
The COG2908 RNA motif is a conserved RNA structure that was discovered by bioinformatics. COG2908 motif RNAs are found in genomic sequences extracted from fresh water environments. They have not, as of 2018, been detected in any classified organism.
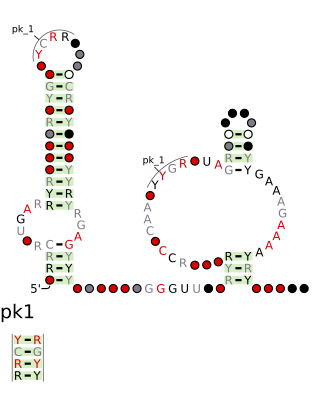
The drum RNA motif is a conserved RNA structure that was discovered by bioinformatics. Drum motifs are found in Bacillota, Bacteroidota, Pseudomonadota, and Spirochaetota, and exhibit multiple highly conserved nucleotide positions, despite their widespread distribution.

The DUF805 RNA motif is a conserved RNA structure that was discovered by bioinformatics. The motif is subdivided into the DUF805 motif and the DUF805b motif, which have similar, but distinct secondary structures. Together, these motifs are found in Bacteroidota, Chlorobiota, and Pseudomonadota.
The Fusobacteriales-1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. Fusobacteriales-1 motif RNAs are found in Fusobacteriales.

The GA-cis RNA motif is a conserved RNA structure that was discovered by bioinformatics. GA-cis motif RNAs are found in one species classified within the phylum Bacillota: specifically, there are 9 predicted copies in Coprocuccus eutactus ATCC 27759.
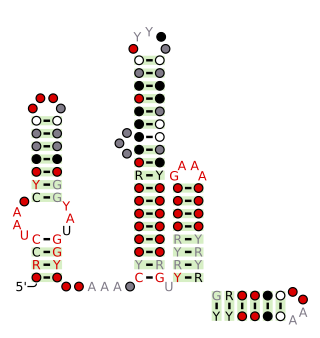
The Latescibacteria, OD1, OP11, TM7 RNA motif is a conserved RNA structure that was discovered by bioinformatics. LOOT motif RNAs are found in multiple bacterial phyla that have only recently been discovered, and are currently not well understood: Latescibacteria, OD1/Parcubacteria, OP11 AND TM7. In some cases, no specific organism has been isolated in the relevant phylum, but the existence of the bacterial phylum is known only through analysis of metagenomic sequences. Curiously, the LOOT motif is not known in any phylum that has been studied for a long time.
lysM RNA motifs are conserved RNA structures that were discovered by bioinformatics. Such bacterial motifs are defined by consistently being upstream of 'lysM' genes, which encode lysin protein domains, a conserved domain that participates in cell wall degradation. lysM motif RNAs likely function as cis-regulatory elements, in view of their positions upstream of protein-coding genes, although this hypothesis is not certain.
The Mahella-1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. Seven Mahella-1 motif RNAs are found in Mahella australiensis, and no other organism has been observed to contain Mahella-1 RNAs.
malK RNA motifs are conserved RNA structures that were discovered by bioinformatics. They are defined by being consistently located upstream of malK genes, which encode an ATPase that is used by transporters whose ligand is likely a kind of sugar. Most of these genes are annotated either as transporting maltose or glycerol-3-phosphate, however the substrate of the transporters associated with malK motif RNAs has not been experimentally determined. All known types of malK RNA motif are generally located nearby to the Shine-Dalgarno sequence of the downstream gene.

The MDR-NUDIX RNA motif is a conserved RNA structure that was discovered by bioinformatics. The MDR-NUDIX motif is found in the poorly studied phylum TM7.

The nhaA-I RNA motif is a conserved RNA structure that was discovered by bioinformatics. nhaA-I motif RNAs are found in Acidobacteriota, alpha-, beta- and Gammaproteobacteria, Verrucomicrobiota and the tentative phylum NC10.

The NLPC-P60 RNA motif is a conserved RNA structure that was discovered by bioinformatics. NLPC-P60 motif RNAs are found in Streptomyces.
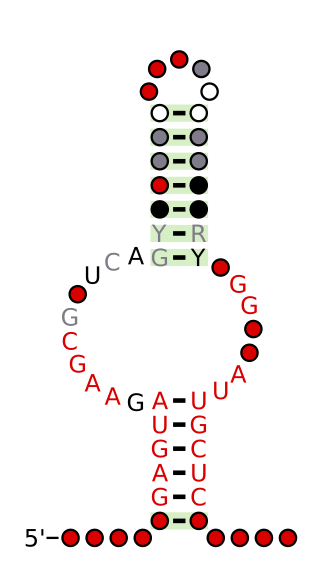
The NMT1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. NMT1 motif RNAs are found in Pseudomonadota. There is also one NMT1 RNA in each of Bacteroidota and Actinomycetota, but these appear to be the result of recent horizontal gene transfer or sequence contamination before or during genome sequencing
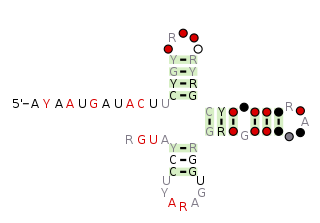
The pemK RNA motif is a conserved RNA structure that was discovered by bioinformatics. pemK motif RNAs are found in organisms within the phylum Bacillota, and is very widespread in this phylum.

The PGK RNA motif is a conserved RNA structure that was discovered by bioinformatics. PGK motif RNAs are found in metagenomic sequences isolated from the gastrointestinal tract of mammals. PGK RNAs have not yet been detected in a classified organism.
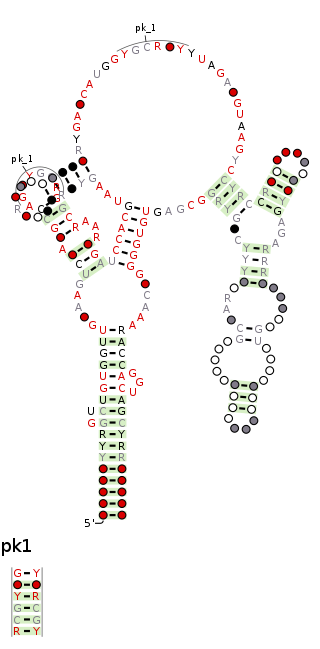
The raiA RNA motif is a conserved RNA structure that was discovered by bioinformatics. raiA motif RNAs are found in Actinomycetota and Bacillota, and have many conserved features—including conserved nucleotide positions, conserved secondary structures and associated protein-coding genes—in both of these phyla. Some conserved features of the raiA RNA motif suggest that they function as cis-regulatory elements, but other aspects of the motif suggest otherwise.

The ssnA RNA motif is a conserved RNA structure that was discovered by bioinformatics. ssnA motif RNAs are found in Clostridiales.

The sul1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. Energetically stable tetraloops often occur in this motif. sul1 motif RNAs are found in Alphaproteobacteria.

The terC RNA motif is a conserved RNA structure that was discovered by bioinformatics. terC motif RNAs are found in Pseudomonadota, within the sub-lineages Alphaproteobacteria and Pseudomonadales.

The xerDC RNA motif is a conserved RNA structure that was discovered by bioinformatics. xerDC motif RNAs are found in Clostridia.
















