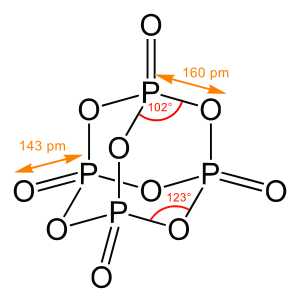
A chemical bond is a lasting attraction between atoms or ions that enables the formation of molecules, crystals, and other structures. The bond may result from the electrostatic force between oppositely charged ions as in ionic bonds, or through the sharing of electrons as in covalent bonds. The strength of chemical bonds varies considerably; there are "strong bonds" or "primary bonds" such as covalent, ionic and metallic bonds, and "weak bonds" or "secondary bonds" such as dipole–dipole interactions, the London dispersion force, and hydrogen bonding.
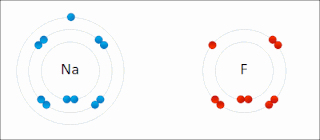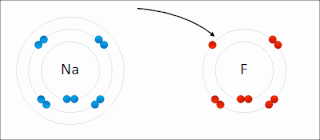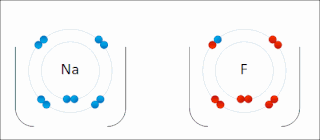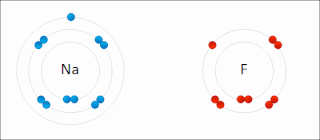
Ionic bonding is a type of chemical bonding that involves the electrostatic attraction between oppositely charged ions, or between two atoms with sharply different electronegativities, and is the primary interaction occurring in ionic compounds. It is one of the main types of bonding, along with covalent bonding and metallic bonding. Ions are atoms with an electrostatic charge. Atoms that gain electrons make negatively charged ions. Atoms that lose electrons make positively charged ions. This transfer of electrons is known as electrovalence in contrast to covalence. In the simplest case, the cation is a metal atom and the anion is a nonmetal atom, but these ions can be more complex, e.g. molecular ions like NH+
4 or SO2−
4. In simpler words, an ionic bond results from the transfer of electrons from a metal to a non-metal to obtain a full valence shell for both atoms.
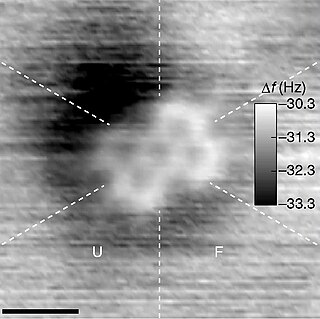
A molecule is a group of two or more atoms held together by attractive forces known as chemical bonds; depending on context, the term may or may not include ions which satisfy this criterion. In quantum physics, organic chemistry, and biochemistry, the distinction from ions is dropped and molecule is often used when referring to polyatomic ions.

X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angles and intensities of these diffracted beams, a crystallographer can produce a three-dimensional picture of the density of electrons within the crystal. From this electron density, the positions of the atoms in the crystal can be determined, as well as their chemical bonds, crystallographic disorder, and various other information.

The structural formula of a chemical compound is a graphic representation of the molecular structure, showing how the atoms are possibly arranged in the real three-dimensional space. The chemical bonding within the molecule is also shown, either explicitly or implicitly. Unlike other chemical formula types, which have a limited number of symbols and are capable of only limited descriptive power, structural formulas provide a more complete geometric representation of the molecular structure. For example, many chemical compounds exist in different isomeric forms, which have different enantiomeric structures but the same molecular formula. There are multiple types of ways to draw these structural formulas such as: Lewis Structures, condensed formulas, skeletal formulas, Newman projections, Cyclohexane conformations, Haworth projections, and Fischer projections.

Structural bioinformatics is the branch of bioinformatics that is related to the analysis and prediction of the three-dimensional structure of biological macromolecules such as proteins, RNA, and DNA. It deals with generalizations about macromolecular 3D structures such as comparisons of overall folds and local motifs, principles of molecular folding, evolution, binding interactions, and structure/function relationships, working both from experimentally solved structures and from computational models. The term structural has the same meaning as in structural biology, and structural bioinformatics can be seen as a part of computational structural biology. The main objective of structural bioinformatics is the creation of new methods of analysing and manipulating biological macromolecular data in order to solve problems in biology and generate new knowledge.
In chemistry, molecular orbital theory is a method for describing the electronic structure of molecules using quantum mechanics. It was proposed early in the 20th century.

Molecular geometry is the three-dimensional arrangement of the atoms that constitute a molecule. It includes the general shape of the molecule as well as bond lengths, bond angles, torsional angles and any other geometrical parameters that determine the position of each atom.
Gas electron diffraction (GED) is one of the applications of electron diffraction techniques. The target of this method is the determination of the structure of gaseous molecules, i.e., the geometrical arrangement of the atoms from which a molecule is built up. GED is one of two experimental methods to determine the structure of free molecules, undistorted by intermolecular forces, which are omnipresent in the solid and liquid state. The determination of accurate molecular structures by GED studies is fundamental for an understanding of structural chemistry.

Valence shell electron pair repulsion (VSEPR) theory, is a model used in chemistry to predict the geometry of individual molecules from the number of electron pairs surrounding their central atoms. It is also named the Gillespie-Nyholm theory after its two main developers, Ronald Gillespie and Ronald Nyholm.

Nuclear magnetic resonance spectroscopy, most commonly known as NMR spectroscopy or magnetic resonance spectroscopy (MRS), is a spectroscopic technique based on re-orientation of atomic nuclei with non-zero nuclear spins in an external magnetic field. This re-orientation occurs with absorption of electromagnetic radiation in the radio frequency region from roughly 4 to 900 MHz, which depends on the isotopic nature of the nucleus and increased proportionally to the strength of the external magnetic field. Notably, the resonance frequency of each NMR-active nucleus depends on its chemical environment. As a result, NMR spectra provide information about individual functional groups present in the sample, as well about connections between nearby nuclei in the same molecule. As the NMR spectra are unique or highly characteristic to individual compounds and functional groups, NMR spectroscopy is one of the most important methods to identify molecular structures, particulary of organic compounds.
In molecular geometry, bond length or bond distance is defined as the average distance between nuclei of two bonded atoms in a molecule. It is a transferable property of a bond between atoms of fixed types, relatively independent of the rest of the molecule.
Nuclear magnetic resonance spectroscopy of proteins is a field of structural biology in which NMR spectroscopy is used to obtain information about the structure and dynamics of proteins, and also nucleic acids, and their complexes. The field was pioneered by Richard R. Ernst and Kurt Wüthrich at the ETH, and by Ad Bax, Marius Clore, Angela Gronenborn at the NIH, and Gerhard Wagner at Harvard University, among others. Structure determination by NMR spectroscopy usually consists of several phases, each using a separate set of highly specialized techniques. The sample is prepared, measurements are made, interpretive approaches are applied, and a structure is calculated and validated.
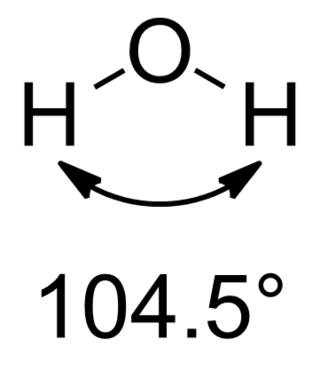
In chemistry, Bent's rule describes and explains the relationship between the orbital hybridization and the electronegativities of substituents. The rule was stated by Henry A. Bent as follows:
Atomic s character concentrates in orbitals directed toward electropositive substituents.
Physical organic chemistry, a term coined by Louis Hammett in 1940, refers to a discipline of organic chemistry that focuses on the relationship between chemical structures and reactivity, in particular, applying experimental tools of physical chemistry to the study of organic molecules. Specific focal points of study include the rates of organic reactions, the relative chemical stabilities of the starting materials, reactive intermediates, transition states, and products of chemical reactions, and non-covalent aspects of solvation and molecular interactions that influence chemical reactivity. Such studies provide theoretical and practical frameworks to understand how changes in structure in solution or solid-state contexts impact reaction mechanism and rate for each organic reaction of interest.

Molecular models of DNA structures are representations of the molecular geometry and topology of deoxyribonucleic acid (DNA) molecules using one of several means, with the aim of simplifying and presenting the essential, physical and chemical, properties of DNA molecular structures either in vivo or in vitro. These representations include closely packed spheres made of plastic, metal wires for skeletal models, graphic computations and animations by computers, artistic rendering. Computer molecular models also allow animations and molecular dynamics simulations that are very important for understanding how DNA functions in vivo.
Nucleic acid NMR is the use of nuclear magnetic resonance spectroscopy to obtain information about the structure and dynamics of nucleic acid molecules, such as DNA or RNA. It is useful for molecules of up to 100 nucleotides, and as of 2003, nearly half of all known RNA structures had been determined by NMR spectroscopy.
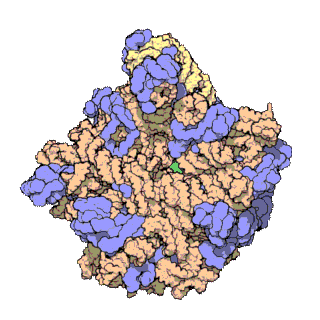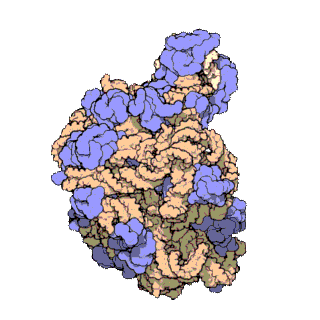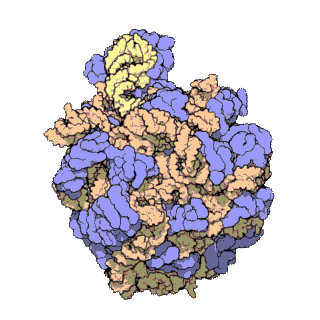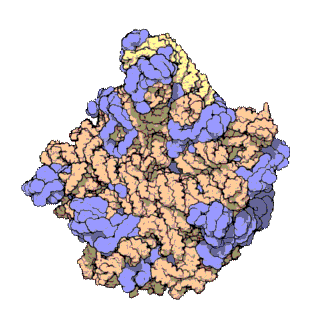
The term macromolecular assembly (MA) refers to massive chemical structures such as viruses and non-biologic nanoparticles, cellular organelles and membranes and ribosomes, etc. that are complex mixtures of polypeptide, polynucleotide, polysaccharide or other polymeric macromolecules. They are generally of more than one of these types, and the mixtures are defined spatially, and with regard to their underlying chemical composition and structure. Macromolecules are found in living and nonliving things, and are composed of many hundreds or thousands of atoms held together by covalent bonds; they are often characterized by repeating units. Assemblies of these can likewise be biologic or non-biologic, though the MA term is more commonly applied in biology, and the term supramolecular assembly is more often applied in non-biologic contexts. MAs of macromolecules are held in their defined forms by non-covalent intermolecular interactions, and can be in either non-repeating structures, or in repeating linear, circular, spiral, or other patterns. The process by which MAs are formed has been termed molecular self-assembly, a term especially applied in non-biologic contexts. A wide variety of physical/biophysical, chemical/biochemical, and computational methods exist for the study of MA; given the scale of MAs, efforts to elaborate their composition and structure and discern mechanisms underlying their functions are at the forefront of modern structure science.
Structural chemistry is a part of chemistry and deals with spatial structures of molecules and solids. For structure elucidation a range of different methods is used. One has to distinguish between methods that elucidate solely the connectivity between atoms (constitution) and such that provide precise three dimensional information such as atom coordinates, bond lengths and angles and torsional angles.

Linnett double-quartet theory (LDQ) is a method of describing the bonding in molecules which involves separating the electrons depending on their spin, placing them into separate 'spin tetrahedra' to minimise the Pauli repulsions between electrons of the same spin. Introduced by J. W. Linnett in his 1961 monograph and 1964 book, this method expands on the electron dot structures pioneered by G. N. Lewis. While the theory retains the requirement for fulfilling the octet rule, it dispenses with the need to force electrons into coincident pairs. Instead, the theory stipulates that the four electrons of a given spin should maximise the distances between each other, resulting in a net tetrahedral electronic arrangement that is the fundamental molecular building block of the theory.