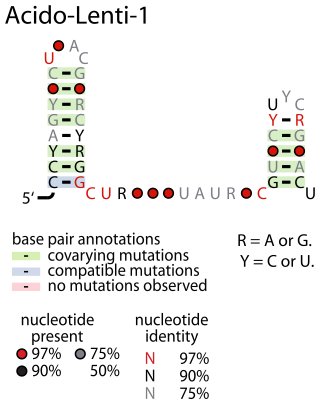
The Acido-Lenti-1 RNA motif describes a predicted non-coding RNA that is found in bacteria within the phyla acidobacteriota and lentisphaerota. It is sometimes found nearby to group II introns, but the reason for this apparent association is unknown.

The Bacillaceae-1 RNA motif is a conserved RNA structure identified by bioinformatics within bacteria in the family bacillaceae. The RNA is presumed to operate as a non-coding RNA, and is sometimes adjacent to operons containing ribosomal RNAs. The most characteristic feature is two terminal loops that have the nucleotide consensus RUCCU, where R is either A or G. The motif might be related to the Desulfotalea-1 RNA motif, as the motifs share some similarity in conserved features, and the Desulfotalea-1 RNA motif is also sometimes adjacent to ribosomal RNA operons.

The Bacteroid-trp RNA motif is a conserved RNA element detected by bioinformatics. It is found in the phylum Bacteroidota in the apparent 5' untranslated regions of genes that encode enzymes used in the synthesis of the amino acid tryptophan. A short open reading frame is found within the motif that encodes at least two tryptophan codons. Similar motifs have been identified regulating tryptophan genes in Pseudomonadota, but not in Bacteroidota. However, the Bacteroid-trp RNA motif likely operates via the same mechanism of attenuation.

The Bacteroidales-1 RNA motif is a conserved RNA structure identified by bioinformatics. It has been identified only in bacteria within the order (biology) Bacteroidales. Its presumed length is marked by a promoter on one end that conforms to an alternate consensus sequence that is common in the phylum Bacteroidota, and its 3′ end is indicated by predicted transcription terminators. It is often located downstream of a gene that encodes the L20 ribosomal subunit, although it is unclear whether there is a functional reason underlying this apparent association.
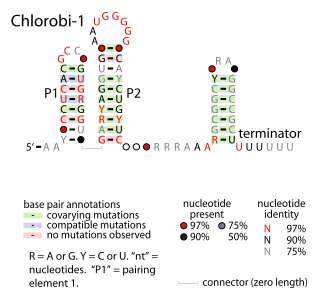
The Chlorobi-1 RNA motif is a conserved RNA secondary structure identified by bioinformatics. It is predicted to be used only by Chlorobiota, a phylum of bacteria. The motif consists of two stem-loops that are followed by an apparent rho-independent transcription terminator. The motif is presumed to function as an independently transcribed non-coding RNA.
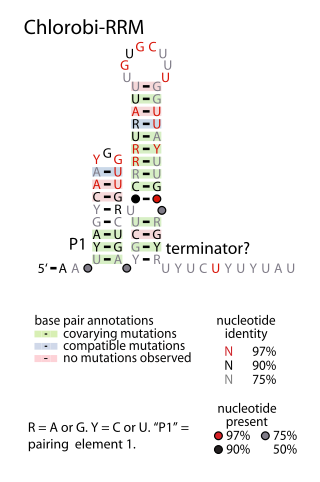
The Chlorobi-RRM RNA motif is a conserved RNA structure identified by bioinformatics. It is found within bacteria in the phylum Chlorobiota, and is exclusively detected in the presumed 5' untranslated regions of genes that encode putative RNA-binding proteins. Since many RNA-binding proteins regulate their own expression in a feedback mechanism by binding or acting up their 5' UTR, it was proposed that the Chlorobi-RRM is a component in an analogous feedback mechanism. Structurally, the motif consists of two stem-loops, the second of which might function as a rho-independent transcription terminator.

The Chloroflexi-1 RNA motif is a conserved RNA structure detected by bioinformatics within the species Chloroflexus aggregans. C. aggregans has three predicted Chloroflexi-1 RNAs, which are located nearby to one another. This arrangement might suggest a repetitive element. C. aggregans is classified as belonging to the bacterial phylum Chloroflexota.

The Collinsella-1 RNA motif denotes a particular conserved RNA structure discovered by bioinformatics. Of the six sequences belonging to this motif that were originally identified, five are from uncultivated bacteria residing in the human gut, while only the sixth is in a cultivated species, Collinsella aerofaciens. The evidence supporting the stem-loops designated as "P1" and "P2" is ambiguous.
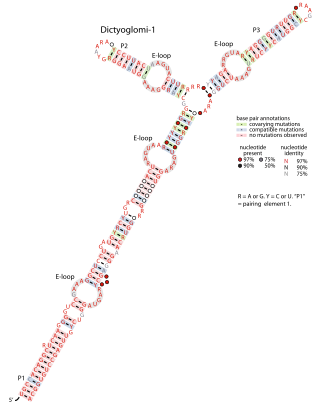
The Dictyoglomi-1 RNA motif is a conserved RNA structure that was discovered via bioinformatics. Only four instances of the RNA were detected, and all are in the bacterial phylum Dictyoglomota, whose members have not been extensively studied. The RNA might have a cis-regulatory role, but the evidence is ambiguous. Because of the few instances of Dictyoglomi-1 RNAs known, it is also unknown whether the RNA structure might extend further in the 5′ or 3′ direction, or in both directions.
The Lnt RNA motif refers to a conserved RNA structure found in certain bacteria. Specifically, Lnt RNAs are known only in species within the phylum Chlorobiota, and are located in the possible 5' untranslated regions of genes that are annotated as encoding apolipoprotein N-acyltransferase enzymes. There is some doubt as to whether the indicated motif is transcribed as RNA, or whether its reverse complement is transcribed. If the reverse complement is transcribed it would potentially in 5' UTRs of genes encoding bacteriochlorophyll A, and would be close to the start codon of those genes.

The pan RNA motif defines a conserved RNA structure that was identified using bioinformatics. pan motif RNAs are present in three phyla: Chloroflexota, Bacillota, and Pseudomonadota, although within the latter phylum they are only known in deltaproteobacteria. A pan RNA is present in the Firmicute Bacillus subtilis, which is one of the most extensively studied bacteria.

The pfl RNA motif refers to a conserved RNA structure present in some bacteria and originally discovered using bioinformatics. pfl RNAs are consistently present in genomic locations that likely correspond to the 5' untranslated regions of protein-coding genes. This arrangement in bacteria is commonly associated with cis-regulatory elements. Moreover, they are in presumed 5' UTRs of multiple non-homologous genes, suggesting that they function only in these locations. Additional evidence of cis-regulatory function came from the observation that predicted rho-independent transcription terminators overlap pfl RNAs. This overlap suggests that the alternate secondary structures of pfl RNA and the transcription terminator stem-loops compete with each other, and this is a common mechanism for cis gene control in bacteria.
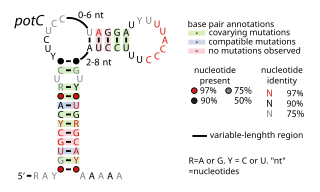
The potC RNA motif is a conserved RNA structure discovered using bioinformatics. The RNA is detected only in genome sequences derived from DNA that was extracted from uncultivated marine bacteria. Thus, this RNA is present in environmental samples, but not yet found in any cultivated organism. potC RNAs are located in the presumed 5' untranslated regions of genes predicted to encode either membrane transport proteins or peroxiredoxins. Therefore, it was hypothesized that potC RNAs are cis-regulatory elements, but their detailed function is unknown.

The SAM-Chlorobi RNA motif is a conserved RNA structure that was identified by bioinformatics. The RNAs are found only in bacteria classified as within the phylum Chlorobiota. These RNAs are always in the 5' untranslated regions of operons that contain metK and ahcY genes. metK genes encode methionine adenosyltransferase, which synthesizes S-adenosyl methionine (SAM), and ahcY genes encode S-adenosylhomocysteine hydrolase, which degrade the related metabolite S-Adenosyl-L-homocysteine (SAH). In fact all predicted metK and ahcY genes within Chlorobiota bacteria as of 2010 are preceded by predicted SAM-Chlorobi RNAs. Predicted promoter sequences are consistently found upstream of SAM-Chlorobi RNAs, and these promoter sequences imply that SAM-Chlorobi RNAs are indeed transcribed as RNAs. The promoter sequences are commonly associated with strong transcription in the phyla Chlorobiota and Bacteroidota, but are not used by most lineages of bacteria. The placement of SAM-Chlorobi RNAs suggests that they are involved in the regulation of the metK/ahcY operon through an unknown mechanism.

The Ssbp, Topoisomerase, Antirestriction, XerDC Integrase RNA motif is a conserved RNA-like structure identified using bioinformatics. STAXI RNAs are located near to genes encoding proteins that interact with DNA or are associated with such proteins. This observation raised the possibility that instances of the STAXI motif function as single-stranded DNA molecules, perhaps during DNA replication or DNA repair. On the other hand, STAXI motifs often contain terminal loops conforming to the stable UNCG tetraloop, but the DNA version of this tetraloop (TNCG) is not especially stable. The STAXI motif consists of a simple pseudoknot structure that is repeated two or more times.
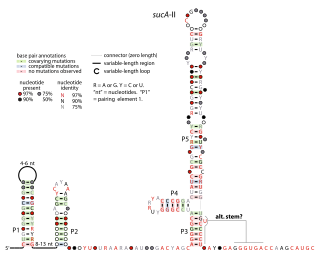
The sucA-II RNA motif is a conserved RNA structure identified by bioinformatics. It is consistently found in the presumed 5' untranslated regions of sucA genes, which encode Oxoglutarate dehydrogenase enzymes that participate in the citric acid cycle. Given this arrangement, sucA-II RNAs might regulate the downstream sucA gene. This genetic arrangement is similar to the previously reported sucA RNA motif. However, sucA-II RNAs are found only in bacteria classified within the genus Pseudomonas, whereas the previously reported motif is found only in betaproteobacteria.

The Whalefall-1 RNA motif refers to a conserved RNA structure that was discovered using bioinformatics. Structurally, the motif consists of two stem-loops, the second of which is often terminated by a CUUG tetraloop, which is an energetically favorable RNA sequence. Whalefall-1 RNAs are found only in DNA extracted from uncultivated bacteria found on whale fall, i.e., a whale carcass. As of 2010, Whalefall-1 RNAs have not been detected in any known, cultivated species of bacteria, and are thus one of several RNAs present in environmental samples.

The yjdF RNA motif is a conserved RNA structure identified using bioinformatics. Most yjdF RNAs are located in bacteria classified within the phylum Bacillota. A yjdF RNA is found in the presumed 5' untranslated region of the yjdF gene in Bacillus subtilis, and almost all yjdF RNAs are found in the 5' UTRs of homologs of this gene. The function of the yjdF gene is unknown, but the protein that it is predicted to encode is classified by the Pfam Database as DUF2992.
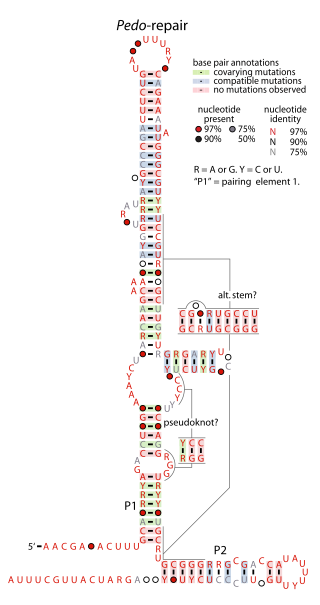
The Pedo-repair RNA motif is a conserved RNA structure identified by using bioinformatics. It has been detected in only one species of bacteria: Pedobacter sp. BAL39, within the phylum Bacteroidota. The motif might be in the 5′ untranslated regions of operons containing genes predicted to be involved in DNA repair or related to restriction enzymes.
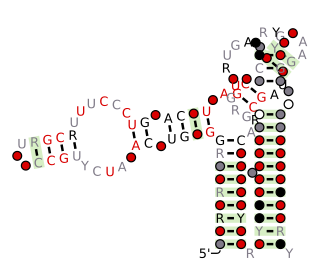
The Flavobacterium-1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. Flavobacterium-1 motif RNAs are found in metagenomic samples from the environment, and only one example of this motif is present in a classified organism. This organism is Flavobacterium sp. SCGC AAA160-P02, which belongs to the bacterial phylum Bacteroidota.



















