another is how gene expression is regulated.

Ujjwal Maulik is an Indian computer scientist and a professor. He is the former chair of the Department of Computer Science and Engineering at Jadavpur University, Kolkata, West Bengal, India. He also held the position of the principal-in-charge and the head of the Department of Computer Science and Engineering at Kalyani Government Engineering College.
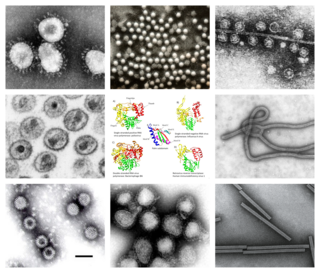
Riboviria is a realm of viruses that includes all viruses that use a homologous RNA-dependent polymerase for replication. It includes RNA viruses that encode an RNA-dependent RNA polymerase, as well as reverse-transcribing viruses that encode an RNA-dependent DNA polymerase. RNA-dependent RNA polymerase (RdRp), also called RNA replicase, produces RNA from RNA. RNA-dependent DNA polymerase (RdDp), also called reverse transcriptase (RT), produces DNA from RNA. These enzymes are essential for replicating the viral genome and transcribing viral genes into messenger RNA (mRNA) for translation of viral proteins.
libRoadRunner is a C/C++ software library that supports simulation of SBML based models.. It uses LLVM to generate extremely high-performance code and is the fastest SBML-based simulator currently available. Its main purpose is for use as a reusable library that can be hosted by other applications, particularly on large compute clusters for doing parameter optimization where performance is critical. It also has a set of Python bindings that allow it to be easily used from Python as well as a set of bindings for Julia.







