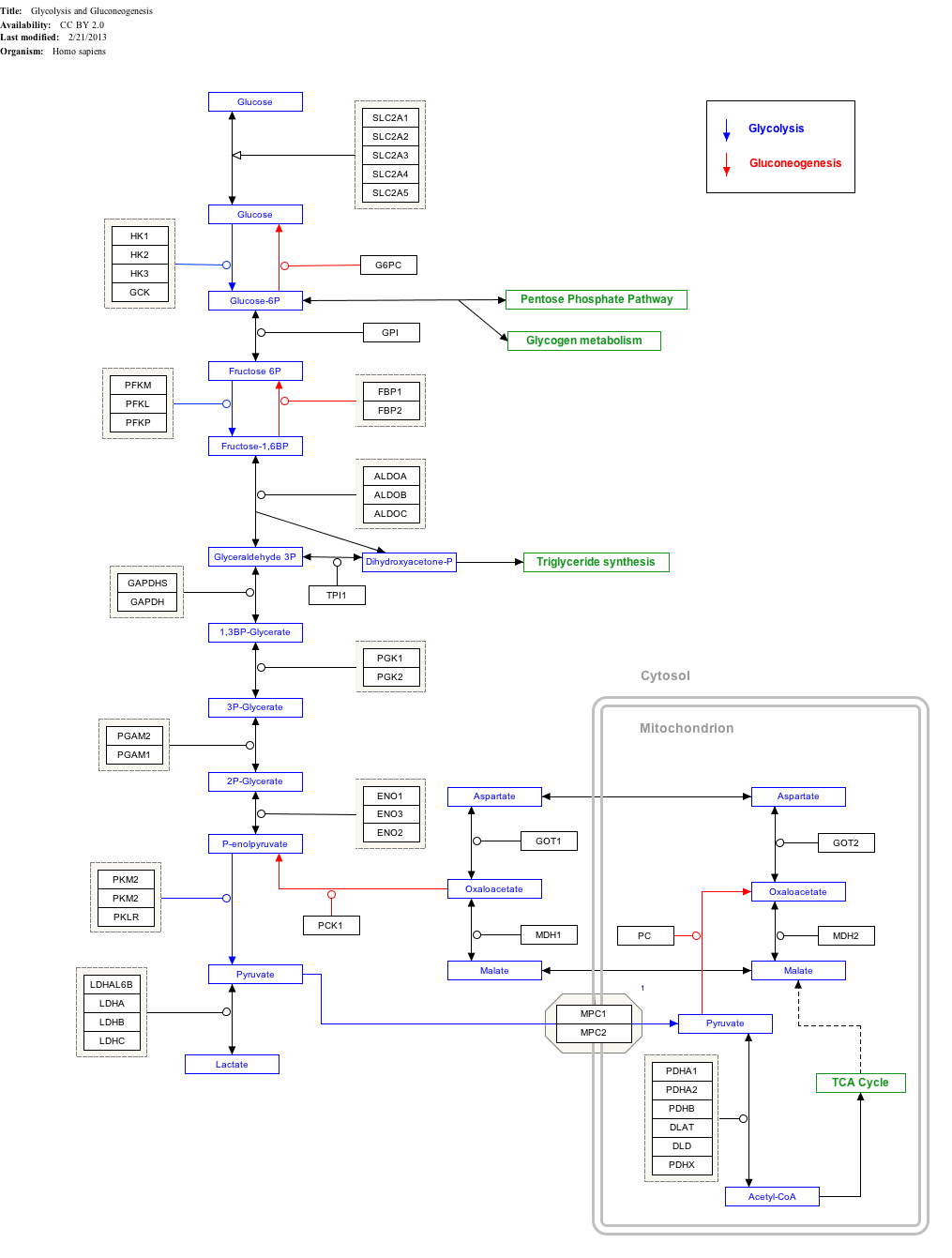
Glycolysis is the metabolic pathway that converts glucose into pyruvate, and in most organisms, occurs in the liquid part of cells, the cytosol. The free energy released in this process is used to form the high-energy molecules adenosine triphosphate (ATP) and reduced nicotinamide adenine dinucleotide (NADH). Glycolysis is a sequence of ten reactions catalyzed by enzymes.

In biochemistry, a kinase is an enzyme that catalyzes the transfer of phosphate groups from high-energy, phosphate-donating molecules to specific substrates. This process is known as phosphorylation, where the high-energy ATP molecule donates a phosphate group to the substrate molecule. This transesterification produces a phosphorylated substrate and ADP. Conversely, it is referred to as dephosphorylation when the phosphorylated substrate donates a phosphate group and ADP gains a phosphate group. These two processes, phosphorylation and dephosphorylation, occur four times during glycolysis.

In biochemistry, phosphorylation is the attachment of a phosphate group to a molecule or an ion. This process and its inverse, dephosphorylation, are common in biology. Protein phosphorylation often activates many enzymes.
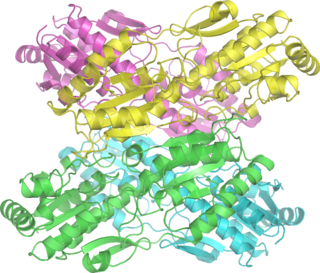
Phosphofructokinase-1 (PFK-1) is one of the most important regulatory enzymes of glycolysis. It is an allosteric enzyme made of 4 subunits and controlled by many activators and inhibitors. PFK-1 catalyzes the important "committed" step of glycolysis, the conversion of fructose 6-phosphate and ATP to fructose 1,6-bisphosphate and ADP. Glycolysis is the foundation for respiration, both anaerobic and aerobic. Because phosphofructokinase (PFK) catalyzes the ATP-dependent phosphorylation to convert fructose-6-phosphate into fructose 1,6-bisphosphate and ADP, it is one of the key regulatory steps of glycolysis. PFK is able to regulate glycolysis through allosteric inhibition, and in this way, the cell can increase or decrease the rate of glycolysis in response to the cell's energy requirements. For example, a high ratio of ATP to ADP will inhibit PFK and glycolysis. The key difference between the regulation of PFK in eukaryotes and prokaryotes is that in eukaryotes PFK is activated by fructose 2,6-bisphosphate. The purpose of fructose 2,6-bisphosphate is to supersede ATP inhibition, thus allowing eukaryotes to have greater sensitivity to regulation by hormones like glucagon and insulin.

Phosphoglucomutase is an enzyme that transfers a phosphate group on an α-D-glucose monomer from the 1 to the 6 position in the forward direction or the 6 to the 1 position in the reverse direction.
Substrate-level phosphorylation is a metabolism reaction that results in the production of ATP or GTP supported by the energy released from another high-energy bond that leads to phosphorylation of ADP or GDP to ATP or GTP (note that the reaction catalyzed by creatine kinase is not considered as "substrate-level phosphorylation"). This process uses some of the released chemical energy, the Gibbs free energy, to transfer a phosphoryl (PO3) group to ADP or GDP. Occurs in glycolysis and in the citric acid cycle.

Triose-phosphate isomerase is an enzyme that catalyzes the reversible interconversion of the triose phosphate isomers dihydroxyacetone phosphate and D-glyceraldehyde 3-phosphate.
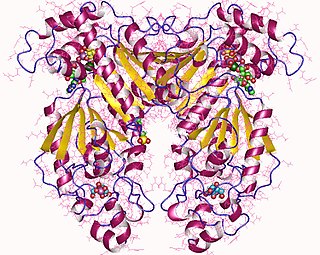
Phosphofructokinase-2 (6-phosphofructo-2-kinase, PFK-2) or fructose bisphosphatase-2 (FBPase-2), is an enzyme indirectly responsible for regulating the rates of glycolysis and gluconeogenesis in cells. It catalyzes formation and degradation of a significant allosteric regulator, fructose-2,6-bisphosphate (Fru-2,6-P2) from substrate fructose-6-phosphate. Fru-2,6-P2 contributes to the rate-determining step of glycolysis as it activates enzyme phosphofructokinase 1 in the glycolysis pathway, and inhibits fructose-1,6-bisphosphatase 1 in gluconeogenesis. Since Fru-2,6-P2 differentially regulates glycolysis and gluconeogenesis, it can act as a key signal to switch between the opposing pathways. Because PFK-2 produces Fru-2,6-P2 in response to hormonal signaling, metabolism can be more sensitively and efficiently controlled to align with the organism's glycolytic needs. This enzyme participates in fructose and mannose metabolism. The enzyme is important in the regulation of hepatic carbohydrate metabolism and is found in greatest quantities in the liver, kidney and heart. In mammals, several genes often encode different isoforms, each of which differs in its tissue distribution and enzymatic activity. The family described here bears a resemblance to the ATP-driven phospho-fructokinases, however, they share little sequence similarity, although a few residues seem key to their interaction with fructose 6-phosphate.

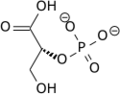
3-Phosphoglyceric acid (3PG, 3-PGA, or PGA) is the conjugate acid of 3-phosphoglycerate or glycerate 3-phosphate (GP or G3P). This glycerate is a biochemically significant metabolic intermediate in both glycolysis and the Calvin-Benson cycle. The anion is often termed as PGA when referring to the Calvin-Benson cycle. In the Calvin-Benson cycle, 3-phosphoglycerate is typically the product of the spontaneous scission of an unstable 6-carbon intermediate formed upon CO2 fixation. Thus, two equivalents of 3-phosphoglycerate are produced for each molecule of CO2 that is fixed. In glycolysis, 3-phosphoglycerate is an intermediate following the dephosphorylation (reduction) of 1,3-bisphosphoglycerate.
A mutase is an enzyme of the isomerase class that catalyzes the movement of a functional group from one position to another within the same molecule. In other words, mutases catalyze intramolecular group transfers. Examples of mutases include bisphosphoglycerate mutase, which appears in red blood cells and phosphoglycerate mutase, which is an enzyme integral to glycolysis. In glycolysis, it changes 3-phosphoglycerate to 2-phosphoglycerate by moving a single phosphate group within a single molecule.
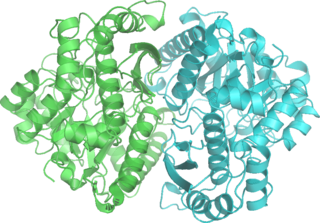
Phosphopyruvate hydratase, usually known as enolase, is a metalloenzyme (EC 4.2.1.11) that catalyses the conversion of 2-phosphoglycerate (2-PG) to phosphoenolpyruvate (PEP), the ninth and penultimate step of glycolysis. The chemical reaction is:

2,3-Bisphosphoglyceric acid (2,3-BPG), also known as 2,3-diphosphoglyceric acid (2,3-DPG), is a three-carbon isomer of the glycolytic intermediate 1,3-bisphosphoglyceric acid (1,3-BPG).

Phosphoglycerate kinase is an enzyme that catalyzes the reversible transfer of a phosphate group from 1,3-bisphosphoglycerate (1,3-BPG) to ADP producing 3-phosphoglycerate (3-PG) and ATP :

Bisphosphoglycerate mutase is an enzyme expressed in erythrocytes and placental cells. It is responsible for the catalytic synthesis of 2,3-Bisphosphoglycerate (2,3-BPG) from 1,3-bisphosphoglycerate. BPGM also has a mutase and a phosphatase function, but these are much less active, in contrast to its glycolytic cousin, phosphoglycerate mutase (PGM), which favors these two functions, but can also catalyze the synthesis of 2,3-BPG to a lesser extent.

Phosphofructokinase (PFK) is a kinase enzyme that phosphorylates fructose 6-phosphate in glycolysis.

Fructose-bisphosphate aldolase, often just aldolase, is an enzyme catalyzing a reversible reaction that splits the aldol, fructose 1,6-bisphosphate, into the triose phosphates dihydroxyacetone phosphate (DHAP) and glyceraldehyde 3-phosphate (G3P). Aldolase can also produce DHAP from other (3S,4R)-ketose 1-phosphates such as fructose 1-phosphate and sedoheptulose 1,7-bisphosphate. Gluconeogenesis and the Calvin cycle, which are anabolic pathways, use the reverse reaction. Glycolysis, a catabolic pathway, uses the forward reaction. Aldolase is divided into two classes by mechanism.
Glucose-1,6-bisphosphate synthase is a type of enzyme called a phosphotransferase and is involved in mammalian starch and sucrose metabolism. It catalyzes the transfer of a phosphate group from 1,3-bisphosphoglycerate to glucose-1-phosphate, yielding 3-phosphoglycerate and glucose-1,6-bisphosphate.
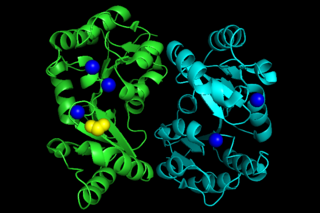
Phosphoglycolate phosphatase(EC 3.1.3.18; systematic name 2-phosphoglycolate phosphohydrolase), also commonly referred to as phosphoglycolate hydrolase, 2-phosphoglycolate phosphatase, P-glycolate phosphatase, and phosphoglycollate phosphatase, is an enzyme responsible for catalyzing the conversion of 2-phosphoglycolate into glycolate and phosphate:

Phosphoglycerate mutase 2 (PGAM2), also known as muscle-specific phosphoglycerate mutase (PGAM-M), is a phosphoglycerate mutase that, in humans, is encoded by the PGAM2 gene on chromosome 7.

2-Phosphoglycolate (chemical formula C2H2O6P3-; also known as phosphoglycolate, 2-PG, or PG) is a natural metabolic product of the oxygenase reaction mediated by the enzyme ribulose 1,5-bisphosphate carboxylase (RuBisCo).