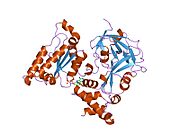
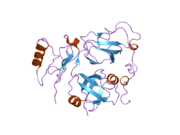
Proteasomes are protein complexes which degrade unneeded or damaged proteins by proteolysis, a chemical reaction that breaks peptide bonds. Enzymes that help such reactions are called proteases.
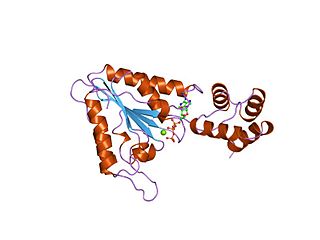
AAA proteins or ATPases Associated with diverse cellular Activities are a protein family sharing a common conserved module of approximately 230 amino acid residues. This is a large, functionally diverse protein family belonging to the AAA+ protein superfamily of ring-shaped P-loop NTPases, which exert their activity through the energy-dependent remodeling or translocation of macromolecules.

Endoplasmic-reticulum-associated protein degradation (ERAD) designates a cellular pathway which targets misfolded proteins of the endoplasmic reticulum for ubiquitination and subsequent degradation by a protein-degrading complex, called the proteasome.
The unfolded protein response (UPR) is a cellular stress response related to the endoplasmic reticulum (ER) stress. It has been found to be conserved between mammalian species, as well as yeast and worm organisms.

Binding immunoglobulin protein (BiPS) also known as 78 kDa glucose-regulated protein (GRP-78) or heat shock 70 kDa protein 5 (HSPA5) is a protein that in humans is encoded by the HSPA5 gene.

26S proteasome non-ATPase regulatory subunit 2, also as known as 26S Proteasome Regulatory Subunit Rpn1, is an enzyme that in humans is encoded by the PSMD2 gene.

Autocrine motility factor receptor, isoform 2 is a protein that in humans is encoded by the AMFR gene.

Homocysteine-responsive endoplasmic reticulum-resident ubiquitin-like domain member 1 protein is a protein that in humans is encoded by the HERPUD1 gene.

E3 ubiquitin-protein ligase synoviolin is an enzyme that in humans is encoded by the SYVN1 gene.

Nuclear factor erythroid 2-related factor 1 (Nrf1) also known as nuclear factor erythroid-2-like 1 (NFE2L1) is a protein that in humans is encoded by the NFE2L1 gene. Since NFE2L1 is referred to as Nrf1, it is often confused with nuclear respiratory factor 1 (Nrf1).
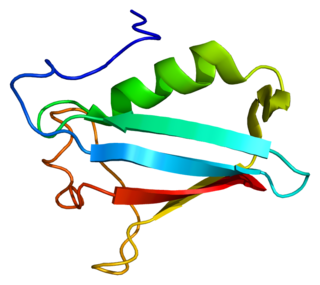
NSFL1 cofactor p47 is a protein that in humans is encoded by the NSFL1C gene.

Derlin-1 also known as degradation in endoplasmic reticulum protein 1 is a membrane protein that in humans is encoded by the DERL1 gene. Derlin-1 is located in the membrane of the endoplasmic reticulum (ER) and is involved in retrotranslocation of specific misfolded proteins and in ER stress. Derlin-1 is widely expressed in thyroid, fat, bone marrow and many other tissues. The protein belongs to the Derlin-family proteins consisting of derlin-1, derlin-2 and derlin-3 that are components in the endoplasmic reticulum-associated protein degradation (ERAD) pathway. The derlins mediate degradation of misfolded lumenal proteins within ER, and are named ‘der’ for their ‘Degradation in the ER’. Derlin-1 is a mammalian homologue of the yeast DER1 protein, a protein involved in the yeast ERAD pathway. Moreover, derlin-1 is a member of the rhomboid-like clan of polytopic membrane proteins.

PNGase also known as N-glycanase 1 or peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase is an enzyme that in humans is encoded by the NGLY1 gene. PNGase is a de-N-glycosylating enzyme that removes N-linked or asparagine-linked glycans (N-glycans) from glycoproteins. More specifically, NGLY1 catalyzes the hydrolysis of the amide bond between the innermost N-acetylglucosamine (GlcNAc) and an Asn residue on an N-glycoprotein, generating a de-N-glycosylated protein, in which the N-glycoylated Asn residue is converted to asp, and a 1-amino-GlcNAc-containing free oligosaccharide. Ammonia is then spontaneously released from the 1-amino GlcNAc at physiological pH (<8), giving rise to a free oligosaccharide with an N,N’-diacetylchitobiose structure at the reducing end.

F-box/LRR-repeat protein 2 is a protein that in humans is encoded by the FBXL2 gene.
The endosomal sorting complexes required for transport (ESCRT) machinery is made up of cytosolic protein complexes, known as ESCRT-0, ESCRT-I, ESCRT-II, and ESCRT-III. Together with a number of accessory proteins, these ESCRT complexes enable a unique mode of membrane remodeling that results in membranes bending/budding away from the cytoplasm. These ESCRT components have been isolated and studied in a number of organisms including yeast and humans. A eukaryotic signature protein, the machinery is found in all eukaryotes and some archaea.

Sec14 is a cytosolic protein found in yeast which plays a role in the regulation of several cellular functions, specifically those related to intracellular transport. Encoded by the Sec14 gene, Sec14p may transport phosphatidylinositol and phosphatidylcholine produced in the endoplasmic reticulum and the Golgi body to other cellular membranes. Additionally, Sec14p potentially plays a role in the localization of lipid raft proteins. Sec14p is an essential gene in yeast, and is homologous in function to phosphatidylinositol transfer protein in mammals. A conditional mutant with non-functional Sec14p presents with Berkeley bodies and deficiencies in protein secretion.

In molecular biology, the CDC48 N-terminal domain is a protein domain found in AAA ATPases including cell division protein 48 (CDC48), VCP-like ATPase and N-ethylmaleimide sensitive fusion protein. It is a substrate recognition domain which binds polypeptides, prevents protein aggregation, and catalyses refolding of permissive substrates. It is composed of two equally sized subdomains. The amino-terminal subdomain (CDC48_N) forms a double-psi beta-barrel whose pseudo-twofold symmetry is mirrored by an internal sequence repeat of 42 residues. The carboxy-terminal subdomain (CDC48_2) forms a novel six-stranded beta-clam fold. Together these subdomains form a kidney-shaped structure, in close agreement with results from electron microscopy. CDC48_N is related to numerous proteins including prokaryotic transcription factors, metabolic enzymes, the protease cofactors UFD1 and PrlF, and aspartic proteinases.

UBX domain protein 6 is a protein in humans that is encoded by the UBXN6 gene.
Raymond Joseph Deshaies is an American biochemist and cell biologist. He is senior vice president of global research at Amgen and a visiting associate at the California Institute of Technology (Caltech). Prior to that, he was a professor of biology at Caltech and an investigator of the Howard Hughes Medical Institute. He is also the co-founder of the biotechnology companies Proteolix and Cleave Biosciences. His research focuses on mechanisms and regulation of protein homeostasis in eukaryotic cells, with a particular focus on how proteins are conjugated with ubiquitin and degraded by the proteasome.
UBXD8 is a protein in the Ubiquitin regulatory X (UBX) domain-containing protein family. The UBX domain contains many eukaryotic proteins that have similarities in amino acid sequence to the tiny protein modifier ubiquitin. UBXD8 engages in a molecular interaction with p97, a protein that is essential for the degradation of membrane proteins associated with the endoplasmic reticulum (ER) through the proteasome. Ubxd8 possesses a UBA domain, alongside the UBX domain, that could interact with polyubiquitin chains. Additionally, it possesses a UAS domain of undetermined function, and this protein is used as a protein sensor that detects long chain unsaturated fatty acids (FAs), having a vital function in regulating the balance of Fatty Acids within cells to maintain cellular homeostasis.