Related Research Articles
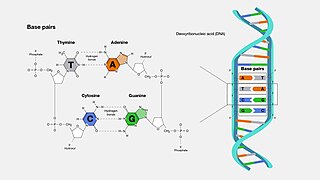
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" base pairs allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding proteins can recognize specific base-pairing patterns that identify particular regulatory regions of genes.

The genetic code is the set of rules used by living cells to translate information encoded within genetic material into proteins. Translation is accomplished by the ribosome, which links proteinogenic amino acids in an order specified by messenger RNA (mRNA), using transfer RNA (tRNA) molecules to carry amino acids and to read the mRNA three nucleotides at a time. The genetic code is highly similar among all organisms and can be expressed in a simple table with 64 entries.

Nucleotides are organic molecules composed of a nitrogenous base, a pentose sugar and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver.

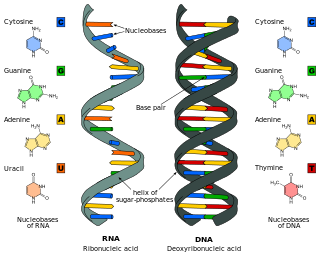
Nucleobases are nitrogen-containing biological compounds that form nucleosides, which, in turn, are components of nucleotides, with all of these monomers constituting the basic building blocks of nucleic acids. The ability of nucleobases to form base pairs and to stack one upon another leads directly to long-chain helical structures such as ribonucleic acid (RNA) and deoxyribonucleic acid (DNA). Five nucleobases—adenine (A), cytosine (C), guanine (G), thymine (T), and uracil (U)—are called primary or canonical. They function as the fundamental units of the genetic code, with the bases A, G, C, and T being found in DNA while A, G, C, and U are found in RNA. Thymine and uracil are distinguished by merely the presence or absence of a methyl group on the fifth carbon (C5) of these heterocyclic six-membered rings. In addition, some viruses have aminoadenine (Z) instead of adenine. It differs in having an extra amine group, creating a more stable bond to thymine.

Har Gobind Khorana was an Indian-American biochemist. While on the faculty of the University of Wisconsin–Madison, he shared the 1968 Nobel Prize for Physiology or Medicine with Marshall W. Nirenberg and Robert W. Holley for research that showed the order of nucleotides in nucleic acids, which carry the genetic code of the cell and control the cell's synthesis of proteins. Khorana and Nirenberg were also awarded the Louisa Gross Horwitz Prize from Columbia University in the same year.

Synthetic biology (SynBio) is a multidisciplinary field of science that focuses on living systems and organisms, and it applies engineering principles to develop new biological parts, devices, and systems or to redesign existing systems found in nature.
Peter G. Schultz is an American chemist. He is the CEO and Professor of Chemistry at The Scripps Research Institute, the founder and former director of GNF, and the founding director of the California Institute for Biomedical Research (Calibr), established in 2012. In August 2014, Nature Biotechnology ranked Schultz the #1 top translational researcher in 2013.
Xenobiology (XB) is a subfield of synthetic biology, the study of synthesizing and manipulating biological devices and systems. The name "xenobiology" derives from the Greek word xenos, which means "stranger, alien". Xenobiology is a form of biology that is not (yet) familiar to science and is not found in nature. In practice, it describes novel biological systems and biochemistries that differ from the canonical DNA–RNA-20 amino acid system. For example, instead of DNA or RNA, XB explores nucleic acid analogues, termed xeno nucleic acid (XNA) as information carriers. It also focuses on an expanded genetic code and the incorporation of non-proteinogenic amino acids into proteins.
Steven Albert Benner is an American chemist. He has been a professor at Harvard University, ETH Zurich, and most recently at the University of Florida, where he was the V.T. & Louise Jackson Distinguished Professor of Chemistry. In 2005, he founded The Westheimer Institute of Science and Technology (TWIST) and the Foundation For Applied Molecular Evolution. Benner has also founded the companies EraGen Biosciences and Firebird BioMolecular Sciences LLC.
Synthetic genomics is a nascent field of synthetic biology that uses aspects of genetic modification on pre-existing life forms, or artificial gene synthesis to create new DNA or entire lifeforms.

Artificial gene synthesis, or simply gene synthesis, refers to a group of methods that are used in synthetic biology to construct and assemble genes from nucleotides de novo. Unlike DNA synthesis in living cells, artificial gene synthesis does not require template DNA, allowing virtually any DNA sequence to be synthesized in the laboratory. It comprises two main steps, the first of which is solid-phase DNA synthesis, sometimes known as DNA printing. This produces oligonucleotide fragments that are generally under 200 base pairs. The second step then involves connecting these oligonucleotide fragments using various DNA assembly methods. Because artificial gene synthesis does not require template DNA, it is theoretically possible to make a completely synthetic DNA molecule with no limits on the nucleotide sequence or size.

Roger Yonchien Tsien was an American biochemist. He was a professor of chemistry and biochemistry at the University of California, San Diego and was awarded the Nobel Prize in Chemistry in 2008 for his discovery and development of the green fluorescent protein, in collaboration with organic chemist Osamu Shimomura and neurobiologist Martin Chalfie. Tsien was also a pioneer of calcium imaging.
Nucleic acid analogues are compounds which are analogous to naturally occurring RNA and DNA, used in medicine and in molecular biology research. Nucleic acids are chains of nucleotides, which are composed of three parts: a phosphate backbone, a pentose sugar, either ribose or deoxyribose, and one of four nucleobases. An analogue may have any of these altered. Typically the analogue nucleobases confer, among other things, different base pairing and base stacking properties. Examples include universal bases, which can pair with all four canonical bases, and phosphate-sugar backbone analogues such as PNA, which affect the properties of the chain . Nucleic acid analogues are also called Xeno Nucleic Acid and represent one of the main pillars of xenobiology, the design of new-to-nature forms of life based on alternative biochemistries.

An expanded genetic code is an artificially modified genetic code in which one or more specific codons have been re-allocated to encode an amino acid that is not among the 22 common naturally-encoded proteinogenic amino acids.

Genetic engineering is the science of manipulating genetic material of an organism. The first artificial genetic modification accomplished using biotechnology was transgenesis, the process of transferring genes from one organism to another, first accomplished by Herbert Boyer and Stanley Cohen in 1973. It was the result of a series of advancements in techniques that allowed the direct modification of the genome. Important advances included the discovery of restriction enzymes and DNA ligases, the ability to design plasmids and technologies like polymerase chain reaction and sequencing. Transformation of the DNA into a host organism was accomplished with the invention of biolistics, Agrobacterium-mediated recombination and microinjection. The first genetically modified animal was a mouse created in 1974 by Rudolf Jaenisch. In 1976 the technology was commercialised, with the advent of genetically modified bacteria that produced somatostatin, followed by insulin in 1978. In 1983 an antibiotic resistant gene was inserted into tobacco, leading to the first genetically engineered plant. Advances followed that allowed scientists to manipulate and add genes to a variety of different organisms and induce a range of different effects. Plants were first commercialized with virus resistant tobacco released in China in 1992. The first genetically modified food was the Flavr Savr tomato marketed in 1994. By 2010, 29 countries had planted commercialized biotech crops. In 2000 a paper published in Science introduced golden rice, the first food developed with increased nutrient value.

Xeno nucleic acids (XNA) are synthetic nucleic acid analogues that have a different sugar backbone than the natural nucleic acids DNA and RNA. As of 2011, at least six types of synthetic sugars have been shown to form nucleic acid backbones that can store and retrieve genetic information. Research is now being done to create synthetic polymerases to transform XNA. The study of its production and application has created a field known as xenobiology.

Phil S. Baran is a Professor in the Department of Chemistry at the Scripps Research Institute and Member of the Skaggs Institute for Chemical Biology. Baran has authored over 130 published scientific articles. He has several patents. His work is focused on synthesizing complex organic compounds, the development of new reactions, and the development of new reagents.

dNaM is an artificial nucleobase containing a 3-methoxy-2-naphthyl group instead of a natural base.

The arylomycins are a class of antibiotics initially isolated from a soil sample obtained in Cape Coast, Ghana. In this initial isolation, two families of closely related arylomycins, A and B, were identified. The family of glycosylated arylomycin C lipopeptides were subsequently isolated from a Streptomyces culture in a screen for inhibitors of bacterial signal peptidase. The initially isolated arylomycins have a limited spectrum of activity against Gram-positive bacteria, including Staphylococcus aureus and Streptococcus pneumoniae. The only activity against Gram-negative bacteria was seen in strains with a compromised outer membrane.

Hachimoji DNA is a synthetic nucleic acid analog that uses four synthetic nucleotides in addition to the four present in the natural nucleic acids, DNA and RNA. This leads to four allowed base pairs: two unnatural base pairs formed by the synthetic nucleobases in addition to the two normal pairs. Hachimoji bases have been demonstrated in both DNA and RNA analogs, using deoxyribose and ribose respectively as the backbone sugar.
References
- ↑ "Star scientist leaves Scripps Research under mysterious circumstances". San Diego Union-Tribune. 2019-08-09. Retrieved 2021-08-31.
- ↑ Malyshev, Denis A.; Dhami, Kirandeep; Quach, Henry T.; Lavergne, Thomas; Ordoukhanian, Phillip (24 July 2012). "Efficient and sequence-independent replication of DNA containing a third base pair establishes a functional six-letter genetic alphabet". Proceedings of the National Academy of Sciences of the United States of America. 109 (30): 12005–12010. Bibcode:2012PNAS..10912005M. doi: 10.1073/pnas.1205176109 . PMC 3409741 . PMID 22773812.
- ↑ Malyshev; et al. (7 May 2014). "A Semi-Synthetic Organism with an Expanded Genetic Alphabet". Nature. 509 (7500): 385–8. Bibcode:2014Natur.509..385M. doi:10.1038/nature13314. PMC 4058825 . PMID 24805238.
- ↑ Andrew Pollack (May 7, 2014). "Scientists Add Letters to DNA's Alphabet, Raising Hope and Fear". New York Times. Retrieved 17 December 2018.
- ↑ Chris Jennewein (29 November 2017). "Scripps Research Creates First 'Semi-Synthetic' E. Coli Bacteria". Times of San Diego. Retrieved 17 December 2018.
- ↑ "Nobel Laureate Signature Award for Graduate Education in Chemistry". American Chemical Society.
- ↑ "Scripps Research chemist receives prestigious Royal Society of Chemistry Award". 8 May 2018. Retrieved 17 December 2018.