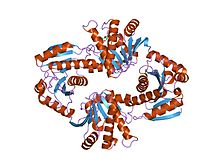
Nicotinamide adenine dinucleotide (NAD) is a coenzyme central to metabolism. Found in all living cells, NAD is called a dinucleotide because it consists of two nucleotides joined through their phosphate groups. One nucleotide contains an adenine nucleobase and the other nicotinamide. NAD exists in two forms: an oxidized and reduced form, abbreviated as NAD+ and NADH (H for hydrogen), respectively.

Poly (ADP-ribose) polymerase (PARP) is a family of proteins involved in a number of cellular processes such as DNA repair, genomic stability, and programmed cell death.

Histone-modifying enzymes are enzymes involved in the modification of histone substrates after protein translation and affect cellular processes including gene expression. To safely store the eukaryotic genome, DNA is wrapped around four core histone proteins, which then join to form nucleosomes. These nucleosomes further fold together into highly condensed chromatin, which renders the organism's genetic material far less accessible to the factors required for gene transcription, DNA replication, recombination and repair. Subsequently, eukaryotic organisms have developed intricate mechanisms to overcome this repressive barrier imposed by the chromatin through histone modification, a type of post-translational modification which typically involves covalently attaching certain groups to histone residues. Once added to the histone, these groups elicit either a loose and open histone conformation, euchromatin, or a tight and closed histone conformation, heterochromatin. Euchromatin marks active transcription and gene expression, as the light packing of histones in this way allows entry for proteins involved in the transcription process. As such, the tightly packed heterochromatin marks the absence of current gene expression.

ADP-ribosylation is the addition of one or more ADP-ribose moieties to a protein. It is a reversible post-translational modification that is involved in many cellular processes, including cell signaling, DNA repair, gene regulation and apoptosis. Improper ADP-ribosylation has been implicated in some forms of cancer. It is also the basis for the toxicity of bacterial compounds such as cholera toxin, diphtheria toxin, and others.

Protein C-ets-1 is a protein that in humans is encoded by the ETS1 gene. The protein encoded by this gene belongs to the ETS family of transcription factors.

Poly [ADP-ribose] polymerase 1 (PARP-1) also known as NAD+ ADP-ribosyltransferase 1 or poly[ADP-ribose] synthase 1 is an enzyme that in humans is encoded by the PARP1 gene. It is the most abundant of the PARP family of enzymes, accounting for 90% of the NAD+ used by the family. PARP1 is mostly present in cell nucleus, but cytosolic fraction of this protein was also reported.

ADP-ribosylation factor 1 is a protein that in humans is encoded by the ARF1 gene.

ADP-ribosylation factor-binding protein GGA1 is a protein that in humans is encoded by the GGA1 gene.

Telomeric repeat-binding factor 1 is a protein that in humans is encoded by the TERF1 gene.

ADP-ribosylation factor 3 is a protein that in humans is encoded by the ARF3 gene.
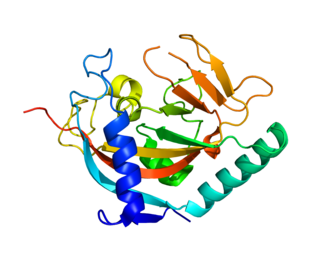
Tankyrase, also known as tankyrase 1, is an enzyme that in humans is encoded by the TNKS gene. It inhibits the binding of TERF1 to telomeric DNA. Tankyrase attracts substantial interest in cancer research through its interaction with AXIN1 and AXIN2, which are negative regulators of pro-oncogenic β-catenin signaling. Importantly, activity in the β-catenin destruction complex can be increased by tankyrase inhibitors and thus such inhibitors are a potential therapeutic option to reduce the growth of β-catenin-dependent cancers.

Histone H1.1 is a protein that in humans is encoded by the HIST1H1A gene.

Poly [ADP-ribose] polymerase 4 is an enzyme that in humans is encoded by the PARP4 gene.

Tankyrase-2 is an enzyme that in humans is encoded by the TNKS2 gene.
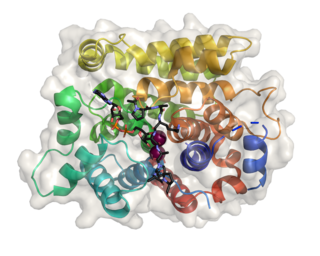
(ADP-ribosyl)hydrolase 3 (ARH3) is an enzyme that in humans is encoded by the ADPRHL2 gene (also called ADPRS). This enzyme reverses the proteins’ post-translational addition of ADP-ribose to serine residues as part of the DNA damage response The enzyme is also known to cleave poly(ADP-ribose) polymers, 1''-O-acetyl-ADP-ribose and alpha-NAD+

A toxin-antitoxin system is a set of two or more closely linked genes that together encode both a "toxin" protein and a corresponding "antitoxin". Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK).

In molecular biology, the (ADP-ribosyl)hydrolase (ARH) family contains enzymes which catalyses the hydrolysis of ADP-ribosyl modifications from proteins, nucleic acids and small molecules.

Poly (ADP-ribose) glycohydrolase is an enzyme that in humans is encoded by the PARG gene.
Parthanatos is a form of programmed cell death that is distinct from other cell death processes such as necrosis and apoptosis. While necrosis is caused by acute cell injury resulting in traumatic cell death and apoptosis is a highly controlled process signalled by apoptotic intracellular signals, parthanatos is caused by the accumulation of Poly(ADP ribose) (PAR) and the nuclear translocation of apoptosis-inducing factor (AIF) from mitochondria. Parthanatos is also known as PARP-1 dependent cell death. PARP-1 mediates parthanatos when it is over-activated in response to extreme genomic stress and synthesizes PAR which causes nuclear translocation of AIF. Parthanatos is involved in diseases that afflict hundreds of millions of people worldwide. Well known diseases involving parthanatos include Parkinson's disease, stroke, heart attack, and diabetes. It also has potential use as a treatment for ameliorating disease and various medical conditions such as diabetes and obesity.