
The centromere links a pair of sister chromatids together during cell division. This constricted region of chromosome connects the sister chromatids, creating a short arm (p) and a long arm (q) on the chromatids. During mitosis, spindle fibers attach to the centromere via the kinetochore.

Meiosis is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately result in four cells with only one copy of each chromosome (haploid). Additionally, prior to the division, genetic material from the paternal and maternal copies of each chromosome is crossed over, creating new combinations of code on each chromosome. Later on, during fertilisation, the haploid cells produced by meiosis from a male and female will fuse to create a cell with two copies of each chromosome again, the zygote.
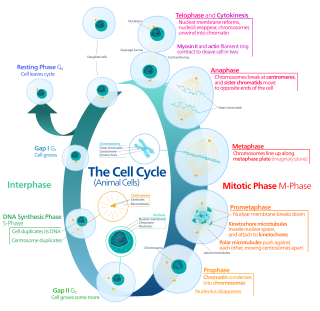
In cell biology, mitosis is a part of the cell cycle in which replicated chromosomes are separated into two new nuclei. Cell division by mitosis gives rise to genetically identical cells in which the total number of chromosomes is maintained. Therefore, mitosis is also known as equational division. In general, mitosis is preceded by S phase of interphase and is often followed by telophase and cytokinesis; which divides the cytoplasm, organelles and cell membrane of one cell into two new cells containing roughly equal shares of these cellular components. The different stages of mitosis altogether define the mitotic (M) phase of an animal cell cycle—the division of the mother cell into two daughter cells genetically identical to each other.

Cell division is the process by which a parent cell divides into two daughter cells. Cell division usually occurs as part of a larger cell cycle in which the cell grows and replicates its chromosome(s) before dividing. In eukaryotes, there are two distinct types of cell division: a vegetative division (mitosis), producing daughter cells genetically identical to the parent cell, and a cell division that produces haploid gametes for sexual reproduction (meiosis), reducing the number of chromosomes from two of each type in the diploid parent cell to one of each type in the daughter cells. In cell biology, mitosis (/maɪˈtoʊsɪs/) is a part of the cell cycle, in which, replicated chromosomes are separated into two new nuclei. Cell division gives rise to genetically identical cells in which the total number of chromosomes is maintained. In general, mitosis is preceded by the S stage of interphase and is often followed by telophase and cytokinesis; which divides the cytoplasm, organelles, and cell membrane of one cell into two new cells containing roughly equal shares of these cellular components. The different stages of mitosis all together define the mitotic (M) phase of animal cell cycle—the division of the mother cell into two genetically identical daughter cells. Meiosis results in four haploid daughter cells by undergoing one round of DNA replication followed by two divisions. Homologous chromosomes are separated in the first division, and sister chromatids are separated in the second division. Both of these cell division cycles are used in the process of sexual reproduction at some point in their life cycle. Both are believed to be present in the last eukaryotic common ancestor.

A couple of homologous chromosomes, or homologs, are a set of one maternal and one paternal chromosome that pair up with each other inside a cell during fertilization. Homologs have the same genes in the same loci where they provide points along each chromosome which enable a pair of chromosomes to align correctly with each other before separating during meiosis. This is the basis for Mendelian inheritance which characterizes inheritance patterns of genetic material from an organism to its offspring parent developmental cell at the given time and area.

Mosaicism or genetic mosaicism is a condition in which a multicellular organism possesses more than one genetic line as the result of genetic mutation. This means that various genetic lines resulted from a single fertilized egg. Mosaicism is one of several possible causes of chimerism, wherein a single organism is composed of cells with more than one distinct genotype.

A kinetochore is a disc-shaped protein structure associated with duplicated chromatids in eukaryotic cells where the spindle fibers attach during cell division to pull sister chromatids apart. The kinetochore assembles on the centromere and links the chromosome to microtubule polymers from the mitotic spindle during mitosis and meiosis. The term kinetochore was first used in a footnote in a 1934 Cytology book by Lester W. Sharp and commonly accepted in 1936. Sharp's footnote reads: "The convenient term kinetochore has been suggested to the author by J. A. Moore", likely referring to John Alexander Moore who had joined Columbia University as a freshman in 1932.
Double minutes are small fragments of extrachromosomal DNA, which have been observed in a large number of human tumors including breast, lung, ovary, colon, and most notably, neuroblastoma. They are a manifestation of gene amplification as a result of chromothripsis, during the development of tumors, which give the cells selective advantages for growth and survival. This selective advantage is as a result of double minutes frequently harboring amplified oncogenes and genes involved in drug resistance. DMs, like actual chromosomes, are composed of chromatin and replicate in the nucleus of the cell during cell division. Unlike typical chromosomes, they are composed of circular fragments of DNA, up to only a few million base pairs in size, and contain no centromere or telomere. Further to this, they often lack key regulatory elements, allowing genes to be constitutively expressed. The term ecDNA may be used to refer to DMs in a more general manner.

An isochromosome is an unbalanced structural abnormality in which the arms of the chromosome are mirror images of each other. The chromosome consists of two copies of either the long (q) arm or the short (p) arm because isochromosome formation is equivalent to a simultaneous duplication and deletion of genetic material. Consequently, there is partial trisomy of the genes present in the isochromosome and partial monosomy of the genes in the lost arm.
Subtelomeres are segments of DNA between telomeric caps and chromatin.

Micronucleus is the name given to the small nucleus that forms whenever a chromosome or a fragment of a chromosome is not incorporated into one of the daughter nuclei during cell division. It usually is a sign of genotoxic events and chromosomal instability. Micronuclei are commonly seen in cancerous cells and may indicate genomic damage events that can increase the risk of developmental or degenerative diseases. Micronuclei form during anaphase from lagging acentric chromosome or chromatid fragments caused by incorrectly repaired or unrepaired DNA breaks or by nondisjunction of chromosomes. This incorrect segregation of chromosomes may result from hypomethylation of repeat sequences present in pericentromeric DNA, irregularities in kinetochore proteins or their assembly, dysfunctional spindle apparatus, or flawed anaphase checkpoint genes. Micronuclei can contribute to genome instability by promoting a catastrophic mutational event called chromothripsis. Many micronucleus assays have been developed to test for the presence of these structures and determine their frequency in cells exposed to certain chemicals or subjected to stressful conditions.
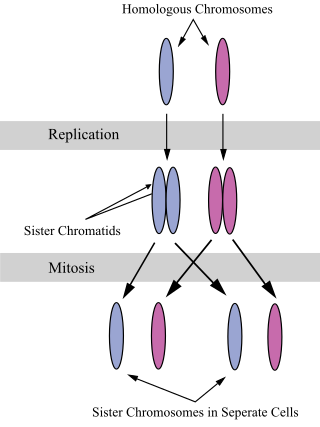
A sister chromatid refers to the identical copies (chromatids) formed by the DNA replication of a chromosome, with both copies joined together by a common centromere. In other words, a sister chromatid may also be said to be 'one-half' of the duplicated chromosome. A pair of sister chromatids is called a dyad. A full set of sister chromatids is created during the synthesis (S) phase of interphase, when all the chromosomes in a cell are replicated. The two sister chromatids are separated from each other into two different cells during mitosis or during the second division of meiosis.
A dicentric chromosome is an abnormal chromosome with two centromeres. It is formed through the fusion of two chromosome segments, each with a centromere, resulting in the loss of acentric fragments and the formation of dicentric fragments. The formation of dicentric chromosomes has been attributed to genetic processes, such as Robertsonian translocation and paracentric inversion. Dicentric chromosomes have important roles in the mitotic stability of chromosomes and the formation of pseudodicentric chromosomes. Their existence has been linked to certain natural phenomena such as irradiation and have been documented to underlie certain clinical syndromes, notably Kabuki syndrome. The formation of dicentric chromosomes and their implications on centromere function are studied in certain clinical cytogenetics laboratories.

A micronucleus test is a test used in toxicological screening for potential genotoxic compounds. The assay is now recognized as one of the most successful and reliable assays for genotoxic carcinogens, i.e., carcinogens that act by causing genetic damage and is recommended by the OECD guideline for the testing of chemicals. There are two major versions of this test, one in vivo and the other in vitro.
The origin and function of meiosis are currently not well understood scientifically, and would provide fundamental insight into the evolution of sexual reproduction in eukaryotes. There is no current consensus among biologists on the questions of how sex in eukaryotes arose in evolution, what basic function sexual reproduction serves, and why it is maintained, given the basic two-fold cost of sex. It is clear that it evolved over 1.2 billion years ago, and that almost all species which are descendants of the original sexually reproducing species are still sexual reproducers, including plants, fungi, and animals.

Neocentromeres are new centromeres that form at a place on the chromosome that is usually not centromeric. They typically arise due to disruption of the normal centromere. These neocentromeres should not be confused with “knobs”, which were also described as “neocentromeres” in maize in the 1950s. Unlike most normal centromeres, neocentromeres do not contain satellite sequences that are highly repetitive but instead consist of unique sequences. Despite this, most neocentromeres are still able to carry out the functions of normal centromeres in regulating chromosome segregation and inheritance. This raises many questions on what is necessary versus what is sufficient for constituting a centromere.
Holocentric chromosomes are chromosomes that possess multiple kinetochores along their length rather than the single centromere typical of other chromosomes. They were first described in cytogenetic experiments in 1935. Since this first observation, the term holocentric chromosome has referred to chromosomes that: i) lack the primary constriction corresponding to the centromere observed in monocentric chromosomes; and ii) possess multiple kinetochores dispersed along the entire chromosomal axis, such that microtubules bind to the chromosome along its entire length and move broadside to the pole from the metaphase plate. Holocentric chromosomes are also termed holokinetic, because, during cell division, the sister chromatids move apart in parallel and do not form the classical V-shaped figures typical of monocentric chromosomes.

The monocentric chromosome is a chromosome that has only one centromere in a chromosome and forms a narrow constriction.
This glossary of genetics is a list of definitions of terms and concepts commonly used in the study of genetics and related disciplines in biology, including molecular biology, cell biology, and evolutionary biology. It is intended as introductory material for novices; for more specific and technical detail, see the article corresponding to each term. For related terms, see Glossary of evolutionary biology.
This glossary of genetics is a list of definitions of terms and concepts commonly used in the study of genetics and related disciplines in biology, including molecular biology, cell biology, and evolutionary biology. It is split across two articles:











