In molecular biology, Small nucleolar RNAs (snoRNAs) are a class of small RNA molecules that primarily guide chemical modifications of other RNAs, mainly ribosomal RNAs, transfer RNAs and small nuclear RNAs. There are two main classes of snoRNA, the C/D box snoRNAs, which are associated with methylation, and the H/ACA box snoRNAs, which are associated with pseudouridylation. SnoRNAs are commonly referred to as guide RNAs but should not be confused with the guide RNAs that direct RNA editing in trypanosomes or the guide RNAs (gRNAs) used by Cas9 for CRISPR gene editing.
Guanylyl transferases are enzymes that transfer a guanosine mono phosphate group, usually from GTP to another molecule, releasing pyrophosphate. Many eukaryotic guanylyl transferases are capping enzymes that catalyze the formation of the 5' cap in the co-transcriptional modification of messenger RNA. Because the 5' end of the RNA molecule ends in a phosphate group, the bond formed between the RNA and the GTP molecule is an unusual 5'-5' triphosphate linkage, instead of the 3'-5' linkages between the other nucleotides that form an RNA strand. In capping enzymes, a highly conserved lysine residue serves as the catalytic residue that forms a covalent enzyme-GMP complex.

RNA-dependent RNA polymerase (RdRp) or RNA replicase is an enzyme that catalyzes the replication of RNA from an RNA template. Specifically, it catalyzes synthesis of the RNA strand complementary to a given RNA template. This is in contrast to typical DNA-dependent RNA polymerases, which all organisms use to catalyze the transcription of RNA from a DNA template.

The Avsunviroidae are a family of viroids. There are four species in three genera. They consist of RNA genomes between 246 and 375 nucleotides in length. They are single-stranded covalent circles and have intramolecular base pairing. All members lack a central conserved region.

Usually found in gram-positive bacteria, the T box leader sequence is an RNA element that controls gene expression through the regulation of translation by binding directly to a specific tRNA and sensing its aminoacylation state. This interaction controls expression of downstream aminoacyl-tRNA synthetase genes, amino acid biosynthesis, and uptake-related genes in a negative feedback loop. The uncharged tRNA acts as the effector for transcription antitermination of genes in the T-box leader family. The anticodon of a specific tRNA base pairs to a specifier sequence within the T-box motif, and the NCCA acceptor tail of the tRNA base pairs to a conserved bulge in the T-box antiterminator hairpin.

U2 spliceosomal snRNAs are a species of small nuclear RNA (snRNA) molecules found in the major spliceosomal (Sm) machinery of virtually all eukaryotic organisms. In vivo, U2 snRNA along with its associated polypeptides assemble to produce the U2 small nuclear ribonucleoprotein (snRNP), an essential component of the major spliceosomal complex. The major spliceosomal-splicing pathway is occasionally referred to as U2 dependent, based on a class of Sm intron—found in mRNA primary transcripts—that are recognized exclusively by the U2 snRNP during early stages of spliceosomal assembly. In addition to U2 dependent intron recognition, U2 snRNA has been theorized to serve a catalytic role in the chemistry of pre-RNA splicing as well. Similar to ribosomal RNAs (rRNAs), Sm snRNAs must mediate both RNA:RNA and RNA:protein contacts and hence have evolved specialized, highly conserved, primary and secondary structural elements to facilitate these types of interactions.

The ykkC/yxkD leader is a conserved RNA structure found upstream of the ykkC and yxkD genes in Bacillus subtilis and related genes in other bacteria. The function of this family is unclear for many years although it has been suggested that it may function to switch on efflux pumps and detoxification systems in response to harmful environmental molecules. The Thermoanaerobacter tengcongensis sequence AE013027 overlaps with that of purine riboswitch suggesting that the two riboswitches may work in conjunction to regulate the upstream gene which codes for TTE0584 (Q8RC62), a member of the permease family.
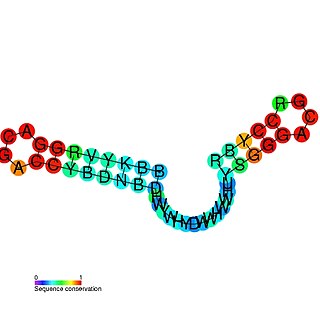
The mini-ykkC RNA motif was discovered as a putative RNA structure that is conserved in bacteria. The motif consists of two conserved stem-loops whose terminal loops contain the RNA sequence ACGR, where R represents either A or G. Mini-ykkC RNAs are widespread in Pseudomonadota, but some are predicted in other phyla of bacteria. It was expected that the RNAs are cis-regulatory elements, because they are typically located upstream of protein-coding genes.
The lacto-2 RNA motif is an RNA structure that is conserved amongst bacteria within the order Lactobacillales. The motif consists of a stem-loop whose stem is interrupted by many internal loops and bulges. Nucleotide identities in many places are conserved, and one internal loop in particular is highly conserved.
Giant, Ornate, Lake- and Lactobacillales-Derived (GOLLD) RNA is a conserved RNA structure present in bacteria. GOLLD RNAs were originally detected based on metagenome sequences of DNA isolated from Lake Gatun in Panama. However, they are known to be present in at least eight strains of cultivated bacteria. GOLLD RNAs are extraordinarily large compared to other RNAs with a conserved, complex secondary structure, and average roughly 800 nucleotides. Such large, complex RNAs are often ribozymes, although the biochemical function of GOLLD RNAs remains unknown. The discovery of large RNAs like GOLLD RNAs among bacteria that are mostly uncultivated under laboratory conditions suggests that many other unusually large RNAs might be found in bacteria that have not yet been studied.

The c4 antisense RNA is a non-coding RNA used by certain phages that infect bacteria. It was initially identified in the P1 and P7 phages of E. coli. The identification of c4 antisense RNAs solved the mystery of the mechanism for regulation of the ant gene, which is an anti-repressor.
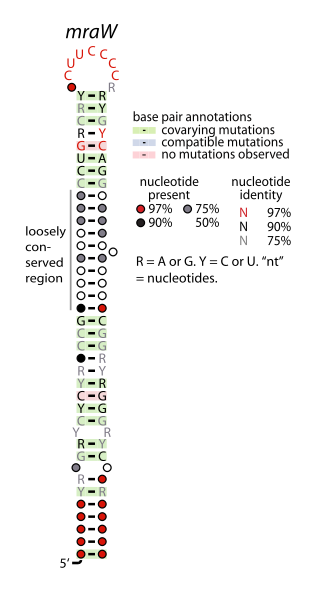

The mraW RNA motif is a conserved, structured RNA found in certain bacteria. Specifically, it is predicted in many, though not all, species of actinobacteria, and especially within the genus Mycobacterium. Structurally, the motif consists of a hairpin with a highly conserved terminal loop sequence. mraW RNAs are consistently in the presumed 5' untranslated regions of mraW genes. These mraW genes likely form operons with immediately downstream ftsI genes, and multiple types of mur genes. These genes are associated with peptidoglycan synthesis, and it was hypothesized that the mraW RNA motif might regulate these genes.
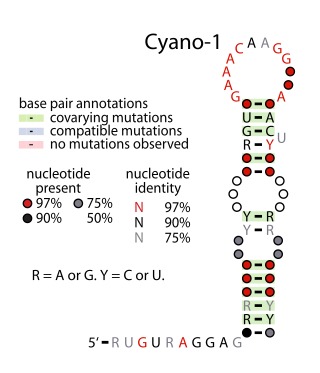
Yfr2 is a family of non-coding RNAs. Members of the Yrf2 family have been identified in almost all studied species of cyanobacteria. The family was identified through a bioinformatics screen of published cyanobacterial genomes, having previously been grouped in a family of Yfr2–5.
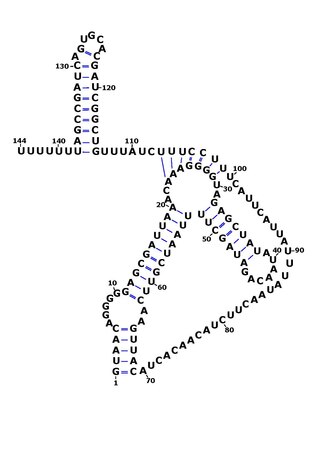
RsaOG is a non-coding RNA that was discovered in the pathogenic bacteria Staphylococcus aureus N315 using a large scale computational screening based on phylogenetic profiling. It was first identified, but not named, in 2005. RsaOG has since been identified in other strains of Staphylococcus aureus under the name of RsaI, it has also been discovered in other members of the Staphylococcus genus but in no other bacteria.
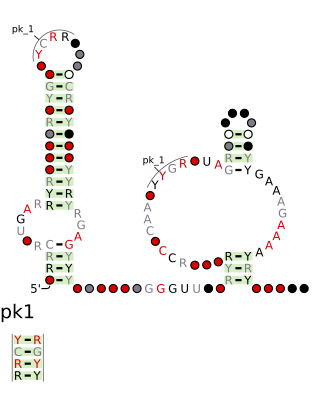
The drum RNA motif is a conserved RNA structure that was discovered by bioinformatics. Drum motifs are found in Bacillota, Bacteroidota, Pseudomonadota, and Spirochaetota, and exhibit multiple highly conserved nucleotide positions, despite their widespread distribution.
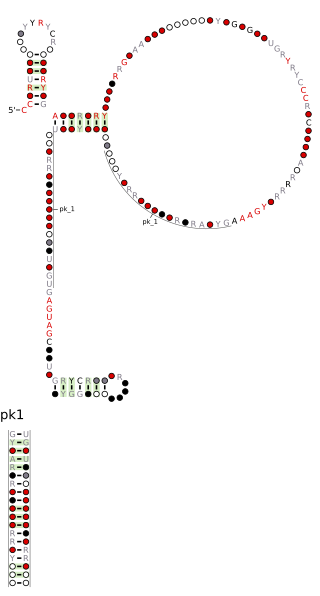
The Pseudomonadales-1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. The Pseudomonadales-1 motif often exhibits an apparent sarcin-ricin loop, a type internal loop common in RNA. Pseudomonadales-1 motif RNAs are found in relatively closely related species of Pseudomonadales. Despite this narrow distribution, the Pseudomonadales-1 RNA motif does not exhibit many invariant nucleotide positions, suggesting that it does not need to be highly conserved at the primary sequence level.
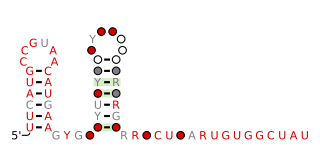
The queA RNA motif is a conserved RNA structure that was discovered by bioinformatics. queA motif RNAs have not yet been found in any classified organism; they are known from metagenomic sequences.
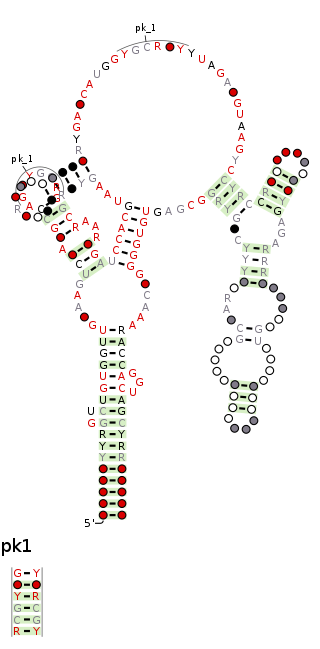
The raiA RNA motif is a conserved RNA structure that was discovered by bioinformatics. raiA motif RNAs are found in Actinomycetota and Bacillota, and have many conserved features—including conserved nucleotide positions, conserved secondary structures and associated protein-coding genes—in both of these phyla. Some conserved features of the raiA RNA motif suggest that they function as cis-regulatory elements, but other aspects of the motif suggest otherwise.

The uup RNA motif is a conserved RNA structure that was discovered by bioinformatics. uup motif RNAs are found in Bacillota and Gammaproteobacteria.
Coronavirus genomes are positive-sense single-stranded RNA molecules with an untranslated region (UTR) at the 5′ end which is called the 5′ UTR. The 5′ UTR is responsible for important biological functions, such as viral replication, transcription and packaging. The 5′ UTR has a conserved RNA secondary structure but different Coronavirus genera have different structural features described below.















