Ribonuclease is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within the EC 2.7 and 3.1 classes of enzymes.

Ribonuclease H is a family of non-sequence-specific endonuclease enzymes that catalyze the cleavage of RNA in an RNA/DNA substrate via a hydrolytic mechanism. Members of the RNase H family can be found in nearly all organisms, from bacteria to archaea to eukaryotes.

Angiogenin (ANG) also known as ribonuclease 5 is a small 123 amino acid protein that in humans is encoded by the ANG gene. Angiogenin is a potent stimulator of new blood vessels through the process of angiogenesis. Ang hydrolyzes cellular RNA, resulting in modulated levels of protein synthesis and interacts with DNA causing a promoter-like increase in the expression of rRNA. Ang is associated with cancer and neurological disease through angiogenesis and through activating gene expression that suppresses apoptosis.
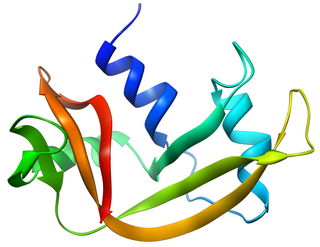
Bovine pancreatic ribonuclease, also often referred to as bovine pancreatic ribonuclease A or simply RNase A, is a pancreatic ribonuclease enzyme that cleaves single-stranded RNA. Bovine pancreatic ribonuclease is one of the classic model systems of protein science. Two Nobel Prizes in Chemistry have been awarded in recognition of work on bovine pancreatic ribonuclease: in 1972, the Prize was awarded to Christian Anfinsen for his work on protein folding and to Stanford Moore and William Stein for their work on the relationship between the protein's structure and its chemical mechanism; in 1984, the Prize was awarded to Robert Bruce Merrifield for development of chemical synthesis of proteins.

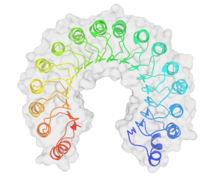
A leucine-rich repeat (LRR) is a protein structural motif that forms an α/β horseshoe fold. It is composed of repeating 20–30 amino acid stretches that are unusually rich in the hydrophobic amino acid leucine. These tandem repeats commonly fold together to form a solenoid protein domain, termed leucine-rich repeat domain. Typically, each repeat unit has beta strand-turn-alpha helix structure, and the assembled domain, composed of many such repeats, has a horseshoe shape with an interior parallel beta sheet and an exterior array of helices. One face of the beta sheet and one side of the helix array are exposed to solvent and are therefore dominated by hydrophilic residues. The region between the helices and sheets is the protein's hydrophobic core and is tightly sterically packed with leucine residues.

Frederic Middlebrook Richards, commonly referred to as Fred Richards, was an American biochemist and biophysicist known for solving the pioneering crystal structure of the ribonuclease S enzyme in 1967 and for defining the concept of solvent-accessible surface. He contributed many key experimental and theoretical results and developed new methods, garnering over 20,000 journal citations in several quite distinct research areas. In addition to the protein crystallography and biochemistry of ribonuclease S, these included solvent accessibility and internal packing of proteins, the first side-chain rotamer library, high-pressure crystallography, new types of chemical tags such as biotin/avidin, the nuclear magnetic resonance (NMR) chemical shift index, and structural and biophysical characterization of the effects of mutations.

In molecular biology, a protein domain is a region of a protein's polypeptide chain that is self-stabilizing and that folds independently from the rest. Each domain forms a compact folded three-dimensional structure. Many proteins consist of several domains, and a domain may appear in a variety of different proteins. Molecular evolution uses domains as building blocks and these may be recombined in different arrangements to create proteins with different functions. In general, domains vary in length from between about 50 amino acids up to 250 amino acids in length. The shortest domains, such as zinc fingers, are stabilized by metal ions or disulfide bridges. Domains often form functional units, such as the calcium-binding EF hand domain of calmodulin. Because they are independently stable, domains can be "swapped" by genetic engineering between one protein and another to make chimeric proteins.
Molecular binding is an attractive interaction between two molecules that results in a stable association in which the molecules are in close proximity to each other. It is formed when atoms or molecules bind together by sharing of electrons. It often, but not always, involves some chemical bonding.

Pancreatic ribonuclease family is a superfamily of pyrimidine-specific endonucleases found in high quantity in the pancreas of certain mammals and of some reptiles.
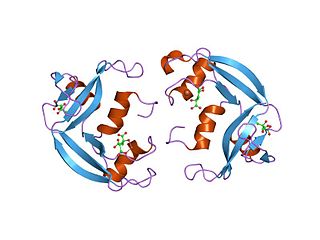
Eosinophil cationic protein (ECP) also known as ribonuclease 3 is a basic protein located in the eosinophil primary matrix. In humans, the eosinophil cationic protein is encoded by the RNASE3 gene.

Eosinophil-derived neurotoxin is an enzyme that in humans is encoded by the RNASE2 gene.

Ribonuclease pancreatic is an enzyme that in humans is encoded by the RNASE1 gene.

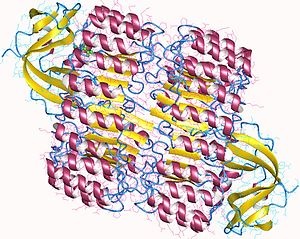
Ribonuclease inhibitor is an enzyme that in humans is encoded by the RNH1 gene.

Ribonuclease 4 is an enzyme that in humans is encoded by the RNASE4 gene.

Leucine-rich repeat neuronal protein 3, also known as neuronal leucine-rich repeat protein 3 (NLRR-3), is a protein that in humans is encoded by the LRRN3 gene.

Leucine rich repeat containing 40 (LRRC40) is a protein that in humans is encoded by the LRRC40 gene.
RiAFP refers to an antifreeze protein (AFP) produced by the Rhagium inquisitor longhorned beetle. It is a type V antifreeze protein with a molecular weight of 12.8 kDa; this type of AFP is noted for its hyperactivity. R. inquisitor is a freeze-avoidant species, meaning that, due to its AFP, R. inquisitor prevents its body fluids from freezing altogether. This contrasts with freeze-tolerant species, whose AFPs simply depress levels of ice crystal formation in low temperatures. Whereas most insect antifreeze proteins contain cysteines at least every sixth residue, as well as varying numbers of 12- or 13-mer repeats of 8.3-12.5kDa, RiAFP is notable for containing only one disulfide bridge. This property of RiAFP makes it particularly attractive for recombinant expression and biotechnological applications.
Ribonuclease E is a bacterial ribonuclease that participates in the processing of ribosomal RNA and the chemical degradation of bulk cellular RNA.

LRRIQ3, which is also known as LRRC44, is a protein that in humans is encoded by the LRRIQ3 gene. It is predominantly expressed in the testes, and is linked to a number of diseases.