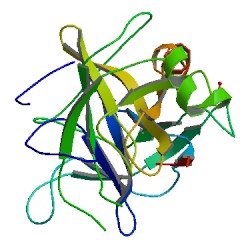
Chymotrypsin (EC 3.4.21.1, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides. Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid (tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme. Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, particularly those containing leucine at the P1 position.

Proteolysis is the breakdown of proteins into smaller polypeptides or amino acids. Uncatalysed, the hydrolysis of peptide bonds is extremely slow, taking hundreds of years. Proteolysis is typically catalysed by cellular enzymes called proteases, but may also occur by intra-molecular digestion.

Trypsin is an enzyme in the first section of the small intestine that starts the digestion of protein molecules by cutting long chains of amino acids into smaller pieces. It is a serine protease from the PA clan superfamily, found in the digestive system of many vertebrates, where it hydrolyzes proteins. Trypsin is formed in the small intestine when its proenzyme form, the trypsinogen produced by the pancreas, is activated. Trypsin cuts peptide chains mainly at the carboxyl side of the amino acids lysine or arginine. It is used for numerous biotechnological processes. The process is commonly referred to as trypsinogen proteolysis or trypsinization, and proteins that have been digested/treated with trypsin are said to have been trypsinized.

The pancreas is an organ of the digestive system and endocrine system of vertebrates. In humans, it is located in the abdomen behind the stomach and functions as a gland. The pancreas is a mixed or heterocrine gland, i.e., it has both an endocrine and a digestive exocrine function. 99% of the pancreas is exocrine and 1% is endocrine. As an endocrine gland, it functions mostly to regulate blood sugar levels, secreting the hormones insulin, glucagon, somatostatin and pancreatic polypeptide. As a part of the digestive system, it functions as an exocrine gland secreting pancreatic juice into the duodenum through the pancreatic duct. This juice contains bicarbonate, which neutralizes acid entering the duodenum from the stomach; and digestive enzymes, which break down carbohydrates, proteins and fats in food entering the duodenum from the stomach.

A protease is an enzyme that catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in numerous biological pathways, including digestion of ingested proteins, protein catabolism, and cell signaling.
In biochemistry, a zymogen, also called a proenzyme, is an inactive precursor of an enzyme. A zymogen requires a biochemical change for it to become an active enzyme. The biochemical change usually occurs in Golgi bodies, where a specific part of the precursor enzyme is cleaved in order to activate it. The inactivating piece which is cleaved off can be a peptide unit, or can be independently-folding domains comprising more than 100 residues. Although they limit the enzyme's ability, these N-terminal extensions of the enzyme or a "prosegment" often aid in the stabilization and folding of the enzyme they inhibit.
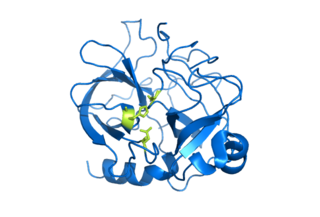
Serine proteases are enzymes that cleave peptide bonds in proteins. Serine serves as the nucleophilic amino acid at the (enzyme's) active site. They are found ubiquitously in both eukaryotes and prokaryotes. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like.

Digestive enzymes take part in the chemical process of digestion, which follows the mechanical process of digestion. Food consists of macromolecules of proteins, carbohydrates, and fats that need to be broken down chemically by digestive enzymes in the mouth, stomach, pancreas, and duodenum, before being able to be absorbed into the bloodstream. Initial breakdown is achieved by chewing (mastication) and the use of digestive enzymes of saliva. Once in the stomach further mechanical churning takes place mixing the food with secreted gastric acid. Digestive gastric enzymes take part in some of the chemical process needed for absorption. Most of the enzymatic activity, and hence absorption takes place in the duodenum.
Trypsinogen is the precursor form of trypsin, a digestive enzyme. It is produced by the pancreas and found in pancreatic juice, along with amylase, lipase, and chymotrypsinogen. It is cleaved to its active form, trypsin, by enteropeptidase, which is found in the intestinal mucosa. Once activated, the trypsin can cleave more trypsinogen into trypsin, a process called autoactivation. Trypsin cleaves the peptide bond on the carboxyl side of basic amino acids such as arginine and lysine.
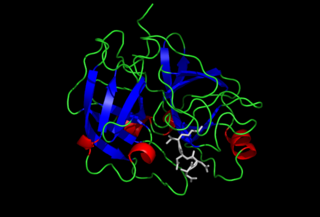
Enteropeptidase is an enzyme produced by cells of the duodenum and is involved in digestion in humans and other animals. Enteropeptidase converts trypsinogen into its active form trypsin, resulting in the subsequent activation of pancreatic digestive enzymes. Absence of enteropeptidase results in intestinal digestion impairment.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.

An oxyanion hole is a pocket in the active site of an enzyme that stabilizes transition state negative charge on a deprotonated oxygen or alkoxide. The pocket typically consists of backbone amides or positively charged residues. Stabilising the transition state lowers the activation energy necessary for the reaction, and so promotes catalysis. For example, proteases such as chymotrypsin contain an oxyanion hole to stabilise the tetrahedral intermediate anion formed during proteolysis and protects substrate's negatively charged oxygen from water molecules. Additionally, it may allow for insertion or positioning of a substrate, which would suffer from steric hindrance if it could not occupy the hole. Enzymes that catalyse multi-step reactions can have multiple oxyanion holes that stabilise different transition states in the reaction.
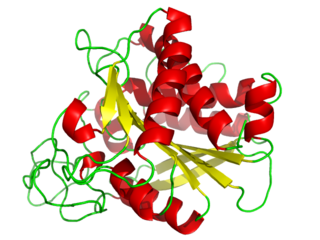
A carboxypeptidase is a protease enzyme that hydrolyzes (cleaves) a peptide bond at the carboxy-terminal (C-terminal) end of a protein or peptide. This is in contrast to an aminopeptidases, which cleave peptide bonds at the N-terminus of proteins. Humans, animals, bacteria and plants contain several types of carboxypeptidases that have diverse functions ranging from catabolism to protein maturation. At least two mechanisms have been discussed.
Pancreatic elastase is a form of elastase that is produced in the acinar cells of the pancreas, initially produced as an inactive zymogen and later activated in the duodenum by trypsin. Elastases form a subfamily of serine proteases, characterized by a distinctive structure consisting of two beta barrel domains converging at the active site that hydrolyze amides and esters amongst many proteins in addition to elastin, a type of connective tissue that holds organs together. Pancreatic elastase 1 is a serine endopeptidase, a specific type of protease that has the amino acid serine at its active site. Although the recommended name is pancreatic elastase, it can also be referred to as elastase-1, pancreatopeptidase, PE, or serine elastase.
Enzyme catalysis is the increase in the rate of a process by an "enzyme", a biological molecule. Most enzymes are proteins, and most such processes are chemical reactions. Within the enzyme, generally catalysis occurs at a localized site, called the active site.
Protein metabolism denotes the various biochemical processes responsible for the synthesis of proteins and amino acids (anabolism), and the breakdown of proteins by catabolism.

Trypsin-1, also known as cationic trypsinogen, is a protein that in humans is encoded by the PRSS1 gene. Trypsin-1 is the main isoform of trypsinogen secreted by pancreas, the others are trypsin-2, and trypsin-3 (meso-trypsinogen).
Alpha-lytic endopeptidase or Alpha-lytic protease is an enzyme isolated from the myxobacterium Lysobacter enzymogenes. This enzyme is a serine protease that catalyses the breakage of peptide bonds using a hydrolysis chemical reaction. Alpha-lytic protease was named based on the observed cleavage specificity for the α position of the tetrapeptide component in gram-positive bacterial cell walls (alanine). Alpha-lytic protease is also capable of digesting elastin and other proteins.

Monitor peptide, also known as pancreatic secretory trypsin inhibitor I (PSTI-I) or pancreatic secretory trypsin inhibitor 61 (PSTI-61), is a peptide that plays an important role in the regulation of the digestive system, specifically the release of cholecystokinin (CCK).














